Abstract
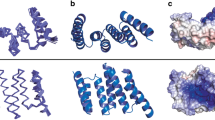
H-NS, a protein found in Gram-negative bacteria, is involved in structuring the bacterial chromosome and acts as a global regulator for the expression of a wide variety of genes. These functions are correlated with both its DNA-binding and oligomerization properties. We have identified the minimal dimerization domain of H-NS, a 46 amino acid–long N-terminal fragment, and determined its structure using heteronuclear NMR spectroscopy. The highly intertwined structure of the dimer, reminiscent of a handshake, defines a new structural fold, which may offer a possibility for discriminating prokaryotic from eukaryotic proteins in drug design. Using mutational analysis, we also show that this N-terminal domain actively contributes to DNA binding, conversely to the current paradigm. Together, our data allows us to propose a model for the action of full length H-NS.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hommais, F. et al. Large-scale monitoring of pleiotropic regulation of gene expression by the prokaryotic nucleoid-associated protein, H-NS. Mol. Microbiol. 40, 20–36 (2001).
Tanaka, K., Muramatsu, S., Yamada, H. & Mizuno, T. Systematic characterization of curved DNA segments randomly cloned from Escherichia coli and their functional significance. Mol. Gen. Genet. 226, 367–376 (1991).
Yamada, H., Yoshida, T., Tanaka, K., Sasakawa, C. & Mizuno, T. Molecular analysis of the Escherichia coli hns gene encoding a DNA-binding protein, which preferentially recognizes curved DNA sequences. Mol. Gen. Genet. 230, 332–336 (1991).
Rimsky, S., Zuber, F., Buckle, M. & Buc, H. A molecular mechanism for the repression of transcription by the H-NS protein. Mol. Microbiol. 42, 1311–1323 (2001).
Tupper, A.E. et al. The chromatin-associated protein H-NS alters DNA topology in vitro. EMBO J. 13, 258–268 (1994).
Falconi, M., Colonna, B., Prosseda, G., Micheli, G. & Gualerzi, C.O. Thermoregulation of Shigella and Escherichia coli EIEC pathogenicity. A temperature-dependent structural transition of DNA modulates accessibility of virF promoter to transcriptional repressor H-NS. EMBO J. 17, 7033–7043 (1998).
Jordi, B.J. & Higgins, C.F. The downstream regulatory element of the proU operon of Salmonella typhimurium inhibits open complex formation by RNA polymerase at a distance. J. Biol. Chem. 275, 12123–12128 (2000).
Williams, R.M., Rimsky, S. & Buc, H. Probing the structure, function, and interactions of the Escherichia coli H-NS and StpA proteins by using dominant negative derivatives. J. Bacteriol. 178, 4335–4343 (1996).
Ueguchi, C., Suzuki, T., Yoshida, T., Tanaka, K. & Mizuno, T. Systematic mutational analysis revealing the functional domain organization of Escherichia coli nucleoid protein H-NS. J. Mol. Biol. 263, 149–162 (1996).
Badaut, C. et al. The degree of oligomerisation of the H-NS nucleoid structuring protein is related to specific binding to DNA. J. Biol. Chem. 277, 41657–41666 (2002).
Shindo, H. et al. Solution structure of the DNA binding domain of a nucleoid-associated protein, H-NS, from Escherichia coli. FEBS Lett. 360, 125–131 (1995).
Smyth, C.P. et al. Oligomerization of the chromatin-structuring protein H-NS. Mol. Microbiol. 36, 962–972 (2000).
Spassky, A. & Buc, H.C. Physico-chemical properties of a DNA binding protein: Escherichia coli factor H1. Eur. J. Biochem. 81, 79–90 (1977).
Falconi, M., Gualtieri, M.T., La Teana, A., Losso, M.A. & Pon, C.L. Proteins from the prokaryotic nucleoid: primary and quaternary structure of the 15-kD Escherichia coli DNA binding protein H-NS. Mol. Microbiol. 2, 323–329 (1988).
Ceschini, S. et al. Multimeric self-assembly equilibria involving the histone-like protein H-NS. A thermodynamic study. J. Biol. Chem. 275, 729–734 (2000).
Ueguchi, C., Suzuki, T., Yoshida, T., Tanaka, K. & Mizuno, T. Systematic mutational analysis revealing the functional domain organization of Escherichia coli nucleoid protein H-NS. J. Mol. Biol. 263, 149–162 (1996).
