Abstract
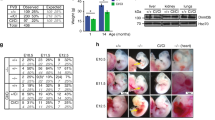
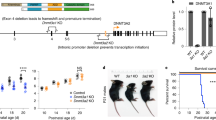
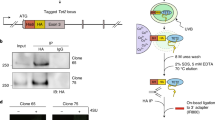
The function of cytosine-C5 methylation, a widespread modification of tRNAs, has remained obscure, particularly in mammals. We have now developed a mouse strain defective in cytosine-C5 tRNA methylation, by disrupting both the Dnmt2 and the NSun2 tRNA methyltransferases. Although the lack of either enzyme alone has no detectable effects on mouse viability, double mutants showed a synthetic lethal interaction, with an underdeveloped phenotype and impaired cellular differentiation. tRNA methylation analysis of the double-knockout mice demonstrated complementary target-site specificities for Dnmt2 and NSun2 and a complete loss of cytosine-C5 tRNA methylation. Steady-state levels of unmethylated tRNAs were substantially reduced, and loss of Dnmt2 and NSun2 was further associated with reduced rates of overall protein synthesis. These results establish a biologically important function for cytosine-C5 tRNA methylation in mammals and suggest that this modification promotes mouse development by supporting protein synthesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Suzuki, M.M. & Bird, A. DNA methylation landscapes: provocative insights from epigenomics. Nat. Rev. Genet. 9, 465–476 (2008).
Mohn, F. & Schubeler, D. Genetics and epigenetics: stability and plasticity during cellular differentiation. Trends Genet. 25, 129–136 (2009).
Jones, P.A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Phizicky, E.M. & Hopper, A.K. tRNA biology charges to the front. Genes Dev. 24, 1832–1860 (2010).
Motorin, Y., Lyko, F. & Helm, M. 5-methylcytosine in RNA: detection, enzymatic formation and biological functions. Nucleic Acids Res. 38, 1415–1430 (2010).
Goll, M.G. et al. Methylation of tRNAAsp by the DNA methyltransferase homolog Dnmt2. Science 311, 395–398 (2006).
Schaefer, M. et al. RNA methylation by Dnmt2 protects transfer RNAs against stress-induced cleavage. Genes Dev. 24, 1590–1595 (2010).
Wilkinson, C.R., Bartlett, R., Nurse, P. & Bird, A.P. The fission yeast gene pmt1+ encodes a DNA methyltransferase homologue. Nucleic Acids Res. 23, 203–210 (1995).
Yoder, J.A. & Bestor, T.H. A candidate mammalian DNA methyltransferase related to pmt1p of fission yeast. Hum. Mol. Genet. 7, 279–284 (1998).
Okano, M., Xie, S. & Li, E. Dnmt2 is not required for de novo and maintenance methylation of viral DNA in embryonic stem cells. Nucleic Acids Res. 26, 2536–2540 (1998).
Jurkowski, T.P. et al. Human DNMT2 methylates tRNAAsp molecules using a DNA methyltransferase-like catalytic mechanism. RNA 14, 1663–1670 (2008).
Sunita, S. et al. Crystal structure of the Escherichia coli 23S rRNA:m5C methyltransferase RlmI (YccW) reveals evolutionary links between RNA modification enzymes. J. Mol. Biol. 383, 652–666 (2008).
Iyer, L.M., Abhiman, S. & Aravind, L. Natural history of eukaryotic DNA methylation systems. Prog. Mol. Biol. Transl. Sci. 101, 25–104 (2011).
Jurkowski, T.P. & Jeltsch, A. On the evolutionary origin of eukaryotic DNA methyltransferases and Dnmt2. PLoS ONE 6, e28104 (2011).
Schaefer, M. & Lyko, F. Solving the Dnmt2 enigma. Chromosoma. 119, 35–40 (2010).
Brzezicha, B. et al. Identification of human tRNA:m5C methyltransferase catalysing intron-dependent m5C formation in the first position of the anticodon of the pre-tRNA Leu (CAA). Nucleic Acids Res. 34, 6034–6043 (2006).
Blanco, S. et al. The RNA–methyltransferase Misu (NSun2) poises epidermal stem cells to differentiate. PLoS Genet. 7, e1002403 (2011).
Motorin, Y. & Grosjean, H. Multisite-specific tRNA:m5C-methyltransferase (Trm4) in yeast Saccharomyces cerevisiae: identification of the gene and substrate specificity of the enzyme. RNA 5, 1105–1118 (1999).
Frye, M. & Watt, F.M. The RNA methyltransferase Misu (NSun2) mediates Myc-induced proliferation and is upregulated in tumors. Curr. Biol. 16, 971–981 (2006).
Squires, J.E. et al. Widespread occurrence of 5-methylcytosine in human coding and non-coding RNA. Nucleic Acids Res. 40, 5023–5033 (2012).
Zhang, X. et al. The tRNA methyltransferase NSun2 stabilizes p16(INK4) mRNA by methylating the 3′-untranslated region of p16. Nat Commun. 3, 712 (2012).
Schaefer, M., Pollex, T., Hanna, K. & Lyko, F. RNA cytosine methylation analysis by bisulfite sequencing. Nucleic Acids Res. 37, e12 (2009).
