Abstract
Inactivation of cell-survival factors is a crucial step in apoptosis. The phosphoinositide 3-kinase (PI3K)-AKT signaling pathway promotes cell growth, proliferation and survival, and its deregulation causes cancer. How this pathway is suppressed to promote apoptosis is poorly understood. Here we report the identification of a CED-3 caspase substrate in Caenorhabditis elegans, CNT-1, that is cleaved during apoptosis to generate an N-terminal phosphoinositide-binding fragment (tCNT-1). tCNT-1 translocates from the cytoplasm to the plasma membrane and blocks AKT binding to phosphatidylinositol (3,4,5)-trisphosphate, thereby disabling AKT activation and its prosurvival activity. Our findings reveal a new mechanism that negatively regulates AKT cell signaling to promote apoptosis and that may restrict cell growth and proliferation in normal cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Steller, H. Mechanisms and genes of cellular suicide. Science 267, 1445–1449 (1995).
Crawford, E.D. & Wells, J.A. Caspase substrates and cellular remodeling. Annu. Rev. Biochem. 80, 1055–1087 (2011).
Rudel, T. & Bokoch, G.M. Membrane and morphological changes in apoptotic cells regulated by caspase-mediated activation of PAK2. Science 276, 1571–1574 (1997).
Metzstein, M.M., Stanfield, G.M. & Horvitz, H.R. Genetics of programmed cell death in C. elegans: past, present and future. Trends Genet. 14, 410–416 (1998).
Nakagawa, A., Shi, Y., Kage-Nakadai, E., Mitani, S. & Xue, D. Caspase-dependent conversion of Dicer ribonuclease into a death-promoting deoxyribonuclease. Science 328, 327–334 (2010).
Breckenridge, D.G. et al. Caenorhabditis elegans drp-1 and fis-2 regulate distinct cell-death execution pathways downstream of ced-3 and independent of ced-9. Mol. Cell 31, 586–597 (2008).
Chen, Y.Z., Mapes, J., Lee, E.S., Skeen-Gaar, R.R. & Xue, D. Caspase-mediated activation of Caenorhabditis elegans CED-8 promotes apoptosis and phosphatidylserine externalization. Nat. Commun. 4, 2726 (2013).
Cully, M., You, H., Levine, A.J. & Mak, T.W. Beyond PTEN mutations: the PI3K pathway as an integrator of multiple inputs during tumorigenesis. Nat. Rev. Cancer 6, 184–192 (2006).
Luo, J., Manning, B.D. & Cantley, L.C. Targeting the PI3K-Akt pathway in human cancer: rationale and promise. Cancer Cell 4, 257–262 (2003).
Staal, S.P., Hartley, J.W. & Rowe, W.P. Isolation of transforming murine leukemia viruses from mice with a high incidence of spontaneous lymphoma. Proc. Natl. Acad. Sci. USA 74, 3065–3067 (1977).
Li, J. et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275, 1943–1947 (1997).
Vivanco, I. & Sawyers, C.L. The phosphatidylinositol 3-Kinase–AKT pathway in human cancer. Nat. Rev. Cancer 2, 489–501 (2002).
Cho, H. et al. Insulin resistance and a diabetes mellitus-like syndrome in mice lacking the protein kinase Akt2 (PKBβ). Science 292, 1728–1731 (2001).
Hers, I., Vincent, E.E. & Tavare, J.M. Akt signalling in health and disease. Cell. Signal. 23, 1515–1527 (2011).
Finch, C.E. & Ruvkun, G. The genetics of aging. Annu. Rev. Genomics Hum. Genet. 2, 435–462 (2001).
Kenyon, C. The plasticity of aging: insights from long-lived mutants. Cell 120, 449–460 (2005).
Wolff, S. & Dillin, A. The trifecta of aging in Caenorhabditis elegans. Exp. Gerontol. 41, 894–903 (2006).
Kimura, K.D., Tissenbaum, H.A., Liu, Y. & Ruvkun, G. daf-2, an insulin receptor-like gene that regulates longevity and diapause in Caenorhabditis elegans. Science 277, 942–946 (1997).
Morris, J.Z., Tissenbaum, H.A. & Ruvkun, G. A phosphatidylinositol-3-OH kinase family member regulating longevity and diapause in Caenorhabditis elegans. Nature 382, 536–539 (1996).
Wolkow, C.A., Munoz, M.J., Riddle, D.L. & Ruvkun, G. Insulin receptor substrate and p55 orthologous adaptor proteins function in the Caenorhabditis elegans daf-2/insulin-like signaling pathway. J. Biol. Chem. 277, 49591–49597 (2002).
Paradis, S., Ailion, M., Toker, A., Thomas, J.H. & Ruvkun, G.A. PDK1 homolog is necessary and sufficient to transduce AGE-1 PI3 kinase signals that regulate diapause in Caenorhabditis elegans. Genes Dev. 13, 1438–1452 (1999).
