Abstract
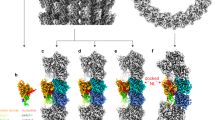
The typical function of kinesins is to transport cargo along microtubules. Binding of ATP to microtubule-attached motile kinesins leads to cargo displacement. To better understand the nature of the conformational changes that lead to the power stroke that moves a kinesin's load along a microtubule, we determined the X-ray structure of human kinesin-1 bound to αβ-tubulin. The structure defines the mechanism of microtubule-stimulated ATP hydrolysis, which releases the kinesin motor domain from microtubules. It also reveals the structural linkages that connect the ATP nucleotide to the kinesin neck linker, a 15–amino acid segment C terminal to the catalytic core of the motor domain, to result in the power stroke. ATP binding to the microtubule-bound kinesin favors neck-linker docking. This biases the attachment of kinesin's second head in the direction of the movement, thus initiating each of the steps taken.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hirokawa, N., Noda, Y., Tanaka, Y. & Niwa, S. Kinesin superfamily motor proteins and intracellular transport. Nat. Rev. Mol. Cell Biol. 10, 682–696 (2009).
Wordeman, L. How kinesin motor proteins drive mitotic spindle function: lessons from molecular assays. Semin. Cell Dev. Biol. 21, 260–268 (2010).
Lawrence, C.J. et al. A standardized kinesin nomenclature. J. Cell Biol. 167, 19–22 (2004).
Howard, J. Mechanics of Motor Proteins and the Cytoskeleton (Sinauer, 2001).
Rice, S. et al. A structural change in the kinesin motor protein that drives motility. Nature 402, 778–784 (1999).
Marx, A., Hoenger, A. & Mandelkow, E. Structures of kinesin motor proteins. Cell Motil. Cytoskeleton 66, 958–966 (2009).
Hirose, K., Akimaru, E., Akiba, T., Endow, S.A. & Amos, L.A. Large conformational changes in a kinesin motor catalyzed by interaction with microtubules. Mol. Cell 23, 913–923 (2006).
Kikkawa, M. & Hirokawa, N. High-resolution cryo-EM maps show the nucleotide binding pocket of KIF1A in open and closed conformations. EMBO J. 25, 4187–4194 (2006).
Sindelar, C.V. & Downing, K.H. An atomic-level mechanism for activation of the kinesin molecular motors. Proc. Natl. Acad. Sci. USA 107, 4111–4116 (2010).
Ma, Y.Z. & Taylor, E.W. Kinetic mechanism of a monomeric kinesin construct. J. Biol. Chem. 272, 717–723 (1997).
Sindelar, C.V. et al. Two conformations in the human kinesin power stroke defined by X-ray crystallography and EPR spectroscopy. Nat. Struct. Biol. 9, 844–848 (2002).
Vale, R.D. & Fletterick, R.J. The design plan of kinesin motors. Annu. Rev. Cell Dev. Biol. 13, 745–777 (1997).
Yang, J.T., Saxton, W.M., Stewart, R.J., Raff, E.C. & Goldstein, L.S. Evidence that the head of kinesin is sufficient for force generation and motility in vitro. Science 249, 42–47 (1990).
Alonso, M.C. et al. An ATP gate controls tubulin binding by the tethered head of kinesin-1. Science 316, 120–123 (2007).
Amos, L.A. & Hirose, K. The structure of microtubule–motor complexes. Curr. Opin. Cell Biol. 9, 4–11 (1997).
Mignot, I. et al. Design and characterization of modular scaffolds for tubulin assembly. J. Biol. Chem. 287, 31085–31094 (2012).
Pecqueur, L. et al. A designed ankyrin repeat protein selected to bind to tubulin caps the microtubule plus end. Proc. Natl. Acad. Sci. USA 109, 12011–12016 (2012).
Nogales, E., Whittaker, M., Milligan, R.A. & Downing, K.H. High-resolution model of the microtubule. Cell 96, 79–88 (1999).
Ravelli, R.B. et al. Insight into tubulin regulation from a complex with colchicine and a stathmin-like domain. Nature 428, 198–202 (2004).
Sack, S., Kull, F.J. & Mandelkow, E. Motor proteins of the kinesin family: structures, variations, and nucleotide binding sites. Eur. J. Biochem. 262, 1–11 (1999).
Grant, B.J., McCammon, J.A., Caves, L.S. & Cross, R.A. Multivariate analysis of conserved sequence-structure relationships in kinesins: coupling of the active site and a tubulin-binding sub-domain. J. Mol. Biol. 368, 1231–1248 (2007).
Chang, Q., Nitta, R., Inoue, S. & Hirokawa, N. Structural basis for the ATP-induced isomerization of kinesin. J. Mol. Biol. 425, 1869–1880 (2013).
Kikkawa, M. et al. Switch-based mechanism of kinesin motors. Nature 411, 439–445 (2001).
Parke, C.L., Wojcik, E.J., Kim, S. & Worthylake, D.K. ATP hydrolysis in Eg5 kinesin involves a catalytic two-water mechanism. J. Biol. Chem. 285, 5859–5867 (2010).
Sack, S. et al. X-ray structure of motor and neck domains from rat brain kinesin. Biochemistry 36, 16155–16165 (1997).
Kull, F.J., Sablin, E.P., Lau, R., Fletterick, R.J. & Vale, R.D. Crystal structure of the kinesin motor domain reveals a structural similarity to myosin. Nature 380, 550–555 (1996).
