Abstract
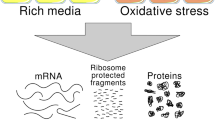
Transcription and mRNA turnover determine the quantitative composition of the cellular transcriptome. The transcriptome in turn serves as a template for the proteome via translation. Treatment of Saccharomyces cerevisiae with the TOR kinase inhibitor rapamycin causes increases and decreases in the mRNA levels of hundreds of genes. We used DNA microarray analysis to monitor simultaneously transcriptome and translational changes for all detectable yeast mRNAs. Notably, genes that are induced in the transcriptome correlate tightly with more efficiently translated mRNAs (based on their relative degree of polyribosome association); similarly, genes that show reduced mRNA levels after rapamycin treatment also show lower translational fitness. Microarray analyses on heat-shocked cells also reveal homodirectional co-regulatory responses. Thus, signal-induced changes in the transcriptome are amplified at the translational level. These results unveil a higher level of coordinated gene regulation that we refer to as 'potentiation.'
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Maniatis, T. & Reed, R. An extensive network of coupling among gene expression machines. Nature 416, 499–506 (2002).
Keene, J.D. Ribonucleoprotein infrastructure regulating the flow of genetic information between the genome and the proteome. Proc. Natl. Acad. Sci. USA 98, 7018–7024 (2001).
Jensen, T.H. & Rosbash, M. Co-transcriptional monitoring of mRNP formation. Nat. Struct. Biol. 10, 10–12 (2003).
Manley, J.L. Nuclear coupling: RNA processing reaches back to transcription. Nat. Struct. Biol. 9, 790–791 (2002).
Proudfoot, N.J., Furger, A. & Dye, M.J. Integrating mRNA processing with transcription. Cell 108, 501–512 (2002).
Fong, N. & Bentley, D.L. Capping, splicing, and 3′ processing are independently stimulated by RNA polymerase II: different functions for different segments of the CTD. Genes Dev. 15, 1783–1795 (2001).
Lei, E.P. & Silver, P.A. Intron status and 3′-end formation control cotranscriptional export of mRNA. Genes Dev. 16, 2761–2766 (2002).
Strasser, K. et al. TREX is a conserved complex coupling transcription with messenger RNA export. Nature 417, 304–308 (2002).
Zenklusen, D., Vinciguerra, P., Wyss, J.C. & Stutz, F. Stable mRNP formation and export require cotranscriptional recruitment of the mRNA export factors Yra1p and Sub2p by Hpr1p. Mol. Cell. Biol. 22, 8241–8253 (2002).
Hentze, M.W. & Kulozik, A.E. A perfect message: RNA surveillance and nonsense-mediated decay. Cell 96, 307–310 (1999).
Maquat, L.E. & Carmichael, G.G. Quality control of mRNA function. Cell 104, 173–176 (2001).
Li, S. & Wilkinson, M.F. Nonsense surveillance in lymphocytes? Immunity 8, 135–141 (1998).
Gygi, S.P., Rochon, Y., Franza, B.R. & Aebersold, R. Correlation between protein and mRNA abundance in yeast. Mol. Cell. Biol. 19, 1720–1730 (1999).
Ideker, T. et al. Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 292, 929–934 (2001).
Mathews, M.B., Sonenberg, N. & Hershey, J.B.W. Origins and principles of translational control. In Translational Control of Gene Expression (eds. Sonenberg, N., Hershey, J.B.W. & Mathews, M.B.) 1–31 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2000).
Pradet-Balade, B., Boulme, F., Beug, H., Mullner, E.W. & Garcia-Sanz, J.A. Translation control: bridging the gap between genomics and proteomics? Trends Biochem. Sci. 26, 225–229 (2001).
Gray, N.K. & Wickens, M. Control of translation initiation in animals. Annu. Rev. Cell. Dev. Biol. 14, 399–458 (1998).
Kuhn, K.M., DeRisi, J.L., Brown, P.O. & Sarnow, P. Global and specific translational regulation in the genomic response of Saccharomyces cerevisiae to a rapid transfer from a fermentable to a nonfermentable carbon source. Mol. Cell. Biol. 21, 916–927 (2001).
Brown, V. et al. Microarray identification of FMRP-associated brain mRNAs and altered mRNA translational profiles in fragile X syndrome. Cell 107, 477–487 (2001).
Zong, Q., Schummer, M., Hood, L. & Morris, D.R. Messenger RNA translation state: the second dimension of high-throughput expression screening. Proc. Natl. Acad. Sci. USA 96, 10632–10636 (1999).
Grolleau, A. et al. Global and specific translational control by rapamycin in T cells uncovered by microarrays and proteomics. J. Biol. Chem. 277, 22175–22184 (2002).
Muckenthaler, M. et al. Relationships and distinctions in iron regulatory networks responding to interrelated signals. Blood 101, 3690–3698 (2002).
Barbet, N.C. et al. TOR controls translation initiation and early G1 progression in yeast. Mol. Biol. Cell 7, 25–42 (1996).
Powers, T. & Walter, P. Regulation of ribosome biogenesis by the rapamycin-sensitive TOR- signaling pathway in Saccharomyces cerevisiae. Mol. Biol. Cell 10, 987–1000 (1999).
Shamji, A.F., Kuruvilla, F.G. & Schreiber, S.L. Partitioning the transcriptional program induced by rapamycin among the effectors of the Tor proteins. Curr. Biol. 10, 1574–1581 (2000).
