Abstract
Riboswitches are metabolite-binding RNA structures that serve as genetic control elements for certain messenger RNAs. These RNA switches have been identified in all three kingdoms of life and are typically responsible for the control of genes whose protein products are involved in the biosynthesis, transport or utilization of the target metabolite. Herein, we report that a highly conserved RNA domain found in bacteria serves as a riboswitch that responds to the coenzyme S-adenosylmethionine (SAM) with remarkably high affinity and specificity. SAM riboswitches undergo structural reorganization upon introduction of SAM, and these allosteric changes regulate the expression of 26 genes in Bacillus subtilis. This and related findings indicate that direct interaction between small metabolites and allosteric mRNAs is an important and widespread form of genetic regulation in bacteria.
Similar content being viewed by others
Main
Riboswitches are structural domains embedded within the noncoding sequences of certain mRNAs that serve as metabolite-responsive genetic control elements1,2,3,4. Riboswitches exhibit two surprising functions that are not typically associated with natural RNAs. First, the mRNA element can adopt distinct structural states wherein one structure serves as a precise binding pocket for its target metabolite. Second, the metabolite-induced allosteric interconversion between structural states causes a change in the level of gene expression by one of several distinct mechanisms. Riboswitches typically can be dissected into two separate domains: one that selectively binds the target (aptamer domain) and another that influences genetic control (expression platform)2,3. The dynamic interplay between these two domains results in metabolite-dependent allosteric control of gene expression.
Three distinct classes of riboswitches have been identified previously, and were shown to selectively recognize coenzyme B12 (ref. 1), thiamine pyrophosphate (TPP)2,4,5 or flavin mononucleotide (FMN)3,4. The aptamer domain of each riboswitch class conforms to a highly conserved consensus sequence and structure. Thus, sequence homology searches have been used to reveal the presence of related riboswitch-like domains in various organisms from bacteria, archaea and eukarya5,6,7,8,9. Most notably, sequence elements that correspond to the riboswitch aptamer domain for thiamine pyrophosphate (TPP) occur within certain mRNAs from many bacteria2,4, but they are also present in mRNAs corresponding to thiamine-related genes from fungi and plants5. These findings support the hypothesis that riboswitches emerged early in evolution, and suggest that metabolite-sensing RNAs could be representatives of an ancient gene regulation system. Specifically, riboswitch domains might have emerged during the RNA world—a proposed phase in evolution in which all biological processes that were necessary for primitive life were guided by RNA10,11.
We report here that a highly conserved RNA domain termed the S box serves as a selective and high-affinity aptamer for SAM. Allosteric modulation of secondary and tertiary structures is induced upon SAM binding to the aptamer domain, and these structural changes are responsible for inducing termination of mRNA transcription. These findings provide additional support that RNA structures can adopt a wide range of sophisticated structures that can function as precision genetic switches.
Results
Identification of a SAM-responsive riboswitch
Each of the compounds sensed by previously identified riboswitches (coenzyme B12, TPP and FMN) is used as a coenzyme by modern protein enzymes. Notably, these coenzymes have substantial structural similarity to RNA and this has been used to support speculation that they might also have been used as coenzymes by ancient ribozymes in an RNA world10,12,13. If modern riboswitches are direct descendants of RNA control systems that originated in the RNA world, then we would expect that the metabolites they sense and the metabolic pathways that they control would be of fundamental importance to modern biochemical processes. To further assess this hypothesis, we have sought to identify additional riboswitches, to determine their biochemical characteristics, and to establish their role in genetic control on a genome-wide level.
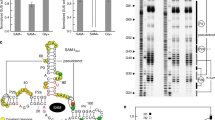
In this effort we have examined the S box14, a highly conserved sequence domain (Fig. 1a) that is located within the 5′-untranslated region (5′ UTR) of certain messenger RNAs in Gram-positive bacteria. Both genetic and sequence analyses suggest that the S-box domain serves as a genetic control element for a regulon composed of 11 transcriptional units. These mRNAs encode as many as 26 different genes in B. subtilis that are involved in sulfur metabolism, methionine biosynthesis, cysteine biosynthesis and SAM biosynthesis. However, the nature of the putative regulatory factor15,16 and the metabolite to which it responds had not been established until now. We prepared an RNA construct corresponding to the first 251 nucleotides of the yitJ mRNA of B. subtilis (Fig. 1b) by in vitro transcription. The yitJ gene product is a putative methylene tetrahydrofolate reductase, an enzyme proposed to be involved in methionine biosynthesis14. The 251 yitJ RNA was subjected to 'in-line probing'17, which reveals locations of structured and unstructured portions of RNA polymers by relying on the variability in rates of spontaneous RNA phosphodiester cleavage caused by differences in structural context. In-line probing also can reveal nucleotides participating in metabolite-induced structural modulation1,2,3,5.