Paradis, S. & Ruvkun, G. Caenorhabditis elegans Akt/PKB transduces insulin receptor-like signals from AGE-1 PI3 kinase to the DAF-16 transcription factor. Genes Dev. 12, 2488–2498 (1998).
Hertweck, M., Gobel, C. & Baumeister, R. C. elegans SGK-1 is the critical component in the Akt/PKB kinase complex to control stress response and life span. Dev. Cell 6, 577–588 (2004).
Lin, K., Dorman, J.B., Rodan, A. & Kenyon, C. daf-16: an HNF-3/forkhead family member that can function to double the life-span of Caenorhabditis elegans. Science 278, 1319–1322 (1997).
Ogg, S. et al. The Fork head transcription factor DAF-16 transduces insulin-like metabolic and longevity signals in C. elegans. Nature 389, 994–999 (1997).
Lee, R.Y., Hench, J. & Ruvkun, G. Regulation of C. elegans DAF-16 and its human ortholog FKHRL1 by the daf-2 insulin-like signaling pathway. Curr. Biol. 11, 1950–1957 (2001).
Lin, K., Hsin, H., Libina, N. & Kenyon, C. Regulation of the Caenorhabditis elegans longevity protein DAF-16 by insulin/IGF-1 and germline signaling. Nat. Genet. 28, 139–145 (2001).
Gil, E.B., Malone Link, E., Liu, L.X., Johnson, C.D. & Lees, J.A. Regulation of the insulin-like developmental pathway of Caenorhabditis elegans by a homolog of the PTEN tumor suppressor gene. Proc. Natl. Acad. Sci. USA 96, 2925–2930 (1999).
Mihaylova, V.T., Borland, C.Z., Manjarrez, L., Stern, M.J. & Sun, H. The PTEN tumor suppressor homolog in Caenorhabditis elegans regulates longevity and dauer formation in an insulin receptor-like signaling pathway. Proc. Natl. Acad. Sci. USA 96, 7427–7432 (1999).
Ogg, S. & Ruvkun, G. The C. elegans PTEN homolog, DAF-18, acts in the insulin receptor-like metabolic signaling pathway. Mol. Cell 2, 887–893 (1998).
Rouault, J.P. et al. Regulation of dauer larva development in Caenorhabditis elegans by daf-18, a homologue of the tumour suppressor PTEN. Curr. Biol. 9, 329–332 (1999).
Dorman, J.B., Albinder, B., Shroyer, T. & Kenyon, C. The age-1 and daf-2 genes function in a common pathway to control the lifespan of Caenorhabditis elegans. Genetics 141, 1399–1406 (1995).
Larsen, P.L., Albert, P.S. & Riddle, D.L. Genes that regulate both development and longevity in Caenorhabditis elegans. Genetics 139, 1567–1583 (1995).
Quevedo, C., Kaplan, D.R. & Derry, W.B. AKT-1 regulates DNA-damage-induced germline apoptosis in C. elegans. Curr. Biol. 17, 286–292 (2007).
Parrish, J. et al. Mitochondrial endonuclease G is important for apoptosis in C. elegans. Nature 412, 90–94 (2001).
Parrish, J.Z. & Xue, D. Functional genomic analysis of apoptotic DNA degradation in C. elegans. Mol. Cell 11, 987–996 (2003).
Stanfield, G.M. & Horvitz, H.R. The ced-8 gene controls the timing of programmed cell deaths in C. elegans. Mol. Cell 5, 423–433 (2000).
Kokel, D., Li, Y., Qin, J. & Xue, D. The nongenotoxic carcinogens naphthalene and para-dichlorobenzene suppress apoptosis in Caenorhabditis elegans. Nat. Chem. Biol. 2, 338–345 (2006).
Conradt, B. & Horvitz, H.R. The C. elegans protein EGL-1 is required for programmed cell death and interacts with the Bcl-2-like protein CED-9. Cell 93, 519–529 (1998).
Xue, D., Shaham, S. & Horvitz, H.R. The Caenorhabditis elegans cell-death protein CED-3 is a cysteine protease with substrate specificities similar to those of the human CPP32 protease. Genes Dev. 10, 1073–1083 (1996).
Park, W.S. et al. Comprehensive identification of PIP3-regulated PH domains from C. elegans to H. sapiens by model prediction and live imaging. Mol. Cell 30, 381–392 (2008).
Tullet, J.M. et al. Direct inhibition of the longevity-promoting factor SKN-1 by insulin-like signaling in C. elegans. Cell 132, 1025–1038 (2008).
Datta, S.R., Brunet, A. & Greenberg, M.E. Cellular survival: a play in three Akts. Genes Dev. 13, 2905–2927 (1999).
