Abstract
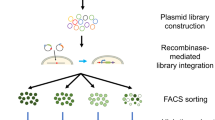
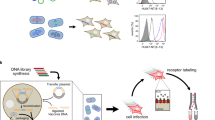
Through the shuffling of predefined modular zinc finger domains with predictable target site recognition in vitro, we have generated a large repertoire of artificial transcription factors with five zinc finger domains (TFZFs). Here we report an effective strategy for the selection of ATF libraries by coupling expression of transcriptional activators of the promoter of interest to the enhanced production of retroviral vector particles transferring the TFZF encoding gene. Using this strategy, we successfully selected specific TFZFs that upregulate the expression of the γ-globin promoter. Selected transcription factors induced the expression of γ-globin when coupled to an activation domain and reduced expression when linked to a repression domain. This new retroviral approach might be used to select other TFZFs but might also be generalized for the selection of other protein and small-molecule interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Beerli, R.R. & Barbas, C.F. III. Engineering polydactyl zinc-finger transcription factors. Nat. Biotechnol. 20, 135–141 (2002).
Dreier, B., Beerli, R.R., Segal, D.J., Flippin, J.D. & Barbas, C.F. III. Development of zinc finger domains for recognition of the 5′-ANN-3′ family of DNA sequences and their use in the construction of artificial transcription factors. J. Biol. Chem. 276, 29466–29478 (2001).
Greisman, H.A. & Pabo, C.O. A general strategy for selecting high-affinity zinc finger proteins for diverse DNA target sites. Science 275, 657–661 (1997).
Liu, P.Q. et al. Regulation of an endogenous locus using a panel of designed zinc finger proteins targeted to accessible chromatin regions. Activation of vascular endothelial growth factor A. J. Biol. Chem. 276, 11323–11334 (2001).
Segal, D.J., Dreier, B., Beerli, R.R. & Barbas, C.F. III. Toward controlling gene expression at will: selection and design of zinc finger domains recognizing each of the 5′-GNN-3′ DNA target sequences. Proc. Natl. Acad. Sci. USA 96, 2758–2763 (1999).
Beerli, R.R., Dreier, B. & Barbas, C.F. III. Positive and negative regulation of endogenous genes by designed transcription factors. Proc. Natl. Acad. Sci. USA 97, 1495–1500 (2000).
Stege, J.T., Guan, X., Ho, T., Beachy, R.N. & Barbas, C.F. III. Controlling gene expression in plants using synthetic zinc finger transcription factors. Plant J. 32, 1077–1086 (2002).
Zhang, L. et al. Synthetic zinc finger transcription factor action at an endogenous chromosomal site. Activation of the human erythropoietin gene. J. Biol. Chem. 275, 33850–33860 (2000).
Blancafort, P., Magnenat, L. & Barbas, C.F. III. Scanning the human genome with combinatorial transcription factor libraries. Nat. Biotechnol. 21, 269–274 (2003).
Magnenat, L., Blancafort, P. & Barbas, C.F. III. In vivo selection of combinatorial libraries and designed affinity maturation of polydactyl zinc finger transcription factors for ICAM-1 provides new insights into gene regulation. J. Mol. Biol. 341, 635–649 (2004).
Chada, K., Magram, J. & Costantini, F. An embryonic pattern of expression of a human fetal globin gene in transgenic mice. Nature 319, 685–689 (1986).
Chada, K. et al. Specific expression of a foreign β-globin gene in erythroid cells of transgenic mice. Nature 314, 377–380 (1985).
Kollias, G., Wrighton, N., Hurst, J. & Grosveld, F. Regulated expression of human Aγ-, β-, and hybrid γβ-globin genes in transgenic mice: manipulation of the developmental expression patterns. Cell 46, 89–94 (1986).
Peterson, K.R. Hemoglobin switching: new insights. Curr. Opin. Hematol. 10, 123–129 (2003).
Rutherford, T. & Nienhuis, A.W. Human globin gene promoter sequences are sufficient for specific expression of a hybrid gene transfected into tissue culture cells. Mol. Cell. Biol. 7, 398–402 (1987).
Li, Q., Peterson, K.R., Fang, X. & Stamatoyannopoulos, G. Locus control regions. Blood 100, 3077–3086 (2002).
Wijgerde, M., Grosveld, F. & Fraser, P. Transcription complex stability and chromatin dynamics in vivo. Nature 377, 209–213 (1995).