(a) Consensus sequence and secondary-structure model of the S-box domain derived from 107 bacterial representatives (see Supplementary Table 1). Red and black positions, nucleotides whose identity as depicted is conserved in >90% or >80% of the representative S-box RNAs, respectively. R, Y and N: purine, pyrimidine and any nucleotide, respectively. P1–P4 identify conserved base pairing. Encircled nucleotides indicate a putative pseudoknot interaction. (b) Sequence and secondary structure model for the 251 yitJ mRNA fragment. Sites of structural modulation upon introduction of SAM are depicted as described. Nucleotide 1, putative transcriptional start site14. Asterisks, nucleotides that were added to the construct to permit efficient transcription in vitro. The first nucleotide of the AUG start codon is 235 (not shown). Other notations are as described in a. (c) Spontaneous cleavage patterns of 251 yitJ in the absence (−) or presence (+) of methionine or SAM as indicated (see Methods for details). NR, T1 and −OH represent no reaction, partial digest with RNase T1, and partial digest with alkali, respectively. Certain fragment bands corresponding to digestion by T1 (which cleaves after guanine residues) are depicted. Blue arrowheads, positions of substantial modulation of spontaneous cleavage; the numbered sites were used for quantification (see Fig. 2b).
Given that previously identified riboswitches recognize coenzymes, we chose to examine whether the 251 yitJ RNA might bind SAM. Indeed, upon separation by PAGE, the pattern of spontaneous RNA cleavage products (Fig. 1c) is indicative of a highly structured RNA element that undergoes conformational modulation upon introduction of SAM to a final concentration of either 0.1 mM or 1 mM. In contrast, no structural modulation is evident upon the introduction of methionine at the same concentrations, suggesting that the RNA might require both the methionine and 5′-deoxyadenosyl moieties of SAM to induce structural reorganization. The locations of the ligand-induced modulations (Fig. 1b) indicate that the conserved core of the S-box RNA serves as a natural aptamer18 for SAM. Similar results were observed with 124 yitJ (data not shown), which encompasses nucleotides 28 through 149 of the mRNA leader plus two guanine residues at the 5′ terminus. The in-line probing data are consistent with the secondary-structure model (Fig. 1a) that has been proposed for S-box RNAs14. Furthermore, the structural modulation revealed by probing indicates that substantia structural modulation occurs at distal parts of the RNA, which could be harnessed for genetic control purposes.
Molecular recognition by a SAM-dependent riboswitch
A genetic switch that responds to metabolites must be able to bind its target with a dissociation constant (Kd) that is relevant to physiological concentrations. Furthermore, the metabolite receptor must be able to discriminate precisely against closely related compounds that are likely to occur in the same milieu, or risk undesirable modulation of gene expression. Therefore, we assessed the affinity of the yitJ RNA for SAM, and the ability of the RNA to discriminate against biologically relevant compounds that are structurally similar to this target (Fig. 2).
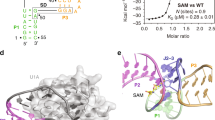
(a) Chemical structures of various compounds used to probe the binding characteristics of the SAM yitJ riboswitch. Other than methionine, each compound as depicted is coupled to an adenosyl moiety (A, inset) via the 5′ carbon (R). (b) Left: Kd plot for the 251 yitJ for SAM (see Methods for details). Right: Kd values for SAM and various analogs. (c) Molecular discrimination determined by equilibrium dialysis (see Methods for details). M1 is a variant of 124 yitJ that carries disruptive mutations in the junction between stems P1 and P2 (Fig. 4a). Notations at top indicate the absence of unlabeled compound (−) or the presence of 25 μM of the compound indicated. Line at a c.p.m. ratio of 1 identifies the bar height expected if a shift in [3H]SAM has not occurred.
