Abstract
IpaH proteins are E3 ubiquitin ligases delivered by the type III secretion apparatus into host cells upon infection of humans by the Gram-negative pathogen Shigella flexneri. These proteins comprise a variable leucine-rich repeat–containing N-terminal domain and a conserved C-terminal domain harboring an invariant cysteine residue that is crucial for activity. IpaH homologs are encoded by diverse animal and plant pathogens. Here we demonstrate that the IpaH C-terminal domain carries the catalytic activity for ubiquitin transfer and that the N-terminal domain carries the substrate specificity. The structure of the IpaH C-terminal domain, determined to 2.65-Å resolution, represents an all-helical fold bearing no resemblance to previously defined E3 ubiquitin ligases. The conserved and essential cysteine residue lies on a flexible, surface-exposed loop surrounded by conserved acidic residues, two of which are crucial for IpaH activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Bhavsar, A.P., Guttman, J.A. & Finlay, B.B. Manipulation of host-cell pathways by bacterial pathogens. Nature 449, 827–834 (2007).
Galán, J.E. & Cossart, P. Host-pathogen interactions: a diversity of themes, a variety of molecular machines. Curr. Opin. Microbiol. 8, 1–3 (2005).
Ardley, H.C. & Robinson, P.A. E3 ubiquitin ligases. Essays Biochem. 41, 15–30 (2005).
Mattoo, S., Lee, Y.M. & Dixon, J.E. Interactions of bacterial effector proteins with host proteins. Curr. Opin. Immunol. 19, 392–401 (2007).
Abramovitch, R.B., Janjusevic, R., Stebbins, C.E. & Martin, G.B. Type III effector AvrPtoB requires intrinsic E3 ubiquitin ligase activity to suppress plant cell death and immunity. Proc. Natl. Acad. Sci. USA 103, 2851–2856 (2006).
Rosebrock, T.R. et al. A bacterial E3 ubiquitin ligase targets a host protein kinase to disrupt plant immunity. Nature 448, 370–374 (2007).
Angot, A. et al. Ralstonia solanacearum requires F-box-like domain-containing type III effectors to promote disease on several host plants. Proc. Natl. Acad. Sci. USA 103, 14620–14625 (2006).
Zhang, Y., Higashide, W.M., McCormick, B.A., Chen, J. & Zhou, D. The inflammation-associated Salmonella SopA is a HECT-like E3 ubiquitin ligase. Mol. Microbiol. 62, 786–793 (2006).
Diao, J., Zhang, Y., Huibregtse, J.M., Zhou, D. & Chen, J. Crystal structure of SopA, a Salmonella effector protein mimicking a eukaryotic ubiquitin ligase. Nat. Struct. Mol. Biol. 15, 65–70 (2008).
Kubori, T., Hyakutake, A. & Nagai, H. Legionella translocates an E3 ubiquitin ligase that has multiple U-boxes with distinct functions. Mol. Microbiol. 67, 1307–1319 (2008).
Buchrieser, C. et al. The virulence plasmid pWR100 and the repertoire of proteins secreted by the type III secretion apparatus of Shigella flexneri. Mol. Microbiol. 38, 760–771 (2000).
Ashida, H., Toyotome, T., Nagai, T. & Sasakawa, C. Shigella chromosomal IpaH proteins are secreted via the type III secretion system and act as effectors. Mol. Microbiol. 63, 680–693 (2007).
Rohde, J.R., Breitkreutz, A., Chenal, A., Sansonetti, P.J. & Parsot, C. Type III secretion effectors of the IpaH family are E3 ubiquitin ligases. Cell Host Microbe 1, 77–83 (2007).
Dohlman, H.G., Song, J., Ma, D., Courchesne, W.E. & Thorner, J. Sst2, a negative regulator of pheromone signaling in the yeast Saccharomyces cerevisiae: expression, localization, and genetic interaction and physical association with Gpa1 (the G-protein alpha subunit). Mol. Cell. Biol. 16, 5194–5209 (1996).
Houben, K. et al. Solution structure of the ubiquitin-conjugating enzyme UbcH5B. J. Mol. Biol. 344, 513–526 (2004).
Dominguez, C. et al. Structural model of the UbcH5B/CNOT4 complex revealed by combining NMR, mutagenesis, and docking approaches. Structure 12, 633–644 (2004).
Winkler, G.S. et al. An altered-specificity ubiquitin-conjugating enzyme/ubiquitin-protein ligase pair. J. Mol. Biol. 337, 157–165 (2004).
Eletr, Z.M. & Kuhlman, B. Sequence determinants of E2–E6AP binding affinity and specificity. J. Mol. Biol. 369, 419–428 (2007).
Beaudenon, S., Dastur, A. & Huibregtse, J.M. Expression and assay of HECT domain ligases. Methods Enzymol. 398, 112–125 (2005).
Sun, S.C. Deubiquitylation and regulation of the immune response. Nat. Rev. Immunol. 8, 501–511 (2008).
Wang, M. & Pickart, C.M. Different HECT domain ubiquitin ligases employ distinct mechanisms of polyubiquitin chain synthesis. EMBO J. 24, 4324–4333 (2005).
