Abstract
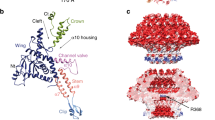
The baseplate of bacteriophage T4 is a multiprotein molecular machine that controls host cell recognition, attachment, tail sheath contraction and viral DNA ejection. We report here the three-dimensional structure of the baseplate–tail tube complex determined to a resolution of 12 Å by cryoelectron microscopy. The baseplate has a six-fold symmetric, dome-like structure ∼520 Å in diameter and ∼270 Å long, assembled around a central hub. A 940 Å–long and 96 Å–diameter tail tube, coaxial with the hub, is connected to the top of the baseplate. At the center of the dome is a needle-like structure that was previously identified as a cell puncturing device. We have identified the locations of six proteins with known atomic structures, and established the position and shape of several other baseplate proteins. The baseplate structure suggests a mechanism of baseplate triggering and structural transition during the initial stages of T4 infection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Eiserling, F.A. & Black, L.W. Pathways in T4 morphogenesis. In Molecular Biology of Bacteriophage T4 (ed. Karam, J.D.) 209–212 (American Society for Microbiology, Washington, DC, 1994).
Kikuchi, Y. & King, J. Genetic control of bacteriophage T4 baseplate morphogenesis. III. Formation of the central plug and overall assembly pathway. J. Mol. Biol. 99, 695–716 (1975).
Crawford, J. & Goldberg, E. The function of tail fibers in triggering baseplate expansion of bacteriophage T4. J. Mol. Biol. 139, 679–690 (1980).
Crowther, R.A., Lenk, E.V., Kikuchi, Y. & King, J. Molecular reorganization in the hexagon to star transition of the baseplate of bacteriophage T4. J. Mol. Biol. 116, 489–523 (1977).
Kanamaru, S. et al. The structure of the bacteriophage T4 cell-puncturing device. Nature 415, 553–557 (2002).
Coombs, D.H. & Arisaka, F. T4 tail structure and function. In Molecular Biology of Bacteriophage T4 (ed. Karam, J.D.) 259–281 (American Society for Microbiology, Washington, DC, 1994).
Watts, N.R.M. & Coombs, D.H. Structure of the bacteriophage T4 baseplate as determined by chemical cross-linking. J. Virol. 64, 143–154 (1990).
Watts, N.R.M., Hainfeld, J. & Coombs, D.H. Localization of the proteins gp7, gp8 and gp10 in the bacteriophage T4 baseplate with colloidal gold:F(ab)2 and undecagold:Fab' conjugates. J. Mol. Biol. 216, 315–325 (1990).
Makhov, A.M. et al. The short tail-fiber of bacteriophage T4: molecular structure and a mechanism for its conformational transition. Virology 194, 117–127 (1993).
Veprintseva, O.D., Deev, A.A., Ivanitski, G.R., Kuninski, A.S. & Khusainov, A.A. Issledovanie aberrantno formy reorganizatsii faga T4 (aberrant form of T4 phage reorganization). Dokl. Akad. Nauk SSSR 254, 496–499 (1980).
Kostyuchenko, V.A. et al. The structure of bacteriophage T4 gene product 9: the trigger for tail contraction. Structure 7, 1213–1222 (1999).
Leiman, P.G. et al. Structure of bacteriophage T4 gene product 11, the interface between the baseplate and short tail fibers. J. Mol. Biol. 301, 975–985 (2000).
van Raaij, M.J., Schoehn, G., Burda, M.R. & Miller, S. Crystal structure of a heat and protease-stable part of the bacteriophage T4 short tail fibre. J. Mol. Biol. 314, 1137–1146 (2001).
Leiman, P.G. et al. Structure and location of gene product 8 in the bacteriophage T4 baseplate. J. Mol. Biol. 328, 821–833 (2003).
Plishker, M.F. & Berget, P.B. Isolation and characterization of precursors in bacteriophage T4 baseplate assembly. III. The carboxyl termini of protein P11 are required for assembly activity. J. Mol. Biol. 178, 699–709 (1984).
Plishker, M.F., Rangwala, S.H. & Berget, P.B. Isolation of bacteriophage T4 baseplate proteins P7 and P8 and in vitro formation of the P10/P7/P8 assembly intermediate. J. Virol. 62, 400–406 (1988).
Zhao, L., Takeda, S., Inoue, M., Leiman, P.G. & Arisaka, F. Stoichiometry and inter-subunit interaction of the wedge initiation complex, gp10-gp11, of bacteriophage T4. Biochim. Biophys. Acta 1479, 286–292 (2000).
