Abstract
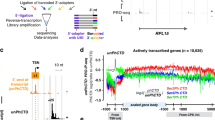
Epigenetic control is an important aspect of gene regulation. Despite detailed understanding of protein-coding gene expression, the transcription of noncoding RNA genes by RNA polymerase III (Pol III) is less well characterized. Here we profile the epigenetic features of Pol III target genes throughout the human genome. This reveals that the chromatin landscape of Pol III–transcribed genes resembles that of Pol II templates in many ways, although there are also clear differences. Our analysis also uncovered an entirely unexpected phenomenon: namely, that Pol II is present at the majority of genomic loci that are bound by Pol III.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Dieci, G. et al. The expanding RNA polymerase III transcriptome. Trends Genet. 23, 614–622 (2007).
Maida, Y. et al. An RNA-dependent RNA polymerase formed by TERT and the RMRP RNA. Nature 461, 230–235 (2009).
Yang, Z. et al. The 7SK small nuclear RNA inhibits the CDK9/cyclin T1 kinase to control transcription. Nature 414, 317–322 (2001).
Nguyen, V.T. et al. 7SK small nuclear RNA binds to and inhibits the activity of CDK9/cyclin T complexes. Nature 414, 322–325 (2001).
Mossink, M.H., van Zon, A., Scheper, R.J., Sonneveld, P. & Wiemer, E.A.C. Vaults: a ribonucleoprotein particle involved in drug resistance? Oncogene 22, 7458–7467 (2003).
Borchert, G.M., Lanier, W. & Davidson, B.L. RNA polymerase III transcribes human microRNAs. Nat. Struct. Mol. Biol. 13, 1097–1101 (2006).
Ozsolak, F. et al. Chromatin structure analyses identify miRNA promoters. Genes Dev. 22, 3172–3183 (2008).
Li, B., Carey, M. & Workman, J.L. The role of chromatin during transcription. Cell 128, 707–719 (2007).
McStay, B. & Grummt, I. The epigenetics of rRNA genes: from molecular to chromosome biology. Annu. Rev. Cell Dev. Biol. 24, 131–157 (2008).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Mikkelsen, T.S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007).
Wang, Z. et al. Combinatorial patterns of histone acetylations and methylations in the human genome. Nat. Genet. 40, 897–903 (2008).
Muse, G.W. et al. RNA polymerase is poised for activation across the genome. Nat. Genet. 39, 1507–1511 (2007).
Zeitlinger, J. et al. RNA polymerase stalling at developmental control genes in the Drosophila melanogaster embryo. Nat. Genet. 39, 1512–1516 (2007).
Bernstein, B.E., Meissner, A. & Lander, E.S. The mammalian epigenome. Cell 128, 669–681 (2007).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Li, J., Moazed, D. & Gygi, S.P. Association of the histone methyltransferase Set2 with RNA polymerase II plays a role in transcription elongation. J. Biol. Chem. 277, 49383–49388 (2002).
Roberts, D.N. & Stewart, A.J. Huff, J. T. & Cairns, B. R. The RNA polymerase III transcriptome revealed by genome-wide localization and activity-occupancy relationships. Proc. Natl. Acad. Sci. USA 100, 14695–14700 (2003).
Kenneth, N.S. et al. TRRAP and GCN5 are used by c-Myc to activate RNA polymerase III transcription. Proc. Natl. Acad. Sci. USA 104, 14917–14922 (2007).
Dittmar, K.A., Goodenbour, J.M. & Pan, T. Tissue-specific differences in human transfer RNA expression. PLoS Genet. 2, e221 (2006).
Bortolin-Cavaille, M.L., Dance, M., Weber, M. & Cavaille, J. C19MC microRNAs are processed from introns of large pol-II, non-protein-coding transcripts. Nucleic Acids Res. 37, 3464–3473 (2009).
Barski, A. et al. Chromatin poises miRNA- and protein-coding genes for expression. Genome Res. 19, 1742–1751 (2009).
Berezikov, E., Cuppen, E. & Plasterk, R.H. Approaches to microRNA discovery. Nat. Genet. 38 (Suppl), S2–S7 (2006).
Jothi, R., Cuddapah, S., Barski, A., Cui, K. & Zhao, K. Genome-wide identification of in vivo protein-DNA binding sites from ChIP-Seq data. Nucleic Acids Res. 36, 5221–5231 (2008).
Listerman, I., Bledau, A.S., Grishina, I. & Neugebauer, K.M. Extragenic accumulation of RNA polymerase II enhances transcription by RNA polymerase III. PLoS Genet. 3, e212 (2007).
Mertens, C. & Roeder, R.G. Different functional modes of p300 in activation of RNA polymerase III transcription from chromatin templates. Mol. Cell. Biol. 28, 5764–5776 (2008).
Lee, D.Y., Hayes, J.J., Pruss, D. & Wolffe, A.P. A positive role for histone acetylation in transcription factor access to nucleosomal DNA. Cell 72, 73–84 (1993).
Ura, K., Kurumizaka, H., Dimitrov, S., Almouzni, G. & Wolffe, A.P. Histone acetylation: influence on transcription, nucleosome mobility and positioning, and linker histone-dependent transcriptional repression. EMBO J. 16, 2096–2107 (1997).
Howe, L., Ranalli, T.A., Allis, C.D. & Ausio, J. Transcriptionally active Xenopus laevis somatic 5 S ribosomal RNA genes are packaged with hyperacetylated histone H4, whereas transcriptionally silent oocyte genes are not. J. Biol. Chem. 273, 20693–20696 (1998).