Wu, P.Y. et al. A conserved catalytic residue in the ubiquitin-conjugating enzyme family. EMBO J. 22, 5241–5250 (2003).
Tolbert, B.S. et al. The active site cysteine of ubiquitin-conjugating enzymes has a significantly elevated pKa: functional implications. Biochemistry 44, 16385–16391 (2005).
Verdecia, M.A. et al. Conformational flexibility underlies ubiquitin ligation mediated by the WWP1 HECT domain E3 ligase. Mol. Cell 11, 249–259 (2003).
Mavris, M. et al. Regulation of transcription by the activity of the Shigella flexneri type III secretion apparatus. Mol. Microbiol. 43, 1543–1553 (2002).
Kim, D.W. et al. The Shigella flexneri effector OspG interferes with innate immune responses by targeting ubiquitin-conjugating enzymes. Proc. Natl. Acad. Sci. USA 102, 14046–14051 (2005).
Breitkreutz, A., Boucher, L. & Tyers, M. MAPK specificity in the yeast pheromone response independent of transcriptional activation. Curr. Biol. 11, 1266–1271 (2001).
Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature 415, 180–183 (2002).
Zhang, R.G. et al. Structure of Thermotoga maritima stationary phase survival protein SurE: a novel acid phosphatase. Structure 9, 1095–1106 (2001).
Kimber, M.S. et al. Data mining crystallization databases: knowledge-based approaches to optimize protein crystal screens. Proteins 51, 562–568 (2003).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution—from diffraction images to an initial model in minutes. Acta Crystallogr. D Biol. Crystallogr. 62, 859–866 (2006).
Schneider, T.R. & Sheldrick, G.M. Substructure solution with SHELXD. Acta Crystallogr. D Biol. Crystallogr. 58, 1772–1779 (2002).
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D Biol. Crystallogr. 59, 2023–2030 (2003).
de la Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997).
Perrakis, A., Morris, R. & Lamzin, V.S. Automated protein model building combined with iterative structure refinement. Nat. Struct. Biol. 6, 458–463 (1999).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Winn, M.D., Isupov, M.N. & Murshudov, G.N. Use of TLS parameters to model anisotropic displacements in macromolecular refinement. Acta Crystallogr. D Biol. Crystallogr. 57, 122–133 (2001).
Winn, M.D., Murshudov, G.N. & Papiz, M.Z. Macromolecular TLS refinement in REFMAC at moderate resolutions. Methods Enzymol. 374, 300–321 (2003).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Acknowledgements
We wish to thank the staff at the Argonne National Laboratory beam line 19-ID for assistance with data collection and A. Edwards for critical reading of the manuscript. We also wish to thank S. Dhe-Paganon and G. Avvakumov at the Structural Genomics Consortium, Toronto, for providing the collection of expression constructs for human E2-conjugating enzymes. We thank D. Briant for insight to E3 ubiquitination assays. This work was supported by US National Institutes of Health Grants GM62414-01, by the Ontario Research and Development Challenge Fund and by a grant from the Canadian Institutes of Health Research Grant. M.T. is supported by grants from the Canadian Institutes of Health Research (MT012466 and MOP-57795), the National Cancer Institute of Canada, the Royal Society and the Scottish Universities Life Sciences Alliance.
Author information
Authors and Affiliations
Contributions
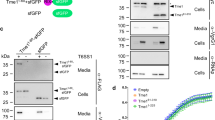
A.U.S. and R.L. determined the structure of the IpaH-CTD and contributed the data for Figures 2, 4 and 6 and Supplementary Figure 1. J.R.R. contributed the data for Figures 1, 2 and 6. T.S. and O.K. purified and crystallized IpaH-CTD. T.S. also contributed the data for Figures 1, 2, 6 and 7. R.D. subcloned most of the constructs. M.E.C. helped with diffraction data collection. A.U.S., J.R.R., N.Y.C., A.J., R.L., M.T., P.J.S., C.P. and A.S. designed experiments, analyzed data and prepared the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1 (PDF 1639 kb)
Rights and permissions
About this article
Cite this article
Singer, A., Rohde, J., Lam, R. et al. Structure of the Shigella T3SS effector IpaH defines a new class of E3 ubiquitin ligases. Nat Struct Mol Biol 15, 1293–1301 (2008). https://doi.org/10.1038/nsmb.1511
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1511
This article is cited by
-
Chromosome-encoded IpaH ubiquitin ligases indicate non-human enteroinvasive Escherichia
Scientific Reports (2022)
-
Subversion of GBP-mediated host defense by E3 ligases acquired during Yersinia pestis evolution
Nature Communications (2022)
-
Efficient production of immunologically active Shigella invasion plasmid antigens IpaB and IpaH using a cell-free expression system
Applied Microbiology and Biotechnology (2022)
-
Substrate-binding destabilizes the hydrophobic cluster to relieve the autoinhibition of bacterial ubiquitin ligase IpaH9.8
Communications Biology (2020)
-
Ubiquitination and degradation of GBPs by a Shigella effector to suppress host defence
Nature (2017)