Abstract
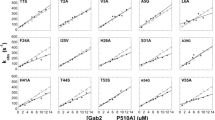
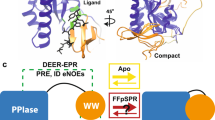
p13suc1 (suc1) has two native states, a monomer and a domain-swapped dimer. The structure of each subunit in the dimer is identical to that of the monomer, except for the hinge loop that connects the exchanging domains. Here we find that single point mutations at sites throughout the protein and ligand binding both shift the position of the equilibrium between monomer and dimer. The hinge loop was shown previously to act as a loaded molecular spring that releases tension present in the monomer by adopting an alternative conformation in the dimer. The results here indicate that the release of strain propagates throughout the entire protein and alters the energetics of regions remote from the hinge. Our data illustrate how the signal conferred by the conformational change of a protein loop, elicited by domain swapping, ligand binding or mutation, can be sensed by a distant active site. This work highlights the potential role of strained loops in proteins: the energy they store can be used for both signal transduction and allostery, and they could steer the evolution of protein function. Finally, a structural mechanism for the role of suc1 as an adapter molecule is proposed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bennett, M.J., Choe, S. & Eisenberg, D. Protein Sci. 3, 1444–1463 (1994).
Bennett, M.J., Choe, S. & Eisenberg, D. Proc. Natl. Acad. Sci. USA 91, 3127–3131 (1994).
Endicott, J.A. et al. EMBO J. 14, 1004–1014 (1995).
Bourne, Y. et al. Proc. Natl. Acad. Sci. USA 92, 10232–10236 (1995).
Khazanovich, N., Bateman, K.S., Chernaia, M., Michalak, M. & James, M.N.G. Structure 4, 299–309 (1996).
Green, S.M., Gittis, A.G., Meeker, A.K. & Lattman, E.E. Nature Strut. Biol. 2, 746–751 (1995).
SaintJean, A.P., Phillips, K.R., Creighton, D.J. & Stone, M.J. Biochemistry 37, 10345–10353 (1998).
Vitagliano, L. et al. J. Mol. Biol. 293, 569–577 (1999).
Qu, C. et al. Struct. Folding Des. 8, 1095–1103 (2000).
Liu, Y., Gotte, G., Libonati, M. & Eisenberg, D. A. Nature Strut. Biol. 8, 211–214. (2001).
Ogihara, N.L. et al. Proc. Natl. Acad. Sci. USA 98, 1404–1409. (2001).
Janowski, R. et al. Nature Strut. Biol. 8, 316–320. (2001).
Murray, A.J., Head, J.G., Barker, J.J. & Brady, R.L. Nature Struct. Biol. 5, 778–782 (1998).
Hayes, M.V., Sessions, R.B., Brady, R.L. & Clarke, A.R. J. Mol. Biol. 285, 1857–1867 (1999).
Rousseau, F., Schymkowitz, J.W.H., Wilkinson, H.R. & Itzhaki, L.S. Proc. Natl. Acad. Sci. USA 98, 5596–5601 (2001).
Landrieu, I. et al. J. Biol. Chem. 276, 1434–1438 (2001).
Schymkowitz, J.W.H., Rousseau, F. & Itzhaki, L.S. J. Mol. Biol. 301, 201–206 (2000).
Hilser, V.J. & Freire, E. J. Mol. Biol. 262, 756–772 (1996).
Freire, E. Proc. Natl. Acad. Sci. USA 96, 10118–10122 (1999).
Freire, E. Proc. Natl. Acad. Sci. USA 97, 11680–11682 (2000).
Todd, M.J. & Freire, E. Proteins 36, 147–156 (1999).
Pines, J. Curr. Biol. 11, 1399–1402 (1996).
Patra, D. & Dunphy, W.G. Genes Dev. 12, 2549–2559 (1998).
Patra, D., Wang, S.X., Kumagai, A. & Dunphy, W.G. J. Biol. Chem. 274, 36839–36842 (1999).
Bourne, Y. et al. Cell 84, 863–874 (1996).
Schymkowitz, J.W.H., Rousseau, F., Irvine, L.R. & Itzhaki, L.S. Struct. Folding Des. 8, 89–100 (2000).
Rousseau, F., Schymkowitz, J.W.H., delPino, M.S. & Itzhaki, L.S. J. Mol. Biol. 284, 503–519 (1998).
Kraulis, P.J. J. Appl. Crystallogr. 24, 946–950 (1991).
Alonso, D.O.V., Alm, E. & Daggett, V. Struct. Folding Des. 8, 101–110.
Parge, H.E., Arvai, A.S., Murtari, D.J., Reed, S.I. & Tainer, J.A. Science 262, 387–394 (1993).
Acknowledgements
This work was supported by the MRC of the UK. L.S.I. was supported by a Career Development Award from the MRC of the UK. J.W.H.S. and F.R. were supported by Marie Curie Training and Mobility of Research Fellowships from the E.C. A.F. is supported by a Human Frontier Science Program long term fellowship. We thank A.R. Fersht and J. Clarke for useful discussions and S. Kazmirski for help with analyzing the crystal structures.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Schymkowitz, J., Rousseau, F., Wilkinson, H. et al. Observation of signal transduction in three-dimensional domain swapping. Nat Struct Mol Biol 8, 888–892 (2001). https://doi.org/10.1038/nsb1001-888
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb1001-888
This article is cited by
-
Interaction of dimeric horse cytochrome c with cyanide ion
JBIC Journal of Biological Inorganic Chemistry (2013)
-
Functional repertoire, molecular pathways and diseases associated with 3D domain swapping in the human proteome
Journal of Clinical Bioinformatics (2012)
-
Structural basis for ADP-mediated transcriptional regulation by P1 and P7 ParA
The EMBO Journal (2009)
-
On the origin of the histone fold
BMC Structural Biology (2007)
-
Cooperative organization in a macromolecular complex
Nature Structural & Molecular Biology (2003)