Abstract
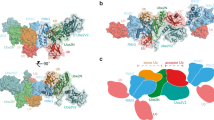
RING E3 ligase–catalyzed formation of K63-linked ubiquitin chains by the Ube2V2–Ubc13 E2 complex is required in many important biological processes. Here we report the structure of the RING-domain dimer of rat RNF4 in complex with a human Ubc13∼Ub conjugate and Ube2V2. The structure has captured Ube2V2 bound to the acceptor (priming) ubiquitin with K63 in a position favorable for attack on the linkage between Ubc13 and the donor (second) ubiquitin held in the active 'folded back' conformation by the RING domain of RNF4. We verified the interfaces identified in the structure by in vitro ubiquitination assays of site-directed mutants. To our knowledge, this represents the first view of synthesis of K63-linked ubiquitin chains in which both substrate ubiquitin and ubiquitin-loaded E2 are juxtaposed to allow E3 ligase–mediated catalysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Metzger, M.B., Pruneda, J.N., Klevit, R.E. & Weissman, A.M. RING-type E3 ligases: master manipulators of E2 ubiquitin-conjugating enzymes and ubiquitination. Biochim. Biophys. Acta 1843, 47–60 (2014).
Scheffner, M. & Kumar, S. Mammalian HECT ubiquitin-protein ligases: biological and pathophysiological aspects. Biochim. Biophys. Acta 1843, 61–74 (2014).
Dou, H., Buetow, L., Sibbet, G.J., Cameron, K. & Huang, D.T. BIRC7–E2 ubiquitin conjugate structure reveals the mechanism of ubiquitin transfer by a RING dimer. Nat. Struct. Mol. Biol. 19, 876–883 (2012).
Plechanovová, A., Jaffray, E.G., Tatham, M.H., Naismith, J.H. & Hay, R.T. Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis. Nature 489, 115–120 (2012).
Pruneda, J.N. et al. Structure of an E3:E2∼Ub complex reveals an allosteric mechanism shared among RING/U-box ligases. Mol. Cell 47, 933–942 (2012).
Berndsen, C.E. & Wolberger, C. New insights into ubiquitin E3 ligase mechanism. Nat. Struct. Mol. Biol. 21, 301–307 (2014).
Tatham, M.H. et al. RNF4 is a poly-SUMO-specific E3 ubiquitin ligase required for arsenic-induced PML degradation. Nat. Cell Biol. 10, 538–546 (2008).
Galanty, Y., Belotserkovskaya, R., Coates, J. & Jackson, S.P. RNF4, a SUMO-targeted ubiquitin E3 ligase, promotes DNA double-strand break repair. Genes Dev. 26, 1179–1195 (2012).
Guzzo, C.M. et al. RNF4-dependent hybrid SUMO-ubiquitin chains are signals for RAP80 and thereby mediate the recruitment of BRCA1 to sites of DNA damage. Sci. Signal. 5, ra88 (2012).
Vyas, R. et al. RNF4 is required for DNA double-strand break repair in vivo. Cell Death Differ. 20, 490–502 (2013).
Yin, Y. et al. SUMO-targeted ubiquitin E3 ligase RNF4 is required for the response of human cells to DNA damage. Genes Dev. 26, 1196–1208 (2012).
Rojas-Fernandez, A. et al. SUMO chain-induced dimerization activates RNF4. Mol. Cell 53, 880–892 (2014).
Hofmann, R.M. & Pickart, C.M. Noncanonical MMS2-encoded ubiquitin-conjugating enzyme functions in assembly of novel polyubiquitin chains for DNA repair. Cell 96, 645–653 (1999).
Eddins, M.J., Carlile, C.M., Gomez, K.M., Pickart, C.M. & Wolberger, C. Mms2–Ubc13 covalently bound to ubiquitin reveals the structural basis of linkage-specific polyubiquitin chain formation. Nat. Struct. Mol. Biol. 13, 915–920 (2006).
Tatham, M.H., Plechanovova, A., Jaffray, E.G., Salmen, H. & Hay, R.T. Ube2W conjugates ubiquitin to alpha-amino groups of protein N-termini. Biochem. J. 453, 137–145 (2013).
Lewis, M.J., Saltibus, L.F., Hau, D.D., Xiao, W. & Spyracopoulos, L. Structural basis for non-covalent interaction between ubiquitin and the ubiquitin conjugating enzyme variant human MMS2. J. Biomol. NMR 34, 89–100 (2006).
McKenna, S. et al. Energetics and specificity of interactions within Ub.Uev.Ubc13 human ubiquitin conjugation complexes. Biochemistry 42, 7922–7930 (2003).