Albig, A.R. & Decker, C.J. The target of rapamycin signaling pathway regulates mRNA turnover in the yeast Saccharomyces cerevisiae. Mol. Biol. Cell 12, 3428–3438 (2001).
Berset, C., Trachsel, H. & Altmann, M. The TOR (target of rapamycin) signal transduction pathway regulates the stability of translation initiation factor eIF4G in the yeast Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 95, 4264–4269 (1998).
Wang, Y. et al. Precision and functional specificity in mRNA decay. Proc. Natl. Acad. Sci. USA 99, 5860–5865 (2002).
Gasch, A.P. et al. Genomic expression programs in the response of yeast cells to environmental changes. Mol. Biol. Cell 11, 4241–4257 (2000).
Schneider, R.J. Translational control during heat shock. In Translational Control of Gene Expression (eds. Sonenberg, N., Hershey, J.W.B. & Mathews, M.B.) 581–593 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2000).
Kedersha, N. et al. Evidence that ternary complex (eIF2-GTP-tRNA(i)(Met))-deficient preinitiation complexes are core constituents of mammalian stress granules. Mol. Biol. Cell 13, 195–210 (2002).
Kimball, S.R., Horetsky, R.L., Ron, D., Jefferson, L.S. & Harding, H.P. Mammalian stress granules represent sites of accumulation of stalled translation initiation complexes. Am. J. Physiol. Cell. Physiol. 284, C273–C284 (2003).
Dunand-Sauthier, I. et al. Sum1, a component of the fission yeast eIF3 translation initiation complex, is rapidly relocalized during environmental stress and interacts with components of the 26S proteasome. Mol. Biol. Cell 13, 1626–1640 (2002).
Sachs, A.B. & Varani, G. Eukaryotic translation initiation: there are (at least) two sides to every story. Nat. Struct. Biol. 7, 356–361 (2000).
Schwartz, D.C. & Parker, R. Interaction of mRNA translation and mRNA degradation in Saccharomyces cerevisiae. In Translational Control of Gene Expression (eds. Sonenberg, N., Hershey, J.W.B. & Mathews, M.B.) 807–825 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2000).
Mitchell, P. & Tollervey, D. mRNA turnover. Curr. Opin. Cell. Biol. 13, 320–325 (2001).
Preiss, T. & Hentze, M.W. Starting the protein synthesis machine: eukaryotic translation initiation. BioEssays 25, 1201–1210 (2003).
Richter, A. et al. Comparison of fluorescent tag DNA labeling methods used for expression analysis by DNA microarrays. Biotechniques 33, 620–628, 630 (2002).
Holstege, F.C. et al. Dissecting the regulatory circuitry of a eukaryotic genome. Cell 95, 717–728 (1998).
Acknowledgements
We thank all members of the EMBL microarray community, past and present, for fruitful interactions throughout this work, and for their contributions to the establishment of microarray technology at EMBL. We also thank M. Altmann, J.E.G. McCarthy, A.B. Sachs, B. Seraphin, G. Kornfeld, T. Lithgow, M. Knop and M. Seedorf for the gift of strains and reagents. This work was funded by grants from the Deutsche Forschungsgemeinschaft to T.P. and M.W.H. and from the Human Frontiers in Science Program to M.W.H.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
41594_2003_BFnsb1015_MOESM1_ESM.pdf
Supplementary Fig. 1. Correlation of mRNA translation state with length of the open reading frame (ORF). Translation state ratios from the control cell measurement in array set I were plotted against ORF size and a Lowess curve was fitted to all data points (solid line). The mRNA frequency in a given length interval (size: 200 amino acids) is shown for comparison (dashed line). The correlation breaks down beyond 2500 amino acids as there is only a statistically insignificant number of scored genes (n=12 in total) in this size region. (PDF 16 kb)
41594_2003_BFnsb1015_MOESM2_ESM.pdf
Supplementary Fig. 2. Selection of mRNA profiles. a, mRNAs were selected for translational state ratio change in response to rapamycin from array data set I as detailed in the main text and Fig. 3A-C. b, mRNAs were selected based on their polysome re-distribution profile from array data set II. Profiles were sorted into a 9x8 matrix using a self-organising map algorithm (GeneSpring 5.0) and then assigned to different candidate groups for translational behaviour as indicated by the coloured frames. (PDF 188 kb)
Rights and permissions
About this article
Cite this article
Preiss, T., Baron-Benhamou, J., Ansorge, W. et al. Homodirectional changes in transcriptome composition and mRNA translation induced by rapamycin and heat shock. Nat Struct Mol Biol 10, 1039–1047 (2003). https://doi.org/10.1038/nsb1015
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb1015
This article is cited by
-
MCHM Acts as a Hydrotrope, Altering the Balance of Metals in Yeast
Biological Trace Element Research (2020)
-
Dynamic changes in eIF4F-mRNA interactions revealed by global analyses of environmental stress responses
Genome Biology (2017)
-
The translational landscape of fission-yeast meiosis and sporulation
Nature Structural & Molecular Biology (2014)
-
Widespread uncoupling between transcriptome and translatome variations after a stimulus in mammalian cells
BMC Genomics (2012)
-
Low level genome mistranslations deregulate the transcriptome and translatome and generate proteotoxic stress in yeast
BMC Biology (2012)