Murray, N., Serjeant, B.E. & Serjeant, G.R. Sickle cell-hereditary persistence of fetal haemoglobin and its differentiation from other sickle cell syndromes. Br. J. Haematol. 69, 89–92 (1988).
Swank, R.A. & Stamatoyannopoulos, G. Fetal gene reactivation. Curr. Opin. Genet. Dev. 8, 366–370 (1998).
Blouin, M.J. et al. Genetic correction of sickle cell disease: insights using transgenic mouse models. Nat. Med. 6, 177–182 (2000).
Gräslund, T., Li, X., Magnenat, L., Popkov, M. & Barbas, C.F. III. Exploring strategies for the design of artificial transcription factors: targeting sites proximal to known regulatory regions for the induction of γ-globin expression and the treatment of sickle cell disease. J. Biol. Chem. 280, 3707–3714 (2005).
May, C. et al. Therapeutic haemoglobin synthesis in β-thalassaemic mice expressing lentivirus-encoded human β-globin. Nature 406, 82–86 (2000).
Blau, C.A. et al. γ-Globin gene expression in chemical inducer of dimerization (CID)-dependent multipotential cells established from human β-globin locus yeast artificial chromosome (β-YAC) transgenic mice. J. Biol. Chem. 280, 36642–36647 (2005).
Collis, P., Antoniou, M. & Grosveld, F. Definition of the minimal requirements within the human β-globin gene and the dominant control region for high level expression. EMBO J. 9, 233–240 (1990).
Powars, D.R., Chan, L. & Schroeder, W.A. The influence of fetal hemoglobin on the clinical expression of sickle cell anemia. Ann. NY Acad. Sci. 565, 262–278 (1989).
Rebar, E.J. et al. Induction of angiogenesis in a mouse model using engineered transcription factors. Nat. Med. 8, 1427–1432 (2002).
Maeder, M.L. et al. Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol. Cell 31, 294–301 (2008).
Lund, C.V., Blancafort, P., Popkov, M. & Barbas, C.F. III. Promoter-targeted phage display selections with preassembled synthetic zinc finger libraries for endogenous gene regulation. J. Mol. Biol. 340, 599–613 (2004).
Gordley, R.M., Smith, J.D., Graslund, T. & Barbas, C.F. III. Evolution of programmable zinc finger-recombinases with activity in human cells. J. Mol. Biol. 367, 802–813 (2007).
Miller, J.C. et al. An improved zinc-finger nuclease architecture for highly specific genome editing. Nat. Biotechnol. 25, 778–785 (2007).
Nomura, W. & Barbas, C.F. III. In vivo site-specific DNA methylation with a designed sequence-enabled DNA methylase. J. Am. Chem. Soc. 129, 8676–8677 (2007).
Szczepek, M. et al. Structure-based redesign of the dimerization interface reduces the toxicity of zinc-finger nucleases. Nat. Biotechnol. 25, 786–793 (2007).
Urnov, F.D. et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435, 646–651 (2005).
Xu, G.L. & Bestor, T.H. Cytosine methylation targeted to pre-determined sequences. Nat. Genet. 17, 376–378 (1997).
Mandell, J.G. & Barbas, C.F. III. Zinc Finger Tools: custom DNA-binding domains for transcription factors and nucleases. Nucleic Acids Res. 34, W516–W523 (2006).
Acknowledgements
Thanks to F.C. Costa and R. Neades for technical assistance with the BMC transfections and expression assays. This study was supported by the Skaggs Institute for Chemical Biology and in part by US National Institutes of Health Grants RO1 DK61803 and R01GM065059 (C.F.B.) and R01 DK061804 (K.R.P.).
Author information
Authors and Affiliations
Contributions
U.T., K.R.P. and C.F.B. designed the research; U.T., K.R.P., B.G. and H.F. performed the experiments; U.T., K.R.P. and C.F.B., wrote the manuscript, which all authors commented on.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Table 1 and Supplementary Methods (PDF 408 kb)
Rights and permissions
About this article
Cite this article
Tschulena, U., Peterson, K., Gonzalez, B. et al. Positive selection of DNA-protein interactions in mammalian cells through phenotypic coupling with retrovirus production. Nat Struct Mol Biol 16, 1195–1199 (2009). https://doi.org/10.1038/nsmb.1677
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1677
This article is cited by
-
Synthetic evolution
Nature Biotechnology (2019)
-
Modular system for the construction of zinc-finger libraries and proteins
Nature Protocols (2010)