Abstract
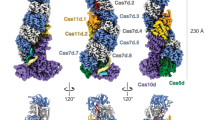
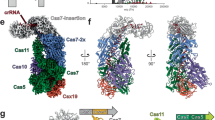
The CRISPR (clustered regularly interspaced short palindromic repeats) immune system in prokaryotes uses small guide RNAs to neutralize invading viruses and plasmids. In Escherichia coli, immunity depends on a ribonucleoprotein complex called Cascade. Here we present the composition and low-resolution structure of Cascade and show how it recognizes double-stranded DNA (dsDNA) targets in a sequence-specific manner. Cascade is a 405-kDa complex comprising five functionally essential CRISPR-associated (Cas) proteins (CasA1B2C6D1E1) and a 61-nucleotide CRISPR RNA (crRNA) with 5′-hydroxyl and 2′,3′-cyclic phosphate termini. The crRNA guides Cascade to dsDNA target sequences by forming base pairs with the complementary DNA strand while displacing the noncomplementary strand to form an R-loop. Cascade recognizes target DNA without consuming ATP, which suggests that continuous invader DNA surveillance takes place without energy investment. The structure of Cascade shows an unusual seahorse shape that undergoes conformational changes when it binds target DNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
van der Oost, J., Jore, M.M., Westra, E.R., Lundgren, M. & Brouns, S.J. CRISPR-based adaptive and heritable immunity in prokaryotes. Trends Biochem. Sci. 34, 401–407 (2009).
Horvath, P. & Barrangou, R. CRISPR/Cas, the immune system of bacteria and archaea. Science 327, 167–170 (2010).
Karginov, F.V. & Hannon, G.J. The CRISPR system: small RNA-guided defense in bacteria and archaea. Mol. Cell 37, 7–19 (2010).
Marraffini, L.A. & Sontheimer, E.J. CRISPR interference: RNA-directed adaptive immunity in bacteria and archaea. Nat. Rev. Genet. 11, 181–190 (2010).
Deveau, H., Garneau, J.E. & Moineau, S. CRISPR/Cas system and its role in phage-bacteria interactions. Annu. Rev. Microbiol. 64, 475–493 (2010).
Mojica, F.J., Diez-Villasenor, C., Garcia-Martinez, J. & Soria, E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J. Mol. Evol. 60, 174–182 (2005).
Pourcel, C., Salvignol, G. & Vergnaud, G. CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiology 151, 653–663 (2005).
Bolotin, A., Quinquis, B., Sorokin, A. & Ehrlich, S.D. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 151, 2551–2561 (2005).
Jansen, R., Embden, J.D., Gaastra, W. & Schouls, L.M. Identification of genes that are associated with DNA repeats in prokaryotes. Mol. Microbiol. 43, 1565–1575 (2002).
Haft, D.H., Selengut, J., Mongodin, E.F. & Nelson, K.E. A guild of 45 CRISPR-associated (Cas) protein families and multiple CRISPR/Cas subtypes exist in prokaryotic genomes. PLOS Comput. Biol. 1, e60 (2005).
Makarova, K.S., Grishin, N.V., Shabalina, S.A., Wolf, Y.I. & Koonin, E.V. A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action. Biol. Direct 1, 7 (2006).
Kunin, V., Sorek, R. & Hugenholtz, P. Evolutionary conservation of sequence and secondary structures in CRISPR repeats. Genome Biol. 8, R61 (2007).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Horvath, P. et al. Diversity, activity, and evolution of CRISPR loci in Streptococcus thermophilus. J. Bacteriol. 190, 1401–1412 (2008).
Garneau, J.E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010).
Wiedenheft, B. et al. Structural basis for DNase activity of a conserved protein implicated in CRISPR-mediated genome defense. Structure 17, 904–912 (2009).
Tang, T.H. et al. Identification of 86 candidates for small non-messenger RNAs from the archaeon Archaeoglobus fulgidus. Proc. Natl. Acad. Sci. USA 99, 7536–7541 (2002).
Hale, C., Kleppe, K., Terns, R.M. & Terns, M.P. Prokaryotic silencing (psi)RNAs in Pyrococcus furiosus. RNA 14, 2572–2579 (2008).
Lillestøl, R.K. et al. CRISPR families of the crenarchaeal genus Sulfolobus: bidirectional transcription and dynamic properties. Mol. Microbiol. 72, 259–272 (2009).
