Abstract
H2A.Z, a widely conserved histone variant, is evicted from chromatin by the histone chaperone ANP32E. However, to date, no deposition chaperone for H2A.Z is known in metazoans. Here, we identify YL1 as a specific H2A.Z-deposition chaperone. The 2.7-Å-resolution crystal structure of the human YL1–H2A.Z–H2B complex shows that YL1 binding, similarly to ANP32E binding, triggers an extension of the H2A.Z αC helix. The interaction with YL1 is, however, more extensive and includes both the extended acidic patch and the entire DNA-binding surface of H2A.Z–H2B. Substitution of only four amino acid residues of H2A is sufficient for the formation of an H2A.Z-like interface specifically recognized by YL1. Collectively, our data reveal the molecular basis of H2A.Z-specific recognition by YL1 and shed light on the mechanism of H2A.Z transfer to the nucleosome by the ATP-dependent chromatin-remodeling complexes SRCAP and P400–TIP60.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Larochelle, M. & Gaudreau, L. H2A.Z has a function reminiscent of an activator required for preferential binding to intergenic DNA. EMBO J. 22, 4512–4522 (2003).
Raisner, R.M. et al. Histone variant H2A.Z marks the 5′ ends of both active and inactive genes in euchromatin. Cell 123, 233–248 (2005).
Papamichos-Chronakis, M., Watanabe, S., Rando, O.J. & Peterson, C.L. Global regulation of H2A.Z localization by the INO80 chromatin-remodeling enzyme is essential for genome integrity. Cell 144, 200–213 (2011).
Clarkson, M.J., Wells, J.R., Gibson, F., Saint, R. & Tremethick, D.J. Regions of variant histone His2AvD required for Drosophila development. Nature 399, 694–697 (1999).
Faast, R. et al. Histone variant H2A.Z is required for early mammalian development. Curr. Biol. 11, 1183–1187 (2001).
Hong, J. et al. The catalytic subunit of the SWR1 remodeler is a histone chaperone for the H2A.Z-H2B dimer. Mol. Cell 53, 498–505 (2014).
Obri, A. et al. ANP32E is a histone chaperone that removes H2A.Z from chromatin. Nature 505, 648–653 (2014).
Suto, R.K., Clarkson, M.J., Tremethick, D.J. & Luger, K. Crystal structure of a nucleosome core particle containing the variant histone H2A.Z. Nat. Struct. Biol. 7, 1121–1124 (2000).
Mizuguchi, G. et al. ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science 303, 343–348 (2004).
Kobor, M.S. et al. A protein complex containing the conserved Swi2/Snf2-related ATPase Swr1p deposits histone variant H2A.Z into euchromatin. PLoS Biol. 2, E131 (2004).
Krogan, N.J. et al. A Snf2 family ATPase complex required for recruitment of the histone H2A variant Htz1. Mol. Cell 12, 1565–1576 (2003).
Luk, E. et al. Stepwise histone replacement by SWR1 requires dual activation with histone H2A.Z and canonical nucleosome. Cell 143, 725–736 (2010).
Wu, W.H. et al. N terminus of Swr1 binds to histone H2AZ and provides a platform for subunit assembly in the chromatin remodeling complex. J. Biol. Chem. 284, 6200–6207 (2009).
Yen, K., Vinayachandran, V. & Pugh, B.F. SWR-C and INO80 chromatin remodelers recognize nucleosome-free regions near +1 nucleosomes. Cell 154, 1246–1256 (2013).
Luk, E. et al. Chz1, a nuclear chaperone for histone H2A.Z. Mol. Cell 25, 357–368 (2007).
Ranjan, A. et al. Nucleosome-free region dominates histone acetylation in targeting SWR1 to promoters for H2A.Z replacement. Cell 154, 1232–1245 (2013).
Wu, W.H. et al. Swc2 is a widely conserved H2AZ-binding module essential for ATP-dependent histone exchange. Nat. Struct. Mol. Biol. 12, 1064–1071 (2005).
Zhou, Z. et al. NMR structure of chaperone Chz1 complexed with histones H2A.Z-H2B. Nat. Struct. Mol. Biol. 15, 868–869 (2008).
Cai, Y. et al. The mammalian YL1 protein is a shared subunit of the TRRAP/TIP60 histone acetyltransferase and SRCAP complexes. J. Biol. Chem. 280, 13665–13670 (2005).
Jensen, K., Santisteban, M.S., Urekar, C. & Smith, M.M. Histone H2A.Z acid patch residues required for deposition and function. Mol. Genet. Genomics 285, 287–296 (2011).
Barbera, A.J. et al. The nucleosomal surface as a docking station for Kaposi's sarcoma herpesvirus LANA. Science 311, 856–861 (2006).
Makde, R.D., England, J.R., Yennawar, H.P. & Tan, S. Structure of RCC1 chromatin factor bound to the nucleosome core particle. Nature 467, 562–566 (2010).
Armache, K.J., Garlick, J.D., Canzio, D., Narlikar, G.J. & Kingston, R.E. Structural basis of silencing: Sir3 BAH domain in complex with a nucleosome at 3.0 Å resolution. Science 334, 977–982 (2011).
Kato, H. et al. A conserved mechanism for centromeric nucleosome recognition by centromere protein CENP-C. Science 340, 1110–1113 (2013).
McGinty, R.K., Henrici, R.C. & Tan, S. Crystal structure of the PRC1 ubiquitylation module bound to the nucleosome. Nature 514, 591–596 (2014).
Hamiche, A. & Richard-Foy, H. Characterization of specific nucleosomal states by use of selective substitution reagents in model octamer and tetramer structures. Methods 19, 457–464 (1999).