Renzoni, D. et al. Structural characterization of the N-terminal oligomerization domain of the bacterial chromatin-structuring protein, H-NS. J. Mol. Biol. 306, 1127–1137 (2001).
Holm, L. & Sanders, C. DALI. Nucleic Acids Res. 25, 231–234 (1997).
Jacquet, M., Cukier-Kahn, R., Pla, J. & Gros, F. A thermostable protein factor acting on in vitro DNA transcription. Biochem. Biophys. Res. Commun. 45, 1597–1607 (1971).
Shindo, H. et al. Identification of the DNA binding surface of H-NS protein from Escherichia coli by heteronuclear NMR spectroscopy. FEBS Lett. 455, 63–639 (1999).
Schroder, O., Tippner, D. & Wagner, R. Toward the three-dimensional structure of the Escherichia coli DNA-binding protein H-NS: a CD and fluorescence study. Biochem. Biophys. Res. Commun. 282, 219–227 (2001).
Rastinejad, F., Perlmann, T., Evans, R.M. & Sigler, P.B. Structural determinants of nuclear receptor assembly on DNA direct repeats. Nature 375, 203–211 (1995).
Williams, R.M. & Rimsky, S. Molecular aspects of the E. coli nucleoid protein, H-NS: a central controller of gene regulatory networks. FEMS Microbiol. Lett. 156, 175–185 (1997).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory Press, Cold Spring Harbor; 1989).
Aumelas, A. et al. Synthesis and solution structure of the antimicrobial peptide protegrin-1. Eur. J. Biochem. 237, 575–583 (1996).
Beechem, J.M., Gratton, E., Ameloot, M.A., Knutson, J.R. & Brand, L. The global analysis of fluorescence intensity and anisotropy decay data: second-generation theory and programs. in Fluorescence Spectroscopy: Principles and Techniques, Vol. 2 (ed. Lakowicz, J.R.) 241–301 (Plenum Publishing, New York; 1991).
Fernando, T. & Royer, C. Role of protein–protein interactions in the regulation of the transcription by trp repressor investigated by fluorescence spectroscopy. Biochemistry 31, 3429–3441 (1992).
Pons, J.L., Malliavin, T. & Delsuc, M. Gifa V4: a complete package for NMR data-set processing. J. Biomol. NMR 8, 445–452 (1996).
Ikura, M. & Bax, A. Isotope-filtered 2D NMR of a protein-peptide complex: study of a skeletal muscle myosin light chain kinase fragment bound to calmodulin. J. Am. Chem. Soc. 114, 2433–2440 (1992).
Brunger, A. X-PLOR (Version 3.1): a system for X-ray crystallography and NMR. (Yale University Press, New Haven; 1992).
Nilges, M., Macias, M.J., O'Donoghue, S.I. & Oschkinat, H. Automated NOESY interpretation with ambiguous distance restraints: the refined NMR solution structure of the pleckstrin homology domain from β-spectrin. J. Mol. Biol. 269, 408–422 (1997).
Kraulis, P. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
J.P. Simore is greatly acknowledged for his help with the 800 NMR spectrometer; H. Buc, for constant support and advice; and M. Buckle, for numerous invaluable suggestions and critical reading of this manuscript. This work was supported by a grant from the Programme de Recherche en Biologie Fondamentale en Microbiologie et Maladies infectieuses, a doctoral fellowship grant (V.B.) from the MNERT and a EEC-TMR grant (M.K.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Bloch, V., Yang, Y., Margeat, E. et al. The H-NS dimerization domain defines a new fold contributing to DNA recognition. Nat Struct Mol Biol 10, 212–218 (2003). https://doi.org/10.1038/nsb904
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb904
This article is cited by
-
Mechanism of phase condensation for chromosome architecture and function
Experimental & Molecular Medicine (2024)
-
c-di-GMP inhibits the DNA binding activity of H-NS in Salmonella
Nature Communications (2023)
-
The environmentally-regulated interplay between local three-dimensional chromatin organisation and transcription of proVWX in E. coli
Nature Communications (2023)
-
A binding cooperativity switch driven by synergistic structural swelling of an osmo-regulatory protein pair
Nature Communications (2019)
-
Identifying the region responsible for Brucella abortus MucR higher-order oligomer formation and examining its role in gene regulation
Scientific Reports (2018)