Leipe, D.D., Wolf, Y.I., Koonin, E.V. & Aravind, L. Classification and evolution of P-loop GTPases and related ATPases. J. Mol. Biol. 317, 41–72 (2002).
Woehlke, G. et al. Microtubule interaction site of the kinesin motor. Cell 90, 207–216 (1997).
Cederquist, G.Y. et al. An inherited TUBB2B mutation alters a kinesin-binding site and causes polymicrogyria, CFEOM and axon dysinnervation. Hum. Mol. Genet. 21, 5484–5499 (2012).
Tischfield, M.A. et al. Human TUBB3 mutations perturb microtubule dynamics, kinesin interactions, and axon guidance. Cell 140, 74–87 (2010).
Cross, R.A. The kinetic mechanism of kinesin. Trends Biochem. Sci. 29, 301–309 (2004).
Hackney, D.D. Kinesin ATPase: rate-limiting ADP release. Proc. Natl. Acad. Sci. USA 85, 6314–6318 (1988).
Coureux, P.D. et al. A structural state of the myosin V motor without bound nucleotide. Nature 425, 419–423 (2003).
Reubold, T.F., Eschenburg, S., Becker, A., Kull, F.J. & Manstein, D.J. A structural model for actin-induced nucleotide release in myosin. Nat. Struct. Biol. 10, 826–830 (2003).
Löwe, J., Li, H., Downing, K.H. & Nogales, E. Refined structure of αβ-tubulin at 3.5 Å resolution. J. Mol. Biol. 313, 1045–1057 (2001).
Pettersen, E.F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Castoldi, M. & Popov, A.V. Purification of brain tubulin through two cycles of polymerization-depolymerization in a high-molarity buffer. Protein Expr. Purif. 32, 83–88 (2003).
Binz, H.K. et al. High-affinity binders selected from designed ankyrin repeat protein libraries. Nat. Biotechnol. 22, 575–582 (2004).
Wong, Y.L. & Rice, S.E. Kinesin's light chains inhibit the head- and microtubule-binding activity of its tail. Proc. Natl. Acad. Sci. USA 107, 11781–11786 (2010).
Wang, W. et al. Kif2C minimal functional domain has unusual nucleotide binding properties that are adapted to microtubule depolymerization. J. Biol. Chem. 287, 15143–15153 (2012).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Strong, M. et al. Toward the structural genomics of complexes: crystal structure of a PE/PPE protein complex from Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 103, 8060–8065 (2006).
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Crystallogr. D Biol. Crystallogr. 66, 133–144 (2010).
Navaza, J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
Nawrotek, A., Knossow, M. & Gigant, B. The determinants that govern microtubule assembly from the atomic structure of GTP-tubulin. J. Mol. Biol. 412, 35–42 (2011).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
We are grateful to J. Skehel and L. Amos for comments on the manuscript. We thank I. Mignot for assistance and D. Mauchand (Unité Commune d'Expérimentation Animale, Institut National de la Recherche Agronomique, Jouy en Josas, France) for providing us with the material from which tubulin was purified. We also thank R. Vale (University of California, San Francisco, San Francisco, California, USA) for giving us the kinesin plasmid. Diffraction data were collected at the European Synchrotron Research Facility (ID29) and at SOLEIL synchrotron (PX1). We thank the machine and beamline groups for making these experiments possible. This work has also benefited from the facilities and expertise of IMAGIF (Centre de Recherche de Gif). We gratefully acknowledge support by the French Fondation pour la Recherche Médicale (DEQ20081213979 to M.K.), Fondation ARC pour la Recherche sur le Cancer (to B.G.) and Agence Nationale de la Recherche (grants ANR-09-BLAN-0071 and ANR-12-BSV8-0002-01 to B.G.) and the Chinese Ministry of Education (grant NCET-10-0604 to C.W.) and Science and Technology Commission of Shanghai Municipality (grant 11JC1413100 to C.W.).
Author information
Authors and Affiliations
Contributions
B.G., C.W. and M.K. designed research; B.G. crystallized the tubulin–DARPin–kinesin complex and determined its structure; W.W. and Q.J. characterized the kinesin-tubulin interaction biochemically; L.P., B.D. and A.P. provided the DARPin; B.G., C.W. and M.K. analyzed the data; B.G., C.W. and M.K. wrote the manuscript with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Table 1 (PDF 1922 kb)
Rights and permissions
About this article
Cite this article
Gigant, B., Wang, W., Dreier, B. et al. Structure of a kinesin–tubulin complex and implications for kinesin motility. Nat Struct Mol Biol 20, 1001–1007 (2013). https://doi.org/10.1038/nsmb.2624
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2624
This article is cited by
-
Anchoring geometry is a significant factor in determining the direction of kinesin-14 motility on microtubules
Scientific Reports (2022)
-
The force required to remove tubulin from the microtubule lattice by pulling on its α-tubulin C-terminal tail
Nature Communications (2022)
-
Magic-angle-spinning NMR structure of the kinesin-1 motor domain assembled with microtubules reveals the elusive neck linker orientation
Nature Communications (2022)
-
Structural basis of mechano-chemical coupling by the mitotic kinesin KIF14
Nature Communications (2021)
-
Tubulin response to intense nanosecond-scale electric field in molecular dynamics simulation
Scientific Reports (2019)