McKenna, S. et al. Noncovalent interaction between ubiquitin and the human DNA repair protein Mms2 is required for Ubc13-mediated polyubiquitination. J. Biol. Chem. 276, 40120–40126 (2001).
Moraes, T.F. et al. Crystal structure of the human ubiquitin conjugating enzyme complex, hMms2–hUbc13. Nat. Struct. Biol. 8, 669–673 (2001).
Spyracopoulos, L., Lewis, M.J. & Saltibus, L.F. Main chain and side chain dynamics of the ubiquitin conjugating enzyme variant human Mms2 in the free and ubiquitin-bound states. Biochemistry 44, 8770–8781 (2005).
VanDemark, A.P., Hofmann, R.M., Tsui, C., Pickart, C.M. & Wolberger, C. Molecular insights into polyubiquitin chain assembly: crystal structure of the Mms2/Ubc13 heterodimer. Cell 105, 711–720 (2001).
Zhang, M. et al. Chaperoned ubiquitylation: crystal structures of the CHIP U box E3 ubiquitin ligase and a CHIP-Ubc13-Uev1a complex. Mol. Cell 20, 525–538 (2005).
Reverter, D. & Lima, C.D. Insights into E3 ligase activity revealed by a SUMO–RanGAP1–Ubc9–Nup358 complex. Nature 435, 687–692 (2005).
Scott, D.C. et al. Structure of a RING E3 trapped in action reveals ligation mechanism for the ubiquitin-like protein NEDD8. Cell 157, 1671–1684 (2014).
Yunus, A.A. & Lima, C.D. Lysine activation and functional analysis of E2-mediated conjugation in the SUMO pathway. Nat. Struct. Mol. Biol. 13, 491–499 (2006).
Berndsen, C.E., Wiener, R., Yu, I.W., Ringel, A.E. & Wolberger, C. A conserved asparagine has a structural role in ubiquitin-conjugating enzymes. Nat. Chem. Biol. 9, 154–156 (2013).
Wu, P.Y. et al. A conserved catalytic residue in the ubiquitin-conjugating enzyme family. EMBO J. 22, 5241–5250 (2003).
Chang, L., Zhang, Z., Yang, J., McLaughlin, S.H. & Barford, D. Molecular architecture and mechanism of the anaphase-promoting complex. Nature 513, 388–393 (2014).
Plechanovová, A. et al. Mechanism of ubiquitylation by dimeric RING ligase RNF4. Nat. Struct. Mol. Biol. 18, 1052–1059 (2011).
Collaborative Computational Project, No. 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D Biol. Crystallogr. 62, 72–82 (2006).
Evans, P.R. in Proc. CCP4 Study Weekend (eds. Wilson, K.S., Davies, G., Ashton, A.W. & Bailey, S.) (CCLRC Daresbury Laboratory, 1997).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–190 (2010).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Acknowledgements
We thank M. Alphey for assistance with data collection and the Division of Signal Transduction Therapy, University of Dundee for the gift of His-UBE1. This work was supported by grants from Cancer Research UK (C434/A13067), the Wellcome Trust (098391/Z/12/Z) and Biotechnology and Biological Sciences Research Council (BB/J016004/1) to R.T.H. Both R.T.H. and J.H.N. (WT100209MA) are supported as Senior Investigators of the Wellcome Trust. J.H.N. is supported as a Royal Society Wolfson Merit Award Holder.
Author information
Authors and Affiliations
Contributions
E.B. cloned, expressed and purified proteins, carried out structural analysis, conducted biochemical experiments and interpreted the data. A.P. cloned, expressed and purified proteins, conducted biochemical experiments and interpreted the data. E.G.J. purified recombinant proteins and carried out biochemical analysis. J.H.N. contributed to structural analysis and data analysis. E.B., A.P., J.H.N. and R.T.H. wrote the paper. R.T.H. conceived the project and contributed to data analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Stimulation of RNF4-mediated synthesis of unanchored K63-linked ubiquitin chains by SUMO-2 chains and biochemical analysis of the isopeptide linked Ubc13~Ub conjugate.
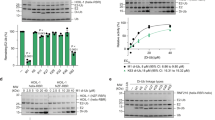
(a) Coomassie-blue stained SDS-PAGE (top panel) and Western blot (bottom panel) analysis of ubiquitination assays with and without 4xSUMO-2. A negative control without ATP is shown with 4xSUMO-2 present. Time points are denoted above the gels.
Ubc13 C87K was used to form a stable isopeptide linked Ubc13~Ub conjugate for crystallography. Due to the identification of non-specific ubiquitination of Ubc13 K92 in the Ubc13 C87A negative control, K92A was also introduced in combination with C87K to prevent non-specific ubiquitination of Ubc13.