Isakoff, S.J. et al. Identification and analysis of PH domain-containing targets of phosphatidylinositol 3-kinase using a novel in vivo assay in yeast. EMBO J. 17, 5374–5387 (1998).
Parent, C.A., Blacklock, B.J., Froehlich, W.M., Murphy, D.B. & Devreotes, P.N. G protein signaling events are activated at the leading edge of chemotactic cells. Cell 95, 81–91 (1998).
Meili, R. et al. Chemoattractant-mediated transient activation and membrane localization of Akt/PKB is required for efficient chemotaxis to cAMP in Dictyostelium. EMBO J. 18, 2092–2105 (1999).
Servant, G. et al. Polarization of chemoattractant receptor signaling during neutrophil chemotaxis. Science 287, 1037–1040 (2000).
Fayard, E., Tintignac, L.A., Baudry, A. & Hemmings, B.A. Protein kinase B/Akt at a glance. J. Cell Sci. 118, 5675–5678 (2005).
Stambolic, V. et al. Negative regulation of PKB/Akt-dependent cell survival by the tumor suppressor PTEN. Cell 95, 29–39 (1998).
Wu, X., Senechal, K., Neshat, M.S., Whang, Y.E. & Sawyers, C.L. The PTEN/MMAC1 tumor suppressor phosphatase functions as a negative regulator of the phosphoinositide 3-kinase/Akt pathway. Proc. Natl. Acad. Sci. USA 95, 15587–15591 (1998).
Kuo, Y.C. et al. Regulation of phosphorylation of Thr-308 of Akt, cell proliferation, and survival by the B55α regulatory subunit targeting of the protein phosphatase 2A holoenzyme to Akt. J. Biol. Chem. 283, 1882–1892 (2008).
Padmanabhan, S. et al. A PP2A regulatory subunit regulates C. elegans insulin/IGF-1 signaling by modulating AKT-1 phosphorylation. Cell 136, 939–951 (2009).
Gao, T., Furnari, F. & Newton, A.C. PHLPP: a phosphatase that directly dephosphorylates Akt, promotes apoptosis, and suppresses tumor growth. Mol. Cell 18, 13–24 (2005).
Brognard, J., Sierecki, E., Gao, T. & Newton, A.C. PHLPP and a second isoform, PHLPP2, differentially attenuate the amplitude of Akt signaling by regulating distinct Akt isoforms. Mol. Cell 25, 917–931 (2007).
Cheng, G.Z. et al. Advances of AKT pathway in human oncogenesis and as a target for anti-cancer drug discovery. Curr. Cancer Drug Targets 8, 2–6 (2008).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Wang, X. et al. C. elegans mitochondrial factor WAH-1 promotes phosphatidylserine externalization in apoptotic cells through phospholipid scramblase SCRM-1. Nat. Cell Biol. 9, 541–549 (2007).
Perrin, A.J. et al. Noncanonical control of C. elegans germline apoptosis by the insulin/IGF-1 and Ras/MAPK signaling pathways. Cell Death Differ. 20, 97–107 (2013).
Mello, C.C., Kramer, J.M., Stinchcomb, D. & Ambros, V. Efficient gene transfer in C.elegans: extrachromosomal maintenance and integration of transforming sequences. EMBO J. 10, 3959–3970 (1991).
Gu, T., Orita, S. & Han, M. Caenorhabditis elegans SUR-5, a novel but conserved protein, negatively regulates LET-60 Ras activity during vulval induction. Mol. Cell. Biol. 18, 4556–4564 (1998).
Tabara, H. et al. The rde-1 gene, RNA interference, and transposon silencing in C. elegans. Cell 99, 123–132 (1999).
Acknowledgements
We thank B. Derry (Hospital for Sick Children) for anti–AKT-1 antibodies, M. Han (University of Colorado) for some RNAi clones, Y. Kohara (Japan National Institute of Genetics) for akt-1 cDNA, M. Valencia and Y. Shi for making some of the constructs, S. Mitani (Tokyo Women's Medical University) and G. Ruvkun (Massachusetts General Hospital) for strains, R.R. Skeen-Gaar for assistance in generating transgenic strains and B.L. Harry, T. Blumenthal, B. Olwin, and J.M. Espinosa for comments on the manuscript. Some of the worm strains used in this study were kindly provided by the Caenorhabditis Genetics Center, which is funded by the US National Institutes of Health. This work was supported by US National Institutes of Health (grants R01 GM59083, R01 GM79097 and R01 GM088241 to D.X.).
Author information
Authors and Affiliations
Contributions
A.N. and D.X. conceived the research and designed experiments. A.N. carried out and analyzed experiments. K.D.S. assisted in some experiments. A.N. and D.X. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Analyses of the cps-2(sm8) mutant and anti–CNT-1 antibodies.