Brouns, S.J. et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321, 960–964 (2008).
Carte, J., Wang, R., Li, H., Terns, R.M. & Terns, M.P. Cas6 is an endoribonuclease that generates guide RNAs for invader defense in prokaryotes. Genes Dev. 22, 3489–3496 (2008).
Haurwitz, R.E., Jinek, M., Wiedenheft, B., Zhou, K. & Doudna, J.A. Sequence- and structure-specific RNA processing by a CRISPR endonuclease. Science 329, 1355–1358 (2010).
Hale, C.R. et al. RNA-guided RNA cleavage by a CRISPR RNA-Cas protein complex. Cell 139, 945–956 (2009).
Marraffini, L.A. & Sontheimer, E.J. CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322, 1843–1845 (2008).
Dickman, M.J. & Hornby, D.P. Enrichment and analysis of RNA centered on ion pair reverse phase methodology. RNA 12, 691–696 (2006).
Waghmare, S.P., Pousinis, P., Hornby, D.P. & Dickman, M.J. Studying the mechanism of RNA separations using RNA chromatography and its application in the analysis of ribosomal RNA and RNA:RNA interactions. J. Chromatogr. A 1216, 1377–1382 (2009).
Raghavan, S.C., Tsai, A., Hsieh, C.L. & Lieber, M.R. Analysis of non-B DNA structure at chromosomal sites in the mammalian genome. Methods Enzymol. 409, 301–316 (2006).
Mojica, F.J., Diez-Villasenor, C., Garcia-Martinez, J. & Almendros, C. Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 155, 733–740 (2009).
Wilson-Sali, T. & Hsieh, T.S. Preferential cleavage of plasmid-based R-loops and D-loops by Drosophila topoisomerase IIIbeta. Proc. Natl. Acad. Sci. USA 99, 7974–7979 (2002).
Thomas, M., White, R.L. & Davis, R.W. Hybridization of RNA to double-stranded DNA: formation of R-loops. Proc. Natl. Acad. Sci. USA 73, 2294–2298 (1976).
Heck, A.J.R. Native mass spectrometry: a bridge between interactomics and structural biology. Nat. Methods 5, 927–933 (2008).
Lorenzen, K., Vannini, A., Cramer, P. & Heck, A.J. Structural biology of RNA polymerase III: mass spectrometry elucidates subcomplex architecture. Structure 15, 1237–1245 (2007).
Zhou, M. et al. Mass spectrometry reveals modularity and a complete subunit interaction map of the eukaryotic translation factor eIF3. Proc. Natl. Acad. Sci. USA 105, 18139–18144 (2008).
Agari, Y., Yokoyama, S., Kuramitsu, S. & Shinkai, A. X-ray crystal structure of a CRISPR-associated protein, Cse2, from Thermus thermophilus HB8. Proteins 73, 1063–1067 (2008).
Ma, J.B. et al. Structural basis for 5′-end-specific recognition of guide RNA by the A. fulgidus Piwi protein. Nature 434, 666–670 (2005).
Li, J., Yang, Z., Yu, B., Liu, J. & Chen, X. Methylation protects miRNAs and siRNAs from a 3′-end uridylation activity in Arabidopsis. Curr. Biol. 15, 1501–1507 (2005).
Houwing, S. et al. A role for Piwi and piRNAs in germ cell maintenance and transposon silencing in Zebrafish. Cell 129, 69–82 (2007).
Marraffini, L.A. & Sontheimer, E.J. Self versus non-self discrimination during CRISPR RNA-directed immunity. Nature 463, 568–571 (2010).
Deveau, H. et al. Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J. Bacteriol. 190, 1390–1400 (2008).
Singleton, M.R., Dillingham, M.S. & Wigley, D.B. Structure and mechanism of helicases and nucleic acid translocases. Annu. Rev. Biochem. 76, 23–50 (2007).
Itoh, T. & Tomizawa, J. Formation of an RNA primer for initiation of replication of ColE1 DNA by ribonuclease H. Proc. Natl. Acad. Sci. USA 77, 2450–2454 (1980).
Baker, T.A. & Kornberg, A. Transcriptional activation of initiation of replication from the E. coli chromosomal origin: an RNA-DNA hybrid near oriC. Cell 55, 113–123 (1988).
Lee, D.Y. & Clayton, D.A. Initiation of mitochondrial DNA replication by transcription and R-loop processing. J. Biol. Chem. 273, 30614–30621 (1998).