(b) Coomassie-blue stained SDS-PAGE analysis of the ability of Ubc13 C87K and Ubc13 C87K K92A to form an isopeptide linked conjugate with ubiquitin. Negative controls containing the C87A mutation were tested in parallel. Time points are shown above the gels.
(c) Coomassie-blue stained SDS-PAGE analysis of ubiquitination assays showing the effect of the introduction of K92A on Ubc13 activity compared to WT Ubc13. Reactions contained 0.1 µM E1, 1 µM Ubc13, 1 µM Ube2V2, 0.55 µM RNF4, 5.5 µM Ub~4xSUMO-2, 20 µM ubiquitin, 3 mM ATP, 5 mM MgCl2, 50 mM Tris pH 7.5, 150 mM NaCl, 0.5 mM TCEP, and 0.1 % NP40. Reactions were incubated at 37 °C. Time points are shown above the gels. The red asterisk indicates formation of di-ubiquitin.
(d) The chromatogram shows separation of the Ube2V2–Ubc13~Ub complex and free ubiquitin by size exclusion chromatography. Elution fractions analysed by SDS-PAGE are denoted with magenta, green and black bars below the chromatogram.
(e) Coomassie stained SDS-PAGE analysis of size exclusion chromatography showing input to the chromatography numbered 1 – 3 and analysis of the elution fractions highlighted in d.
.
Supplementary Figure 2 Comparison of the active site of the Ubc13~Ub–RING complexes with and without Ube2V2 and the UbcH5A~Ub–RING complex (4AP4).
E2’s from the Ubc13~Ub–RING complexes with and without Ube2V2 and the UbcH5A~Ub–RING complex (4AP4) (Plechanovová, A. et al., Nature. 489, 115–20, 2012) were superimposed. Ubc13, Ub and RING-RING from the Ubc13~Ub–RING complex are coloured in pale green, pale orange and pale pink respectively. UbcH5A, Ub and RING-RING from the UbcH5A~Ub–RING complex are coloured in teal, grey and purple respectively. Ube2V2, Ubc13, Ub and RING-RING from the Ube2V2–Ubc13~Ub–RING complex are coloured in blue, dark green, yellow and dark pink respectively. Ub* is coloured in orange.
(a) Molecular detail of the active site of the Ubc13~Ub–RING complex and the UbcH5A~Ub–RING complex where the E2’s are superimposed.
(b) Molecular detail of the active site of the Ube2V2–Ubc13~Ub–RING complex and the Ubc13~Ub–RING complex where Ubc13 from each complex is superimposed.
Supplementary Figure 3 Analysis of the crystal packing of the Ube2V2–Ubc13~Ub–RING complex, revealing specificity for K63-linked polyubiquitination and electron density at ubiquitin Q62, K63 and E64.
The complex polymerises in the crystal such that Ub* (orange) from a symmetry related complex binds to Ube2V2 (dark blue), which orients Ub* such that K63 is presented to the isopeptide linked Ubc13~Ub conjugate.
(a) The crystal packing of three Ube2V2–Ubc13~Ub–RING complexes is shown.
Phases were generated from a molecular replacement solution, which had Q62, K63, and E64 deleted from both the acceptor ubiquitin (Ub*) and the donor ubiquitin. All b-factors were deleted and reset to single value of 20. The structure was refined with a single overall b-factor then refined again with individual b-factors.
(b) Difference electron density (Fo–Fc) contoured at 1.5σ is shown in dark blue for Ub*(orange).
(c) 2Fo–Fc electron density contoured at 0.5σ is shown in grey for Ub*.
(d) Difference electron density (Fo–Fc) contoured at 1.5σ is shown in dark blue for the donor ubiquitin (yellow). Ubc13 is coloured green and Ube2V2 is coloured blue.
(e) 2Fo–Fc electron density contoured at 0.5σ is shown in grey for the donor ubiquitin (yellow).
Supplementary Figure 4 Comparison of the Ube2V2–Ubc13~Ub–RING complex with the MMS2–Ubc13~Ub complex (2GMI) and the UEV1A–Ubc13–CHIP complex (2C2V).
Ube2V2, Ubc13 and donor Ub from the Ube2V2–Ubc13~Ub–RING complex are coloured in blue, green and yellow respectively. The linear fusion dimer of the RING domain of RNF4 is coloured in dark and pale pink. Ub* is coloured in orange. MMS2, Ubc13 and Ub from the MMS2–Ubc13~Ub complex (2GMI) (Eddins, M. J. et al., Nat. Struct. Mol. Biol. 13, 915–20, 2006) are coloured in cyan, grey and magenta respectively. Ubiquitin from the MMS2–Ubc13~Ub complex has been transformed using the symmetry operator X+1/2, –Y+1/2, –Z to show its role as the acceptor ubiquitin.