(a) Embryonic cell corpses were scored in the indicated animals. The y axis represents the average number of cell corpses and error bars are S.D. “ex” indicates an extrachromosomal array carrying the indicated construct. Statistical significance values were determined by two-way ANOVA, followed by Bonferroni comparison (n = 15 embryos). *, P < 0.001. All other points had P > 0.05. (b) RT-PCR analysis of N2 and cps-2(sm8) animals. Primer sets used for PCR are indicated with arrowheads in the cartoon. Left panel shows cnt-1 and right panel shows rpl-26 as a control. (c) Immunoblotting of N2 and cps-2(sm8) animals. Left panel is probed with anti-CNT-1 antibody and right panel is probed with anti-α-tubulin antibody. (d) Immunoblotting of recombinant GST-tCNT-1a and GST-tCNT-1b. Left panel is probed with anti-CNT-1 antibody and right panel is probed with anti-GST antibody. (e) The uncropped image of α-tubulin blot shown in Fig. 2c.
Supplementary Figure 3 Further analysis of the roles of AKT kinase genes, age-1 and daf-16 in C. elegans cell death.
Cell corpses were scored in the indicated strains. The y axis represents average number of cell corpses scored and error bars represent S.D. Statistical significance values were determined by two-way ANOVA, followed by Bonferroni comparison (n = 15 embryos for each stage). *, P < 0.001; **, P < 0.05.
Supplementary Figure 4 Analyses of the importance of AKT-1 and AKT-2 kinase activities and CNT-1 in cell death in C. elegans.
Cell corpses were scored in the indicated strains. The y axis represents average number of cell corpses scored and error bars represent S.D. Statistical significance values were determined by two-way ANOVA, followed by Bonferroni comparison (n = 15 embryos for each stage). *, P < 0.001.
Supplementary Figure 5 Expression levels of CNT-1a(K284A) and tCNT-1a(K284A) are comparable to those of wild-type proteins both in vitro and in vivo.
(a) Autoradiogram of GST-CNT-1a, GST-CNT-1a(K284A), GST-tCNT-1a, and GST-tCNT-1a(K284A) synthesized in rabbit reticulocyte lysate and labeled with 35S-Methionine (*). (b) Immunoblotting of worm lysates using an anti-CNT-1 antibody. Ten transgenic animals of the indicated genotype for each lane were subjected to immunoblotting.
Supplementary Figure 6 The expression level of CNT-1 in C. elegans is higher than that of AKT-1.
(a) Left, immunoblotting of 100 N2 embryos at 1.5-fold stage and 1 pmol of GST-CNT-1a purified from bacteria using an anti-CNT-1 antibody. Right, quantification of the relative CNT-1 amount, which was determined from 3 independent experiments, including the one shown in the left panel, as described in Supplementary Fig. 2d. The data are presented as relative CNT-1 amount (CNT-1a+CNT-1b) and S.D. (b) Left, immunoblotting of 100 N2 embryos at 1.5-fold stage and 1 pmol of AKT-1-His6 purified from bacteria using an anti-AKT-1 antibody. Right, the relative AKT-1 amount was determined as in a. Please see Online Methods for detail. Statistical significance values were determined by two-tailed Student’s t-test (n = 3 independent experiments). *, P < 0.001 (a). P > 0.05 (b).
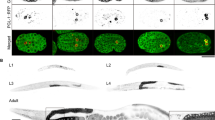
Supplementary Figure 7 Analyses of CNT-1 protein localization in various embryos.
(a-f) The CNT-1 protein is expressed in all cells during embryogenesis. Embryos with the indicated genotype and developmental stage were stained with an anti-CNT-1 antibody. FITC (left) and DIC (right) images are shown. (g-l) CED-3 uncleavable form of CNT-1a does not translocate to the plasma membrane after activation of CED-3 and apoptosis. Transgenic or non-transgenic embryos with the indicated genotype were stained with an anti-CNT-1 antibody. In all panels, -HS, no heat-shock treatment. +HS, with heat-shock treatment. Scale bars represent 10 mm.
Supplementary Figure 8 The cnt-1(tm2313) deletion does not affect life span or thermotolerance.
Animals with the indicated genotype were subjected to either life span analysis (a) or heat stress resistance analysis (b) (see Online Methods).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–5 and Supplementary Note (PDF 2803 kb)
Rights and permissions
About this article
Cite this article
Nakagawa, A., Sullivan, K. & Xue, D. Caspase-activated phosphoinositide binding by CNT-1 promotes apoptosis by inhibiting the AKT pathway. Nat Struct Mol Biol 21, 1082–1090 (2014). https://doi.org/10.1038/nsmb.2915
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2915
This article is cited by
-
Programmed cell death and clearance of cell corpses in Caenorhabditis elegans
Cellular and Molecular Life Sciences (2016)