Yu, K., Chedin, F., Hsieh, C.L., Wilson, T.E. & Lieber, M.R. R-loops at immunoglobulin class switch regions in the chromosomes of stimulated B cells. Nat. Immunol. 4, 442–451 (2003).
Gnatt, A.L., Cramer, P., Fu, J., Bushnell, D.A. & Kornberg, R.D. Structural basis of transcription: an RNA polymerase II elongation complex at 3.3 A resolution. Science 292, 1876–1882 (2001).
Noy, A., Perez, A., Marquez, M., Luque, F.J. & Orozco, M. Structure, recognition properties, and flexibility of the DNA.RNA hybrid. J. Am. Chem. Soc. 127, 4910–4920 (2005).
Han, D. & Krauss, G. Characterization of the endonuclease SSO2001 from Sulfolobus solfataricus P2. FEBS Lett. 583, 771–776 (2009).
Tahallah, N., Pinkse, M., Maier, C.S. & Heck, A.J. The effect of the source pressure on the abundance of ions of noncovalent protein assemblies in an electrospray ionization orthogonal time-of-flight instrument. Rapid Commun. Mass Spectrom. 15, 596–601 (2001).
van den Heuvel, R.H. et al. Improving the performance of a quadrupole time-of-flight instrument for macromolecular mass spectrometry. Anal. Chem. 78, 7473–7483 (2006).
Maxam, A.M. & Gilbert, W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 65, 499–560 (1980).
Oostergetel, G.T., Keegstra, W. & Brisson, A. Automation of specimen selection and data acquisition for protein electron crystallography. Ultramicroscopy 74, 47–59 (1998).
van Heel, M. et al. Single-particle electron cryo-microscopy: towards atomic resolution. Q. Rev. Biophys. 33, 307–369 (2000).
Acknowledgements
We thank M.H. Lai for experimental contributions, and E. Schaible, P. Zwart and M. Bokhove for technical support and for assistance with post processing of SAXS data. This work was financially supported by The Netherlands Organisation for Scientific Research (NWO) Vici grant to J.v.d.O (865.05.001), Veni grants to S.J.J.B. (863.08.014) and E.v.D. (700.58.402), NWO TOP grant to E.J.B., and UK Engineering and Physical Sciences Research Council and Biotechnology and Biological Sciences Research Council grants to M.J.D. M.L. was financially supported by the Wenner-Gren Foundation, E.R.W. by Spinoza resources awarded to W.M. de Vos, A.P.L.S. by the Research Councils UK, B.W. by the Life Sciences Research Foundation of HHMI and Ü.P. by the Deutsche Forschungsgemeinschaft PU 435/1-1. We thank The Netherlands Proteomics Center for financial support.
Author information
Authors and Affiliations
Contributions
M.M.J., M.L., E.R.W., M.R.B., J.J.B. and J.v.d.O. purified Cascade and performed binding assays and plaque assays. E.v.D., A.B. and A.J.R.H. conducted protein MS experiments. J.B.B. and E.J.B. performed EM. S.P.W., A.P.L.S. and M.J.D. conducted RNA MS experiments. B.W., K.Z. and J.A.D. performed SAXS. Ü.P., R. Wurm and R. Wagner did the footprint analyses. All the authors analyzed the data, and S.J.J.B., M.M.J. and J.v.d.O. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Tables 1–3 and Supplementary Methods (PDF 4696 kb)
Rights and permissions
About this article
Cite this article
Jore, M., Lundgren, M., van Duijn, E. et al. Structural basis for CRISPR RNA-guided DNA recognition by Cascade. Nat Struct Mol Biol 18, 529–536 (2011). https://doi.org/10.1038/nsmb.2019
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2019
This article is cited by
-
Insight into the molecular mechanism of the transposon-encoded type I-F CRISPR-Cas system
Journal of Genetic Engineering and Biotechnology (2023)
-
Insertion sequence transposition inactivates CRISPR-Cas immunity
Nature Communications (2023)
-
The energy landscape for R-loop formation by the CRISPR–Cas Cascade complex
Nature Structural & Molecular Biology (2023)
-
Dynamic interplay between target search and recognition for a Type I CRISPR-Cas system
Nature Communications (2023)
-
Dynamic modulation of enzyme activity by synthetic CRISPR–Cas6 endonucleases
Nature Chemical Biology (2022)