The Kd of 251 yitJ for SAM was determined by using in-line probing to monitor the extent of structural modulation over a range of different ligand types (Fig. 2a) and concentrations (Fig. 2b, left). Although the Kd of 251 yitJ for SAM is ∼200 nM, the minimized aptamer domain represented by 124 yitJ exhibits a Kd of ∼4 nM under our assay conditions. Such improvements in binding affinity by minimized aptamer domains have been observed previously2. This most likely reflects greater structural pre-organization of the ligand binding form of the aptamer domain owing to the elimination of the adjoining expression platform, which otherwise would permit alternative folding to occur.
As expected, the 124 yitJ RNA shows a high level of molecular discrimination against analogs of SAM. For example, the RNA exhibits ∼100-fold discrimination against SAH (Fig. 2b, right), which is produced upon utilization of SAM as a coenzyme for methyl-ation reactions19. Thus, the aptamer must form a binding pocket for SAM that can sense the absence of a single methyl group and an associated loss of positive charge. Similarly, the RNA discriminates nearly 10,000-fold against SAC, another biological compound that differs from SAH by the absence of a single methylene group.
This pattern of molecular discrimination was confirmed by using equilibrium dialysis (Fig. 2c). Specifically, tritiated SAM is equally distributed between chambers A and B of a dialysis apparatus when no RNA is introduced into chamber B, or when a nonfunctional mutant is added. In contrast, the addition of excess 124 yitJ RNA to chamber B causes a substantial shift in the distribution of tritium to this side of the membrane. This asymmetric distribution of tritium can be restored by the introduction of excess un-labeled SAM, but not by the addition of other analogs at a concentration of 25 μM. Although the concentrations of unlabeled SAH and SAC in these competition assays are above or near the Kd for these compounds, respectively, a measurable reversal of tritium distribution of the chambers should not be apparent until a higher concentration of competitor is used (see Methods). As expected, the use of 200 μM SAH yields a c.p.m. ratio of 2.7 (data not shown), which indicates that SAH can compete for RNA binding sites when a sufficient concentration of this analog is used.
Tight binding was also observed when the 124 yitJ was tested by using a Scatchard analysis with tritiated SAM (Fig. 3). These data are consistent with a Kd value in the low nanomolar range and also indicate that only one SAM molecule is bound by each RNA. After exhaustive equilibration with excess RNA, 10–15% of the tritium remains in chamber A, indicating that between 20% and 30% of tritiated material is not in the form of SAM (most likely demethylated). Upon correction for 20% unbindable tritium, the Scatchard data correspond to a Kd for SAM of ∼10 nM. This assessment of binding affinity indicates that the Kd for the 124 yitJ aptamer is as much as 1,000-fold improved as compared with that reported recently for a related RNA20. Normal concentrations of SAM in bacteria are typically in the low micromolar range21. Some of this coenzyme pool is probably bound by enzymes, thus the high affinity exhibited by this riboswitch might be needed to sense the concentration of free SAM inside cells. However, it is not yet clear whether the actual Kd value is most meaningful for genetic control, or whether the rate constant for riboswitch-metabolite association establishes whether the associated gene is expressed.
The variable r represents the ratio of bound ligand concentration versus the total RNA concentration and the variable [LF] represents the concentration of free ligand (see Methods for details).
SAM binding by an mRNA is required for genetic regulation
The secondary structure model for the SAM-binding aptamer domain was established using phylogenetic data (see Supplementary Table 1 and ref. 14). To provide further support for this model, we examined the influence of disruptive and compensatory mutations (Fig. 4a) on the binding function of the 124 yitJ RNA, and on SAM-mediated genetic control of a lacZ reporter gene when fused with variant riboswitches based on these mutant aptamers. As expected, mutations that alter the conserved core of the aptamer (M1) or that disrupt base pairing in each of the four major base-paired regions (M2, M4, M6 and M8) largely result in a loss of SAM binding function as determined by equilibrium dialysis (Fig. 4b). Compensatory mutations that restore base pairing in these stems (M3, M5, M7 and M9) restore at least partial binding activity.
(a) Sequence and secondary structure model for the 124 yitJ RNA (see Methods for experimental details). (b) Analysis of SAM-binding function by equilibrium dialysis in the presence of wild-type (WT) and mutant RNAs as denoted. Details are described in the legend to Figure 2c, except that 300 nM RNA was used and all assays were conducted without the addition of unlabeled analogs. (c) In vivo control of β-galactosidase expression in B. subtilis cells transformed with various riboswitch constructs as indicated (see Methods for details). β-galactosidase activities were measured as described2. Cells were grown in glucose minimal media in 0.25 μg ml−1 methionine (−) or in 50 μg ml−1 methionine (+). M6–M9 were not examined in vivo.
