Abstract
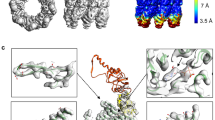
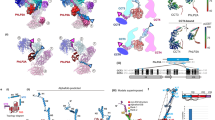
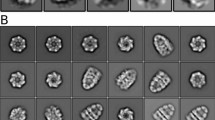
All chaperonins mediate ATP-dependent polypeptide folding by confining substrates within a central chamber. Intriguingly, the eukaryotic chaperonin TRiC (also called CCT) uses a built-in lid to close the chamber, whereas prokaryotic chaperonins use a detachable lid. Here we determine the mechanism of lid closure in TRiC using single-particle cryo-EM and comparative protein modeling. Comparison of TRiC in its open, nucleotide-free, and closed, nucleotide-induced states reveals that the interdomain motions leading to lid closure in TRiC are radically different from those of prokaryotic chaperonins, despite their overall structural similarity. We propose that domain movements in TRiC are coordinated through unique interdomain contacts within each subunit and, further, these contacts are absent in prokaryotic chaperonins. Our findings show how different mechanical switches can evolve from a common structural framework through modification of allosteric networks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Dobson, C.M. Principles of protein folding, misfolding and aggregation. Semin. Cell Dev. Biol. 15, 3–16 (2004).
Gregersen, N., Bross, P., Jorgensen, M.M., Corydon, T.J. & Andresen, B.S. Defective folding and rapid degradation of mutant proteins is a common disease mechanism in genetic disorders. J. Inherit. Metab. Dis. 23, 441–447 (2000).
Frydman, J. Folding of newly translated proteins in vivo: the role of molecular chaperones. Annu. Rev. Biochem. 70, 603–647 (2001).
Hartl, F.U. & Hayer-Hartl, M. Molecular chaperones in the cytosol: from nascent chain to folded protein. Science 295, 1852–1858 (2002).
Sigler, P.B. et al. Structure and function in GroEL-mediated protein folding. Annu. Rev. Biochem. 67, 581–608 (1998).
Spiess, C., Meyer, A.S., Reissmann, S. & Frydman, J. Mechanism of the eukaryotic chaperonin: protein folding in the chamber of secrets. Trends Cell Biol. 14, 598–604 (2004).
Frydman, J. et al. Function in protein folding of TRiC, a cytosolic ring complex containing TCP-1 and structurally related subunits. EMBO J. 11, 4767–4778 (1992).
Kerner, M.J. et al. Proteome-wide analysis of chaperonin-dependent protein folding in Escherichia coli . Cell 122, 209–220 (2005).
Tian, G., Vainberg, I.E., Tap, W.D., Lewis, S.A. & Cowan, N.J. Specificity in chaperonin-mediated protein folding. Nature 375, 250–253 (1995).
Fenton, W.A. & Horwich, A.L. Chaperonin-mediated protein folding: fate of substrate polypeptide. Q. Rev. Biophys. 36, 229–256 (2003).
Brinker, A. et al. Dual function of protein confinement in chaperonin-assisted protein folding. Cell 107, 223–233 (2001).
Bukau, B. & Horwich, A.L. The Hsp70 and Hsp60 chaperone machines. Cell 92, 351–366 (1998).
Saibil, H.R. & Ranson, N.A. The chaperonin folding machine. Trends Biochem. Sci. 27, 627–632 (2002).
Valpuesta, J.M., Martin-Benito, J., Gomez-Puertas, P., Carrascosa, J.L. & Willison, K.R. Structure and function of a protein folding machine: the eukaryotic cytosolic chaperonin CCT. FEBS Lett. 529, 11–16 (2002).
Gutsche, I., Essen, L.O. & Baumeister, W. Group II chaperonins: new TRiC(k)s and turns of a protein folding machine. J. Mol. Biol. 293, 295–312 (1999).
Leroux, M.R. & Hartl, F.U. Protein folding: versatility of the cytosolic chaperonin TRiC/CCT. Curr. Biol. 10, R260–R264 (2000).
Meyer, A.S. et al. Closing the folding chamber of the eukaryotic chaperonin requires the transition state of ATP hydrolysis. Cell 113, 369–381 (2003).
Kafri, G., Willison, K.R. & Horovitz, A. Nested allosteric interactions in the cytoplasmic chaperonin containing TCP-1. Protein Sci. 10, 445–449 (2001).
Reissmann, S., Parnot, C., Booth, C.R., Chiu, W. & Frydman, J. Essential function of the built-in lid in the allosteric regulation of eukaryotic and archaeal chaperonins. Nat. Struct. Mol. Biol. 14, 432–440 (2007).
Yoshida, T. et al. Archaeal group II chaperonin mediates protein folding in the cis-cavity without a detachable GroES-like co-chaperonin. J. Mol. Biol. 315, 73–85 (2002).