Tse, C., Sera, T., Wolffe, A.P. & Hansen, J.C. Disruption of higher-order folding by core histone acetylation dramatically enhances transcription of nucleosomal arrays by RNA polymerase III. Mol. Cell. Biol. 18, 4629–4638 (1998).
Kundu, T.K., Wang, Z. & Roeder, R.G. Human TFIIIC relieves chromatin-mediated repression of RNA polymerase III transcription and contains an intrinsic histone acetyltransferase activity. Mol. Cell. Biol. 19, 1605–1615 (1999).
Hernandez, N. TBP, a universal eukaryotic transcription factor? Genes Dev. 7, 1291–1308 (1993).
Gomez-Roman, N., Grandori, C., Eisenman, R.N. & White, R.J. Direct activation of RNA polymerase III transcription by c-Myc. Nature 421, 290–294 (2003).
Ghavi-Helm, Y. et al. Genome-wide location analysis reveals a role of TFIIS in RNA polymerase III transcription. Genes Dev. 22, 1934–1947 (2008).
Lunyak, V.V. Boundaries. Boundaries...Boundaries??? Curr. Opin. Cell Biol. 20, 281–287 (2008).
Haldar, D. & Kamakaka, R.T. tRNA genes as chromatin barriers. Nat. Struct. Mol. Biol. 13, 192–193 (2006).
Donze, D. & Kamakaka, R.T. RNA polymerase III and RNA polymerase II promoter complexes are heterochromatin barriers in Saccharomyces cerevisiae. EMBO J. 20, 520–531 (2001).
Oki, M. & Kamakaka, R.T. Barrier function at HMR. Mol. Cell 19, 707–716 (2005).
Scott, K.C., Merrett, S.L. & Willard, H.F. A heterochromatin barrier partitions the fission yeast centromere into discrete chromatin domains. Curr. Biol. 16, 119–129 (2006).
Noma, K., Cam, H.P., Maraia, R.J. & Grewal, S.I.S. A role for TFIIIC transcription factor complex in genome organization. Cell 125, 859–872 (2006).
Daniels, G.R. & Deininger, P.L. Repeat sequence families derived from mammalian tRNA genes. Nature 317, 819–822 (1985).
Ferrigno, O. et al. Transposable B2 SINE elements can provide mobile RNA polymerase II promoters. Nat. Genet. 28, 77–81 (2001).
West, A.G., Huang, S., Gaszner, M., Litt, M.D. & Felsenfeld, G. Recruitment of histone modifications by USF proteins at a vertebrate barrier element. Mol. Cell 16, 453–463 (2004).
Willoughby, D.A., Vilalta, A. & Oshima, R.G. An Alu element from the K18 gene confers position-independent expression in transgenic mice. J. Biol. Chem. 275, 759–768 (2000).
Geyer, P.K. The role of insulator elements in defining domains of gene expression. Curr. Opin. Genet. Dev. 7, 242–248 (1997).
Chepelev, I., Wei, G., Tang, Q. & Zhao, K. Detection of single nucleotide variations in expressed exons of the human genome using RNA-Seq. Nucleic Acids Res. 37, e106 (2009).
Daly, N.L. et al. Deregulation of RNA polymerase III transcription in cervical epithelium in response to high-risk human papillomavirus. Oncogene 24, 880–888 (2005).
Winter, A.G. et al. RNA polymerase III transcription factor TFIIIC2 is overexpressed in ovarian tumors. Proc. Natl. Acad. Sci. USA 97, 12619–12624 (2000).
Acknowledgements
We thank D.E. Schones for assistance with the Solexa Pipeline analysis and helpful discussions. This work was supported by the Intramural Research Program of the US National Institutes of Health, National Heart, Lung, and Blood Institute (K.Z.) and Cancer Research UK (R.J.W.). A.B. was additionally supported by 1K22HL098691-01 Career Transition Award from the National Heart, Lung, and Blood Institute, US National Institutes of Health.
Author information
Authors and Affiliations
Contributions
A.B., I.C., R.J.W. and K.Z. designed the study and wrote the paper; A.B., D.L., S.C., A.B.F., J.B. and K.C. performed the experiments; I.C. and A.B. analyzed the data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2, Supplementary Tables 1 and 2 and Supplementary Methods (PDF 1099 kb)
Supplementary Table 3
Novel pol III, TFIIIB and TFIIIC binding sites in HeLa cells (XLS 66 kb)
Rights and permissions
About this article
Cite this article
Barski, A., Chepelev, I., Liko, D. et al. Pol II and its associated epigenetic marks are present at Pol III–transcribed noncoding RNA genes. Nat Struct Mol Biol 17, 629–634 (2010). https://doi.org/10.1038/nsmb.1806
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1806
This article is cited by
-
Selective gene expression maintains human tRNA anticodon pools during differentiation
Nature Cell Biology (2024)
-
Chromatin remodeling by Pol II primes efficient Pol III transcription
Nature Communications (2023)
-
A cancer-associated RNA polymerase III identity drives robust transcription and expression of snaR-A noncoding RNA
Nature Communications (2022)
-
Regulatory networking of the three RNA polymerases helps the eukaryotic cells cope with environmental stress
Current Genetics (2021)
-
Yeast PAF1 complex counters the pol III accumulation and replication stress on the tRNA genes
Scientific Reports (2019)