It has been shown14 that a growth medium rich in methionine leads to repression of B. subtilis genes that carry the S-box domain. This is most likely due to the ability of the cell to convert methionine into an ample supply of SAM. In the current study, we find that, in all cases tested, the binding function of the mutant correlates with its ability to downregulate an appended reporter gene when presented with excess methionine in otherwise minimal growth media (Fig. 4c). These findings are consistent with the hypothesis that SAM binding to the mRNA is necessary for the genetic regulation of S-box mRNAs.
Transcription termination by SAM riboswitches in B. subtilis
Previous studies suggest that bacterial riboswitches can control gene expression by modulating either transcription termination or translation initiation2,3,4, whereas several putative riboswitches in eukaryotes might use one of several different mechanisms5. In B. subtilis, the SAM-binding aptamer domains typically reside immediately upstream from a putative transcription terminator hairpin14. This suggests that SAM binding most likely induces transcription termination as described previously for FMN- and TPP-dependent riboswitches3,4.
This hypothesis was examined by conducting in vitro transcription in the absence or presence of SAM using 11 DNA templates corresponding to the mRNA leader sequences of the S-box regulon. These assays were simplified by using T7 RNA polymerase instead of the native B. subtilis RNA polymerase. Previously, it was observed that an FMN-dependent riboswitch induces transcription termination even when T7 RNA polymerase is used as a surrogate for the bacterial polymerase3. In this study, we found that the yitJ, yoaD and metK leader constructs exhibit modest transcription termination upon the addition of SAM. More dramatically, the termination product from the metI leader construct increases from ∼12% to nearly 75% upon introduction of SAM (Fig. 5a). In all instances, little or no modulation of transcription termination occurs when the analogs SAH or SAC are added to the reaction. The remaining seven S-box representatives did not exhibit substantial modulation with T7 RNA polymerase, presumably because it serves as an imperfect substitute for the native polymerase. Indeed, SAM-dependent transcription termination is observed with many of these mRNA leader sequences when Escherichia coli or B. subtilis polymerases are used in the assay19. Most likely, the differences between the extent of termination with T7 polymerase (Fig. 5) and the extent of reporter modulation (Fig. 4c) could be at least partially offset inside cells by the use of bacterial polymerase.
(a) In vitro transcription using T7 RNA polymerase results in increased termination of four mRNA leader sequences. Reactions were conducted in the absence (−) or presence (+) of 50 μM of the effector as indicated for each lane. For example, the metI template includes the 5′ UTR and coding sequences through mRNA position 242 (relative to transcription start), whereas the termination site is expected to occur at position 189. Below each gel is indicated the percentage of transcription termination (T) at the expected location relative to the total amount of expected termination plus full length RNA (FL). (b) Sequence and structural model for the metI riboswitch in two structural states. Green and pink residues correspond to the P1 (anti-anti-terminator) and the terminator stems, respectively. The orange residues correspond to the anti-terminator stem. Sequences boxed in red define the location and identity of mutations used to examine the proposed mechanism of genetic control. Gel: analysis of mutant metI riboswitches wherein disruptive (Ma, Mab and Mc) or the corresponding compensatory mutations (Mabc) have been inserted. The metI mutant templates and wild-type control template (WT) are identical to the templates used in a, except that the FL product is 220 nucleotides. Other notations are as described in a.
The mechanism of SAM-induced termination (Fig. 5b) most likely involves the ligand-mediated formation of alternative hairpin structures that permit transcriptional read-through (anti-terminator formation without SAM) or that cause termination (terminator formation with SAM). Such a transcription terminator mechanism has been invoked recently to explain the function of an S-box RNA of the B. subtilis metE gene22. Independently, we examined this mechanism by generating several mutant metI constructs that carry disruptive or compensatory changes in the expression platform (Fig. 5b). SAM causes an additional ∼20% yield in transcription termination in a mutant (Mabc) that carries six mutations relative to the wild-type metI riboswitch, which retains proper terminator and anti-terminator base complementation. However, incomplete representation of these six mutations that do not permit normal pairing interactions to occur allows little or no SAM-mediated transcription modulation. Furthermore, mutations that disrupt terminator stem formation (Ma) yield lower levels of termination, whereas mutations that disrupt anti-terminator stem formation (Mab, Mc) yield higher levels of termination (Fig. 5b). These findings indicate that the RNA structural modulation induced by SAM binding mediates genetic control by sequestering an anti-terminator sequence, and thus favors the formation of a transcriptional terminator hairpin.