Kanamaru, S., Gassner, N.C., Ye, N., Takeda, S. & Arisaka, F. The C-terminal fragment of the precursor tail lysozyme of bacteriophage T4 stays as a structural component of the baseplate after cleavage. J. Bacteriol. 181, 2739–2744 (1999).
Thomassen, E. et al. The structure of the receptor-binding domain of the bacteriophage T4 short tail fibre reveals a knitted trimeric metal-binding fold. J. Mol. Biol. 331, 361–373 (2003).
Meezen, E. & Wood, W.B. The sequence of gene product interaction in bacteriophage T4 core assembly. J. Mol. Biol. 58, 685–692 (1971).
Duda, R.L., Gingery, M. & Eiserling, F.A. Potential length determiner and DNA injection protein is extruded from bacteriophage T4 tail tubes in vitro. Virology 151, 296–314 (1986).
Abuladze, N.K., Gingery, M., Tsai, J. & Eiserling, F.A. Tail length determination in bacteriophage T4. Virology 199, 301–310 (1994).
Moody, M.F. & Makowski, L. X-ray diffraction study of tail-tubes from bacteriophage T2L. J. Mol. Biol. 150, 217–244 (1981).
Crowther, R.A. Mutants of bacteriophage T4 that produce infective fibreless particles. J. Mol. Biol. 137, 159–174 (1980).
Kellenberger, E. et al. Functions and properties related to the tail fibers of bacteriophage T4. Virology 26, 419–440 (1965).
Kellenberger, E., Stauffer, E., Haner, M., Lustig, A. & Karamata, D. Mechanism of the long tail-fiber deployment of bacteriophages T-even and its role in adsorption, infection and sedimentation. Biophys. Chem. 59, 41–59 (1996).
Moody, M.F. Sheath of bacteriophage T4. III. Contraction mechanism deduced from partially contracted sheaths. J. Mol. Biol. 80, 613–635 (1973).
King, J. & Mykolajewycz, N. Bacteriophage T4 tail assembly: proteins of the sheath, core, and baseplate. J. Mol. Biol. 75, 339–358 (1973).
Baker, T.S., Olson, N.H. & Fuller, S.D. Adding the third dimension to virus life cycles: three-dimensional reconstruction of icosahedral viruses from cryo-electron micrographs. Microbiol. Mol. Biol. Rev. 63, 862–922 (1999).
Conway, J.F. & Steven, A.C. Methods for reconstructing density maps of “single” particles from cryoelectron micrographs to subnanometer resolution. J. Struct. Biol. 128, 106–118 (1999).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Wriggers, W. & Chacón, P. Modeling tricks and fitting techniques for multiresolution structures. Structure 9, 779–788 (2001).
Rossmann, M.G., Bernal, R. & Pletnev, S.V. Combining electron microscopic with X-ray crystallographic structures. J. Struct. Biol. 136, 190–200 (2001).
Acknowledgements
We are grateful for a Keck Foundation Grant for the purchase of the CM300 FEG electron microscope. We thank S. Wilder and C. Towell for help in the preparation of the manuscript. The work was supported by a US National Science Foundation grant to M.G.R., a Howard Hughes Medical Institute grant and Russian Fund for Basic Research grant to V.V.M. and a Russian Fund for Basic Research grant to V.A.K. This work is an initiation of a T4 structural genomics project supported by a Human Frontiers Science Program grant to M.G.R., F.A. and V.V.M.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Kostyuchenko, V., Leiman, P., Chipman, P. et al. Three-dimensional structure of bacteriophage T4 baseplate. Nat Struct Mol Biol 10, 688–693 (2003). https://doi.org/10.1038/nsb970
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb970
This article is cited by
-
Biogenesis and structure of a type VI secretion baseplate
Nature Microbiology (2018)
-
Recent Advancements in 3-D Structure Determination of Bacteriophages: from Negative Stain to CryoEM
Journal of the Indian Institute of Science (2018)
-
Artificial bio-nanomachines based on protein needles derived from bacteriophage T4
Biophysical Reviews (2018)
-
Most of it started with T4 phage and was then taken over
Biophysical Reviews (2018)
-
Foreword to ‘Multiscale structural biology: biophysical principles and mechanisms underlying the action of bio-nanomachines’, a special issue in Honour of Fumio Arisaka’s 70th birthday
Biophysical Reviews (2018)