Iizuka, R. et al. Role of the helical protrusion in the conformational change and molecular chaperone activity of the archaeal group II chaperonin. J. Biol. Chem. 279, 18834–18839 (2004).
Rivenzon-Segal, D., Wolf, S.G., Shimon, L., Willison, K.R. & Horovitz, A. Sequential ATP-induced allosteric transitions of the cytoplasmic chaperonin containing TCP-1 revealed by EM analysis. Nat. Struct. Mol. Biol. 12, 233–237 (2005).
Boisvert, D.C., Wang, J., Otwinowski, Z., Horwich, A.L. & Sigler, P.B. The 2.4 crystal structure of the bacterial chaperonin GroEL complexed with ATPγS. Nat. Struct. Biol. 3, 170–177 (1996).
Braig, K. et al. The crystal structure of the bacterial chaperonin GroEL at 2.8. Nature 371, 578–586 (1994).
Ditzel, L. et al. Crystal structure of the thermosome, the archaeal chaperonin and homolog of CCT. Cell 93, 125–138 (1998).
Shomura, Y. et al. Crystal structures of the group II chaperonin from Thermococcus strain KS-1: steric hindrance by the substituted amino acid, and inter-subunit rearrangement between two crystal forms. J. Mol. Biol. 335, 1265–1278 (2004).
Llorca, O. et al. The 'sequential allosteric ring' mechanism in the eukaryotic chaperonin-assisted folding of actin and tubulin. EMBO J. 20, 4065–4075 (2001).
Llorca, O. et al. Eukaryotic chaperonin CCT stabilizes actin and tubulin folding intermediates in open quasi-native conformations. EMBO J. 19, 5971–5979 (2000).
Llorca, O. et al. Eukaryotic type II chaperonin CCT interacts with actin through specific subunits. Nature 402, 693–696 (1999).
Xu, Z., Horwich, A.L. & Sigler, P.B. The crystal structure of the asymmetric GroEL-GroES-(ADP)7 chaperonin complex. Nature 388, 741–750 (1997).
Kubota, H., Hynes, G., Carne, A., Ashworth, A. & Willison, K. Identification of six Tcp-1-related genes encoding divergent subunits of the TCP-1-containing chaperonin. Curr. Biol. 4, 89–99 (1994).
Sali, A. & Blundell, T.L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Liou, A.K. & Willison, K.R. Elucidation of the subunit orientation in CCT (chaperonin containing TCP1) from the subunit composition of CCT micro-complexes. EMBO J. 16, 4311–4316 (1997).
Saibil, H. Molecular chaperones: containers and surfaces for folding, stabilising or unfolding proteins. Curr. Opin. Struct. Biol. 10, 251–258 (2000).
Young, J.C., Agashe, V.R., Siegers, K. & Hartl, F.U. Pathways of chaperone-mediated protein folding in the cytosol. Nat. Rev. Mol. Cell Biol. 5, 781–791 (2004).
Brooks, B. & Karplus, M. Harmonic dynamics of proteins: normal modes and fluctuations in bovine pancreatic trypsin inhibitor. Proc. Natl. Acad. Sci. USA 80, 6571–6575 (1983).
Chacon, P., Tama, F. & Wriggers, W. Mega-Dalton biomolecular motion captured from electron microscopy reconstructions. J. Mol. Biol. 326, 485–492 (2003).
Go, N., Noguti, T. & Nishikawa, T. Dynamics of a small globular protein in terms of low-frequency vibrational modes. Proc. Natl. Acad. Sci. USA 80, 3696–3700 (1983).
Hinsen, K. The molecular modeling toolkit: a new approach to molecular simulations. J. Comput. Chem. 21, 79–85 (2000).
Ma, J. & Karplus, M. The allosteric mechanism of the chaperonin GroEL: a dynamic analysis. Proc. Natl. Acad. Sci. USA 95, 8502–8507 (1998).
Tama, F. & Brooks, C.L., III. Diversity and identity of mechanical properties of icosahedral viral capsids studied with elastic network normal mode analysis. J. Mol. Biol. 345, 299–314 (2005).
Heller, M. et al. NMR studies on the substrate-binding domains of the thermosome: structural plasticity in the protrusion region. J. Mol. Biol. 336, 717–729 (2004).
Gomez-Puertas, P., Martin-Benito, J., Carrascosa, J.L., Willison, K.R. & Valpuesta, J.M. The substrate recognition mechanisms in chaperonins. J. Mol. Recognit. 17, 85–94 (2004).
Pappenberger, G. et al. Crystal structure of the CCTγ apical domain: implications for substrate binding to the eukaryotic cytosolic chaperonin. J. Mol. Biol. 318, 1367–1379 (2002).