Riboswitches and fundamental biochemical control
Our results provide more direct biochemical evidence that the S-box motif serves as a SAM-binding aptamer, and that this aptamer functions as part of a SAM-dependent riboswitch that controls the expression of the yitJ mRNA. This evidence is consistent with the findings of two recent reports19,22, which implicate the existence of SAM-dependent riboswitches on a total of 11 mRNAs in B. subtilis. The 11 transcriptional units that comprise the regulon controlled by SAM riboswitches14 seem to encompass at least 26 genes that are central to sulfur metabolism, amino acid metabolism and SAM biosynthesis. Although all 11 transcriptional units from B. subtilis carry a consensus S-box element, a recent report indicates that gene expression from one of these (cysH) is not modulated by addition of methionine to the medium, as are other S-box RNAs23. In contrast, a recent study has demonstrated that the cysH RNA undergoes SAM-induced transcription termination when high concentrations of the ligand are used19, which suggests that the binding affinity of this riboswitch might be shifted higher relative to other examples in B. subtilis.
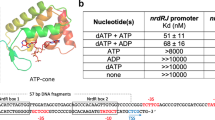
We find that the aptamer domain from B. subtilis cysH does bind SAM with an affinity that is more than two orders of magnitude poorer than that of yitJ from the same organism (Fig. 6a). However, the cysH homolog from B. anthracis exhibits a Kd that matches that of yitJ (Fig. 6b), implying that the B. subtilis cysH aptamer has suffered one or more mutations that have somewhat degraded binding affinity. This loss of binding affinity would explain why the gene is not modulated in vivo despite the fact that high concentrations of SAM induce transcription termination in vitro.
(a) Structural modulation of the B. subtilis cysH aptamer as determined by in-line probing. Inset: apparent Kd values determined by monitoring structural modulation over a range of SAM or SAM analog concentrations. Two guanosyl residues (asterisks) were included at the 5′ terminus of the RNA construct to facilitate in vitro transcription. Nucleotide numbers are given relative to the putative transcription start site. In-line probing was conducted with an RNA extending to nucleotide 117, whereas the remainder of the RNA is shown to depict the putative transcription terminator stem. Experiments were similar to those described in Figures 1b (see legend for details) and 2b. (b) Structural modulation of the B. anthracis cysH aptamer as determined by in-line probing. The transcription start point of the B. anthracis cysH mRNA has not been determined, and so numbering of nucleotides begins immediately after the two inserted guanosyl residues (asterisks). In-line probing was conducted with an RNA extending to nucleotide 112. See a for additional details.
Discussion
B. subtilis and several other Gram-positive bacteria carry all known classes of riboswitches, including newly discovered riboswitches that respond to guanine24 and lysine (N.S., J.K. Wickiser, S. Nakamura, M.S. Ebert and R.R.B., unpublished data. Current biochemical and bioinformatics data indicate that B. subtilis has at least 68 genes (nearly 2% of its total genetic complement) under riboswitch control24. Moreover, each of these mRNAs is responding to biological compounds that are universal in biology. The fact that genetic control elements for fundamental metabolic processes are formed by RNA indicates that this polymer has the structural sophistication needed to precisely monitor chemical environments and transduce metabolite-binding events into genetic responses. A more detailed analysis of riboswitch structures at the atomic level would be of great utility in determining how metabolite binding promotes allosteric reorganization of RNA genetic switches.
Riboswitches ligands such as SAM and guanine seem to serve as master control molecules whose concentrations are being monitored to ensure homeostasis of a much wider set of metabolic pathways. Riboswitches also seem to permit metabolite surveillance and genetic control with the same level of precision and efficiency as that exhibited by protein factors, and thus could have emerged late in the evolution of modern biochemical architectures. However, given their fundamental role in metabolic maintenance and the widespread phylogenetic distribution of certain riboswitches, we speculate that aptamer domains similar to these might have been the primary mechanism by which RNA-world organisms detected metabolites and controlled biochemical pathways before the emergence of proteins.
Methods
DNA oligonucleotides and chemicals.