Sprinzl, M. & Vassilenko, K.S. Compilation of tRNA sequences and sequences of tRNA genes. Nucleic Acids Res. 33, D139–D140 (2005).
Motorin, Y. & Helm, M. RNA nucleotide methylation. Wiley Interdiscip. Rev. RNA 2, 611–631 (2011).
Phizicky, E.M. & Alfonzo, J.D. Do all modifications benefit all tRNAs? FEBS Lett. 584, 265–271 (2010).
Alexandrov, A. et al. Rapid tRNA decay can result from lack of nonessential modifications. Mol. Cell 21, 87–96 (2006).
Chernyakov, I., Whipple, J.M., Kotelawala, L., Grayhack, E.J. & Phizicky, E.M. Degradation of several hypomodified mature tRNA species in Saccharomyces cerevisiae is mediated by Met22 and the 5′-3′ exonucleases Rat1 and Xrn1. Genes Dev. 22, 1369–1380 (2008).
Chan, C.T. et al. A quantitative systems approach reveals dynamic control of tRNA modifications during cellular stress. PLoS Genet. 6, e1001247 (2010).
Shoemaker, C.J., Eyler, D.E. & Green, R. Dom34:Hbs1 promotes subunit dissociation and peptidyl-tRNA drop-off to initiate no-go decay. Science 330, 369–372 (2010).
Ivanov, P., Emara, M.M., Villen, J., Gygi, S.P. & Anderson, P. Angiogenin-induced tRNA fragments inhibit translation initiation. Mol. Cell 43, 613–623 (2011).
Thiagarajan, D., Dev, R.R. & Khosla, S. The DNA methyltranferase Dnmt2 participates in RNA processing during cellular stress. Epigenetics 6, 103–113 (2011).
Chan, C.T. et al. Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins. Nat. Commun. 3, 937 (2012).
Sampath, P. et al. A hierarchical network controls protein translation during murine embryonic stem cell self-renewal and differentiation. Cell Stem Cell 2, 448–460 (2008).
Rai, K. et al. Dnmt2 functions in the cytoplasm to promote liver, brain, and retina development in zebrafish. Genes Dev. 21, 261–266 (2007).
He, C. Grand challenge commentary: RNA epigenetics? Nat. Chem. Biol. 6, 863–865 (2010).
Todaro, G.J. & Green, H. Quantitative studies of the growth of mouse embryo cells in culture and their development into established lines. J. Cell Biol. 17, 299–313 (1963).
Kellner, S., Seidu-Larry, S., Burhenne, J., Motorin, Y. & Helm, M. A multifunctional bioconjugate module for versatile photoaffinity labeling and click chemistry of RNA. Nucleic Acids Res. 39, 7348–7360 (2011).
Acknowledgements
We thank M. Bewerunge-Hudler and O. Heil for mRNA-expression profiling, C. Falckenhayn and G. Raddatz for analyzing 454 sequencing results, J. Hummel-Eisenbeiß and K. Hanna for technical assistance, and J. Burhenne for providing LC-MS. This work was supported by grants from the Deutsche Forschungsgemeinschaft (SPP1463 to F.L., FOR1082 to M.S., M.H. and F.L. and STO 859/2-1 to G.S.). S.H. is supported by a bridging project grant from the DKFZ-ZMBH Alliance, and F.T. is supported by the Institute of Genetics and Biophysics A. Buzzati-Traverso, C.N.R., Italy.
Author information
Authors and Affiliations
Contributions
F.L. conceived the study. F.T. performed the mouse husbandry and the mouse phenotyping. M.F. contributed the NSun2−/− mice. F.T., R.L. and T.M. performed the RNA bisulfite sequencing. S.K. performed the LC-MS analysis. F.T. established the MEF cell lines. F.T., R.L., T.M. and S.H. characterized the MEF cell lines. F.T., M.S., S.H., S.K., M.H., G.S. and F.L. designed the experiments and interpreted the results. F.L. wrote the manuscript with contributions from F.T. and G.S. as well as input from the other coauthors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 1–3 (PDF 228 kb)
Rights and permissions
About this article
Cite this article
Tuorto, F., Liebers, R., Musch, T. et al. RNA cytosine methylation by Dnmt2 and NSun2 promotes tRNA stability and protein synthesis. Nat Struct Mol Biol 19, 900–905 (2012). https://doi.org/10.1038/nsmb.2357
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2357
This article is cited by
-
Roles and regulation of tRNA-derived small RNAs in animals
Nature Reviews Molecular Cell Biology (2024)
-
tRNA epitranscriptomic alterations associated with opioid-induced reward-seeking and long-term opioid withdrawal in male mice
Neuropsychopharmacology (2024)
-
Small RNA modifications: regulatory molecules and potential applications
Journal of Hematology & Oncology (2023)
-
Transcriptome-wide 5-methylcytosine modification profiling of long non-coding RNAs in A549 cells infected with H1N1 influenza A virus
BMC Genomics (2023)
-
Epitranscriptome marks detection and localization of RNA modifying proteins in mammalian ovarian follicles
Journal of Ovarian Research (2023)