Feldman, D.E., Spiess, C., Howard, D.E. & Frydman, J. Tumorigenic mutations in VHL disrupt folding in vivo by interfering with chaperonin binding. Mol. Cell 12, 1213–1224 (2003).
Kubota, S., Kubota, H. & Nagata, K. Cytosolic chaperonin protects folding intermediates of Gβ from aggregation by recognizing hydrophobic β-strands. Proc. Natl. Acad. Sci. USA 103, 8360–8365 (2006).
Rommelaere, H., De Neve, M., Melki, R., Vandekerckhove, J. & Ampe, C. The cytosolic class II chaperonin CCT recognizes delineated hydrophobic sequences in its target proteins. Biochemistry 38, 3246–3257 (1999).
Spiess, C., Miller, E.J., McClellan, A.J. & Frydman, J. Identification of the TRiC/CCT substrate binding sites uncovers the function of subunit diversity in eukaryotic chaperonins. Mol. Cell 24, 25–37 (2006).
Ferreyra, R.G. & Frydman, J. Purification of the cytosolic chaperonin TRiC from bovine testis. Methods Mol. Biol. 140, 153–160 (2000).
Booth, C.R. et al. A 9 Å single particle reconstruction from CCD captured images on a 200 kV electron cryomicroscope. J. Struct. Biol. 147, 116–127 (2004).
Ludtke, S.J., Baldwin, P.R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Brink, J. et al. Experimental verification of conformational variation of human fatty acid synthase as predicted by normal mode analysis. Structure 12, 185–191 (2004).
Ludtke, S.J., Chen, D.H., Song, J.L., Chuang, D.T. & Chiu, W. Seeing GroEL at 6 Å resolution by single particle electron cryomicroscopy. Structure 12, 1129–1136 (2004).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Eramian, D. et al. A composite score for predicting errors in protein structure models. Protein Sci. 15, 1653–1666 (2006).
Topf, M., Baker, M.L., Marti-Renom, M.A., Chiu, W. & Sali, A. Refinement of protein structures by iterative comparative modeling and CryoEM density fitting. J. Mol. Biol. 357, 1655–1668 (2006).
Jiang, W., Baker, M.L., Ludtke, S.J. & Chiu, W. Bridging the information gap: computational tools for intermediate resolution structure interpretation. J. Mol. Biol. 308, 1033–1044 (2001).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
Acknowledgements
The authors would like to thank M.L. Baker for his discussions and assistance quantifying the similarity between cryo-EM reconstructions and high-resolution structures. This research was supported by the US National Institutes of Health (NIH), the NIH Roadmap Initiative for Medical Research, the US National Science Foundation and the Robert Welch Foundation.
Author information
Authors and Affiliations
Contributions
W.C. and J.F. designed and led the project; C.R.B. and A.S.M. carried out the protein preparations and cryo-EM measurements; C.R.B. and S.J.L. carried out the single-particle reconstruction and initial modeling; Y.C. carried out the normal mode analysis and refined the comparative models together with M.T. and A.S; C.R.B., A.S.M., W.C. and J.F. interpreted the data and wrote the manuscript; all authors made contributions to the final manuscript.
Corresponding authors
Supplementary information
Supplementary Figures
Supplementary Figures 1–4 (PDF 11600 kb)
Supplementary Video 1
Monomer transition from open to closed conformation. (MPG 6585 kb)
Supplementary Video 2
Octamer transition from open to closed conformation. (MPG 25537 kb)
Supplementary Video 3
Top view of the lowest frequency non-degenerate mode (mode 1) from normal mode analysis (NMA). (MPG 4955 kb)
Supplementary Video 4
Side view of the lowest frequency non-degenerate mode (mode 1) from NMA. (MPG 5570 kb)
Rights and permissions
About this article
Cite this article
Booth, C., Meyer, A., Cong, Y. et al. Mechanism of lid closure in the eukaryotic chaperonin TRiC/CCT. Nat Struct Mol Biol 15, 746–753 (2008). https://doi.org/10.1038/nsmb.1436
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1436
This article is cited by
-
Pathway and mechanism of tubulin folding mediated by TRiC/CCT along its ATPase cycle revealed using cryo-EM
Communications Biology (2023)
-
Staggered ATP binding mechanism of eukaryotic chaperonin TRiC (CCT) revealed through high-resolution cryo-EM
Nature Structural & Molecular Biology (2016)
-
A human CCT5 gene mutation causing distal neuropathy impairs hexadecamer assembly in an archaeal model
Scientific Reports (2014)
-
Symmetry-free cryo-EM structures of the chaperonin TRiC along its ATPase-driven conformational cycle
The EMBO Journal (2012)
-
Mechanism of nucleotide sensing in group II chaperonins
The EMBO Journal (2012)