Synthetic DNAs were purchased from the HHMI Keck Foundation Biotechnology Resource Center at Yale University (New Haven, Connecticut, USA). Preparation of RNAs by in vitro transcription was carried out25 and the products were purified as described previously3. SAM, various analogs of SAM and S-adenosyl-L-methionine-methyl-3H ([3H]SAM) were purchased from Sigma.
DNA constructs.
A yitJ DNA construct encompassing nucleotides −380 to +14 relative to the translation start site was prepared using primers that generated EcoRI and BamH1 restriction sites upon PCR amplification of B. subtilis chromosomal DNA (strain 168). The product was cloned into pDG1661 (ref. 26; Bacillus Genetic Stock Center, Columbus, Ohio, USA) using these restriction sites to place the riboswitch immediately upstream of the lacZ reporter gene. Mutants were created by using the appropriate mutagenic primers and the QuikChange site-directed mutagenesis kit (Stratagene). All sequences were confirmed by DNA sequencing.
In-line probing assays.
For each probing reaction, ∼1 nM 5′-32P-labeled RNA was incubated for ∼40 h at 25 °C in 50 mM Tris-HCl (pH 8.3 at 25 °C), 20 mM MgCl2, 100 mM KCl, with added compounds as indicated for each experiment. Additional experimental details are similar to those described elsewhere1,2,3. The Kd of 251 yitJ for SAM or for other analogs was determined by plotting the normalized fraction of RNA cleaved at regions 1–6 (see Fig. 1c) versus the logarithm of the concentration of SAM in molar units. The dashed line indicates the concentration needed to induce half maximal modulation of cleavage activity.
Equilibrium dialysis and Scatchard analyses.
Assays used 100 nM of [3H]SAM (14.5 μCi mmol−1, ∼7,000 counts per minute (c.p.m.)) added to side A of an equilibrium dialysis chamber1,2, and were done in the absence (none) or the presence of 3 μM RNA on the B side of the chamber as indicated. Equilibrations were carried out for ∼10 h in the absence (−) of unlabeled analogs, and then were subsequently incubated in the presence of 25 μM unlabeled compounds (added to side B) as indicated. Additional experimental details are as described1,2.
Scatchard data was generated by adding RNA (124 yitJ) to 30 μl in line probing buffer to a final concentration of 200 nM in chamber B of DispoEquilibrium Dialyzers (ED-1, Harvard Bioscience; 5,000 Da molecular-mass cut-off membrane). [3H]SAM (14.5 μCi mmol−1, ∼7,000 c.p.m.) was added to chamber A to concentrations ranging from 10 nM to 2.5 μM (total concentration of SAM = [LT]). The solutions were equilibrated for 11 h at room temperature, whereupon aliquots were removed for scintillation counting. The ratio of c.p.m. of chamber B relative to chamber A was utilized for determination of the concentrations of free ([LF]) and bound ([LB]) [3H]SAM, using the relationships [LT] = [LB] + [LF] and c.p.m. A / c.p.m. B = [LT] / [LF]. The data from four independent analyses were then assembled into a standard Scatchard plot of r ([LB] / (200 nM 124 yitJ)) versus r / [LF], where the x-intercept reflects the number of ligand-binding sites and the slope is equal to −1 / Kd. The data depicted in Figure 3 were derived in four separate experiments and were plotted after correction for ∼20% tritium that is not in the form of SAM.
Expectations for the action of SAM analogs in competition assays (Fig. 2b) were established by adapting the formula used to assess competitive inhibition with enzymes27. Analogs that function as competitors will effectively raise the apparent Kd of SAM according to the following formula: apparent Kd for SAM = measured Kd of SAM × (1 + [analog] / Kd of the analog). For example, when present at 25 μM under our assay conditions, both SAH and SAC are not expected to degrade the apparent Kd for SAM such that a loss of tritium shift would occur.
In vivo analysis of riboswitch function.
Mutations M1–M9 were generated in plasmids containing fusions of the yitJ 5′ UTR upstream from a lacZ reporter gene. Templates for preparation of mutant RNAs for in vitro studies were then created by PCR, and the mutant DNA constructs were integrated into the bacterial chromosome for in vivo studies. Transformations of pDG1661 variants (see DNA constructs) into B. subtilis strain 1A234 (obtained from the Bacillus Genetic Stock Center) were carried out as described28. The correct transformants were identified by selecting for chloramphenicol (5 μg ml−1) resistance and screening for spectinomycin (100 μg ml−1) sensitivity. Proper site-specific genomic insertion by double crossover recombination was confirmed by PCR using amyE-specific primers.