(a) Ubc13 molecules from both complexes have been superimposed to highlight the different orientation of Ube2V2 and MMS2 with respect to Ubc13 and the resultant effect on the positioning of the acceptor Ub K63.
(b) MMS2 and Ube2V2 molecules from the two complexes have been superimposed to show slight differences in the positioning of the acceptor Ub with respect to MMS2 and Ube2V2. This is likely due to differences in crystal packing.
(c) The Ube2V2–Ubc13~Ub–RING complex is coloured the same as in a. UEV1A and Ubc13 from the UEV1A–Ubc13–CHIP complex (2C2V) (Zhang, M. et al., Mol. Cell. 20, 525–38, 2005) are coloured in dark blue and cyan respectively. One protomer of the CHIP dimer is coloured in grey and the other is coloured in purple. These two structures have been superimposed using Ubc13 showing the similarity between the Ubc13–Ube2V2 and the Ubc13–UEV1A interfaces. Although CHIP is a U-box containing E3 ligase, it adopts the same orientation with respect to the Ubc13–UEV1A complex as the RING domain dimer of RNF4 in the Ube2V2–Ubc13~Ub–RING complex.
Supplementary Figure 5 Raw data for ITC experiments for the Ubc13-Ube2V2 interaction.
ITC experimental data was collected for the WT Ubc13–Ube2V2 interaction and mutants.
(a) The top panel for each experiment shows the raw data (black line) and the background fitting (red line). Data were fitted using MicroCal Origin software. The bottom panel for each experiment shows integrated heats of injection (black squares) and the best fit using a one binding site model (black line).
(b) ITC derived Kd measurements (± error) of Ube2V2–Ubc13 interaction for WT and mutants.
Supplementary Figure 6 Mutational analysis of the Ube2V2–Ubc13~Ub–RING complex.
Ubc13 (green) of the Ube2V2–Ubc13~Ub–RING complex was superimposed onto UbcH5A (green) of the UbcH5A~Ub–RING complex (4AP4) (Plechanovová, A. et al., Nature. 489, 115–20, 2012). Ube2V2 is blue and Ub* is orange. Both structures contain donor ubiquitin molecules (yellow) and the RING domain dimer of RNF4 (dark and pale pink).
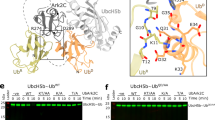
(a) Detailed molecular interface between Ube2V2 (blue) α1 helix and Ubc13 (green) from the Ube2V2–Ubc13~Ub–RING complex.
(b) Detailed surface of UbcH5A (green) from the UbcH5A~Ub–RING complex (4AP4) similar to the surface shown in a for Ubc13.
(c) Coomassie-blue stained SDS-PAGE analysis of ubiquitination assays containing WT or mutants of Ubc13 and Ube2V2. The red asterisk indicates formation of di-ubiquitin.
(d) Sequence alignment for a selection of E2s containing residues H77 to K92 for Ubc13. Conserved regions are coloured in red while non-conserved regions are coloured in black.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 2201 kb)
Supplementary Data Set 1
Individual SYPRO orange–stained SDS-PAGE for the lysine-discharge assay (PDF 5784 kb)
Supplementary Data Set 2
Individual rate calculations for the lysine-discharge assay (XLS 252 kb)
Rights and permissions
About this article
Cite this article
Branigan, E., Plechanovová, A., Jaffray, E. et al. Structural basis for the RING-catalyzed synthesis of K63-linked ubiquitin chains. Nat Struct Mol Biol 22, 597–602 (2015). https://doi.org/10.1038/nsmb.3052
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3052
This article is cited by
-
Concerted SUMO-targeted ubiquitin ligase activities of TOPORS and RNF4 are essential for stress management and cell proliferation
Nature Structural & Molecular Biology (2024)
-
Structural snapshots along K48-linked ubiquitin chain formation by the HECT E3 UBR5
Nature Chemical Biology (2024)
-
Cryo-EM structures of Uba7 reveal the molecular basis for ISG15 activation and E1-E2 thioester transfer
Nature Communications (2023)
-
Structural and functional asymmetry of RING trimerization controls priming and extension events in TRIM5α autoubiquitylation
Nature Communications (2022)
-
Structure of UBE2K–Ub/E3/polyUb reveals mechanisms of K48-linked Ub chain extension
Nature Chemical Biology (2022)