Cells were grown with shaking at 37 °C either in rich media (2XYT broth or tryptose blood agar base) or defined media (0.5% (w/v) glucose, 2 g l−1 (NH4)2SO4, 18.3 g l−1 K2HPO4·3H2O, 6 g l−1 KH2PO4, 1 g l−1 sodium citrate, 0.2 g l−1 MgSO4·7H2O, 5 μM MnCl2, 0.5 mM CaCl2, 50 μg tryptophan and 50 μg ml−1 glutamate. Methionine was added to 50 μg ml−1 for routine growth. Growth under methionine-limiting conditions was established by incubation under routine growth conditions in defined medium to an A595 of 0.1, at which time the cells were pelleted by centrifugation, resuspended in minimal media, split into two aliquots and supplemented with either 50 μg ml−1 (with methionine) or 0.25 μg ml−1 (without methionine) (Fig. 4c). Cultures were incubated for an additional 3 h before β-galactosidase assays were carried out.
In vitro transcription termination assays.
Transcription reactions (10 μl) containing ∼30 pmol of specific template DNA, 200 μM each NTP, 5 μCi [α-32P]UTP (1 Ci = 37 GBq) and 50 units of T7 RNA polymerase (New England Biolabs) were incubated in the presence of 50 mM Tris-HCl (pH 7.5 at 23 °C), 15 mM MgCl2, 2 mM spermidine, 5 mM DTT at 37 °C for 2 h. SAM and its analogs were added to a final concentration of 50 μM. Transcription templates were generated for all 11 riboswitch domains in the S-box regulon of B. subtilis by using PCR with corresponding primers that in each case produced transcripts beginning with GG, encompassing the putative natural transcription start14, and including the first 13 codons of the adjoining open reading frame. Transcription products were separated by denaturing 6% PAGE and visualized by PhosphorImager. Termination yields were approximated by determining the ratio of RNAs in the termination band relative to the combined terminated and full-length RNAs.
Note: Supplementary information is available on the Nature Structural Biology website.
References
Nahvi, A. et al. Genetic control by a metabolite binding mRNA. Chem. Biol. 9, 1043–1049 (2002).
Winkler, W., Nahvi, A. & Breaker, R.R. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature 419, 952–956 (2002).
Winkler, W.C., Cohen-Chalamish, S. & Breaker, R.R. An mRNA structure that controls gene expression by binding FMN. Proc. Natl. Acad. Sci. USA 99, 15908–15913 (2002).
Mironov, A.S. et al. Sensing small molecules by nascent RNA: a mechanism to control transcription in bacteria. Cell 111, 747–756 (2002).
Sudarsan, N., Barrick, J.E. & Breaker, R.R. Metabolite-binding RNA domains are present in the genes of eukaryotes. RNA 644–647 (2003).
Gelfand, M.S., Mironov, A.A., Jomantas, J., Kozlov, Y.I. & Perumov, D.A. A conserved RNA structure element involved in the regulation of bacterial riboflavin synthesis genes. Trends Genet. 15, 439–442 (1999).
Miranda-Rios, J., Navarro, M. & Soberón, M. A conserved RNA structure (thi box) is involved in regulation of thiamin biosynthetic gene expression in bacteria. Proc. Natl. Acad. Sci. USA 98, 9736–9741 (2001).
Stormo, G.D. & Ji, Y. Do mRNAs act as direct sensors of small molecules to control their expression? Proc. Natl. Acad. Sci. USA 98, 9465–9467 (2001).
Rodionov, D.A., Vitreschak, A.G., Mironov, A.A. & Gelfand, M.S. Comparative genomics of thiamin biosynthesis in procaryotes. New genes and regulatory mechanisms. J. Biol. Chem. 277, 48949–48959 (2002).
Benner, S.A., Ellington, A.D. & Tauer, A. Modern metabolism as a palimpsest of the RNA world. Proc. Natl. Acad. Sci. USA 86, 7054–7058 (1989).
Joyce, G.F. The antiquity of RNA-based evolution. Nature 418, 214–221 (2002).
White III, H.B. Coenzymes as fossils of an earlier metabolic state. J. Mol. Evol. 7, 101–104 (1976).
Jeffares, D.C., Poole, A.M. & Penny, D. Relics from the RNA world. J. Mol. Evol. 46, 18–36 (1998).
Grundy, F.J. & Henkin, T.M. The S box regulon: a new global transcription termination control system for methionine and cysteine biosynthesis genes in Gram-positive bacteria. Mol. Microbiol. 30, 737–749 (1998).
Henkin, T.M. Transcription termination control in bacteria. Curr. Opin. Microbiol. 3, 149–153 (2000).
Grundy, F.J. & Henkin, T.M. The T box and S box transcription termination control systems. Frontiers Biosci. 8, D20–31 (2003).
Soukup, G.A. & Breaker, R.R. Relationship between internucleotide linkage geometry and the stability of RNA. RNA 5, 1308–1325 (1999).
Gold, L., Poliski, B., Uhlenbeck, O.C. & Yarus, M. Diversity of oligonucleotide functions. Annu. Rev. Biochem. 64, 763–797 (1995).
Takusagawa, F., Fujioka, M., Spies, A. & Schowen, R.L. In Comprehensive Biological Catalysis Vol. 1 (ed. Sinnott, M.) 1–30 (Academic, New York, 1998).
McDaniel, B.A.M., Grundy, F.J., Artsimovitch, I. & Henkin, T.M. Transcription termination control of the S box system: direct measurement of S-adenosylmethionine by the leader RNA. Proc. Natl. Acad. Sci. USA 100, 3083–3088 (2003).
Posnick, L.M. & Samson, L.D. Influence of S-adenosylmethionine pool size on spontaneous mutation, dam methylation, and cell growth of Escherichia coli. J. Bacteriol. 181, 6756–6762 (1999).
Epshtein, V., Mironov, A.S. & Nudler, E. The riboswitch-mediated control of sulfur metabolism in bacteria. Proc. Natl. Acad. Sci. USA 100, 5052–5056 (2003).
Mansilla, M.C., Albanesi, D. & De Mendoza, D. Transcriptional control of the sulfur-regulated cysH operon, containing genes involved in L-cysteine biosynthesis in Bacillus subtilis. J. Bacteriol. 182, 5885–5892 (2000).
Mandal, M., Boese, B., Barrick, J.E., Winkler, W.C. & Breaker, R.R. Metabolite-sensing riboswitches control fundamental biochemical pathways in bacteria. Cell 113, 577–586 (2003).
Seetharaman, S., Zivartz, M., Sudarsan, N. & Breaker, R.R. Immobilized RNA switches for the analysis of complex chemical and biological mixtures. Nat. Biotechnol. 19, 336–341 (2001).
Guérout-Fleury, A.M., Frandsen, N. & Stragier, P. Plasmids for ectopic integration in Bacillus subtilis. Gene 180, 57–61 (1996).
Fersht, A. In Structure and Mechanism in Protein Science 113 (Freeman, New York, 1999).
Jarmer, H., Berka, R., Knudsen, S. & Saxild, H.H. Transcriptome analysis documents induced competence of Bacillus subtilis during nitrogen limiting conditions. FEMS Microbiol. Lett. 206, 197–200 (2002).
Acknowledgements
We thank J.K. Wickiser for assistance with the Scatchard analysis, other members of the Breaker laboratory for helpful discussions, and D. Söll at Yale University for the generous gift of B. subtilis strain 168. We also thank A. Miranker and B. Koo for assistance in confirming the purity of SAM using mass spectroscopy. This work was supported by grants from the US National Institutes of Health and the US National Science Foundation. R.R.B is also grateful for support from the David and Lucile Packard Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Winkler, W., Nahvi, A., Sudarsan, N. et al. An mRNA structure that controls gene expression by binding S-adenosylmethionine. Nat Struct Mol Biol 10, 701–707 (2003). https://doi.org/10.1038/nsb967
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb967
This article is cited by
-
Magnesium ions mediate ligand binding and conformational transition of the SAM/SAH riboswitch
Communications Biology (2023)
-
Observation of structural switch in nascent SAM-VI riboswitch during transcription at single-nucleotide and single-molecule resolution
Nature Communications (2023)
-
Structure-based insights into recognition and regulation of SAM-sensing riboswitches
Science China Life Sciences (2023)
-
Getting to the bottom of lncRNA mechanism: structure–function relationships
Mammalian Genome (2022)
-
Identification of 11 candidate structured noncoding RNA motifs in humans by comparative genomics
BMC Genomics (2021)