Ouararhni, K. et al. The histone variant mH2A1.1 interferes with transcription by down-regulating PARP-1 enzymatic activity. Genes Dev. 20, 3324–3336 (2006).
Shuaib, M., Ouararhni, K., Dimitrov, S. & Hamiche, A. HJURP binds CENP-A via a highly conserved N-terminal domain and mediates its deposition at centromeres. Proc. Natl. Acad. Sci. USA 107, 1349–1354 (2010).
Luger, K., Rechsteiner, T.J. & Richmond, T.J. Expression and purification of recombinant histones and nucleosome reconstitution. Methods Mol. Biol. 119, 1–16 (1999).
Hamiche, A., Kang, J.G., Dennis, C., Xiao, H. & Wu, C. Histone tails modulate nucleosome mobility and regulate ATP-dependent nucleosome sliding by NURF. Proc. Natl. Acad. Sci. USA 98, 14316–14321 (2001).
Diebold, M.L., Fribourg, S., Koch, M., Metzger, T. & Romier, C. Deciphering correct strategies for multiprotein complex assembly by co-expression: application to complexes as large as the histone octamer. J. Struct. Biol. 175, 178–188 (2011).
Simpson, R.T., Thoma, F. & Brubaker, J.M. Chromatin reconstituted from tandemly repeated cloned DNA fragments and core histones: a model system for study of higher order structure. Cell 42, 799–808 (1985).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D Biol. Crystallogr. 66, 22–25 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Blanc, E. et al. Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT. Acta Crystallogr. D Biol. Crystallogr. 60, 2210–2221 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Acknowledgements
This work was supported by institutional funds from the Centre Nationale de la Recherche Scientifique (CNRS, France), the Institut National de la Santé et de la Recherche Médicale (INSERM, France), the Université de Strasbourg (UDS) and the Université de Grenoble Alpes and by grants from the Ligue Nationale Contre le Cancer Equipe Labellisée (A.H.), the Fondation ARC pour la Recherche sur le Cancer (C.R.), the UDS Institute for Advanced Study (USIAS) (A.H.), IdEX Attractivité UDS (C.R.), the French National Research Agency (ANR; VariZome, contract ANR-12-BSV8-0018-01; Nucleoplat, contract NT09_476241), the French National Cancer Institute (INCA) (INCa_4496 and INCa_4454), la Fondation pour la Recherche Medicale, the French Infrastructure for Integrated Structural Biology (FRISBI; ANR-10-INSB-05-01) and by Instruct as part of the European Strategy Forum on Research Infrastructures (ESFRI). We thank the members of the European Synchrotron Radiation Facility/European Molecular Biology Laboratory joint structural biology group and staff of the synchrotron SOLEIL for use of beamline facilities and for help during data collection. We thank S. Tan (Pennsylvania State University) for fruitful discussions, C. Birck (IGBMC) for help with biophysical characterization and C. Da Veiga for help with ITC experiments.
Author information
Authors and Affiliations
Contributions
C.M.L., M.M. and K.O. contributed equally to this work. C.M.L. performed complex purification and characterization, designed and analyzed the human mutants and contributed to the text. K.O. performed complex purification, chromatin assembly and eviction assays and analysis of yeast mutants. C.P. and A.O. built constructs and performed preliminary complex purification. I.S. generated baculovirus constructs and performed purification. M.I. performed preliminary pulldown assays. C.R. and M.M. solved the YL1 ZID–H2A.Z–H2B structure and designed the mutants. E.E. and M.M. performed the ITC experiments. S.D. provided important reagents. A.H., C.R. and S.D. designed experiments, analyzed data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 YL1 specifically associates with the H2A.Z complex.
a) Silver staining of proteins associated with double immuno-affinity (FLAG-HA) purified e-H2A.com and e-H2A.Z.com complexes.
b) Western blotting analysis of the e-H2A.com and e-H2A.Z.com complexes shown in (a). Each complex was probed for the presence of the tagged protein (anti-HA) as well as for YL1, ANP32E, H2A and H2A.Z.
c) Mass spectrometry identification of additional proteins associated with e-H2A.com shown in (a). For each identified protein the number of unique peptides is given.
d) Mass spectrometry identification of additional proteins associated with e-H2A.Z.com shown in (a). For each identified protein the number of unique peptides is given.
e) Mass spectrometry identification of additional proteins associated with e-YL1.com shown in Figure 1a. For each identified protein the number of unique peptides is given.
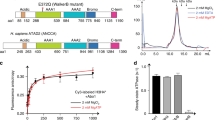
f) Alignment of the YL1–ZID sequences from vertebrates, insects, plants and fungi. Conservation of the residues is represented by shades of blue. Numbering above and underneath the sequences corresponds to YL1 sequences from human and S. cerevisiae (yeast SWC2), respectively. Secondary structure elements of human YL1, as seen in the structure, are displayed above the alignment (helices, cylinders; coils, lines). The various regions of the YL1–ZID described in the text are indicated (ZIDN, ZIDM1, ZIDM2, ZIDC). Stars above and underneath the alignment mark residues of human YL1 and yeast SWC2 that were mutated in our study: m1 mutant (dark blue), m2 mutant (light blue), and m1m2 mutant (combination of m1 and m2 mutants).
Supplementary Figure 2 Alignment of full-length YL1 and stereo view of the elongated αC helix.
a) Sequences of YL1 proteins from different organisms have been aligned. For clarity, fungi sequences have not been included since they are twice as long as their other homologs. See alignment of the YL1–ZID region in Supplementary Figure 1f for incorporation of the fungi sequences. Conservation is shown with different hues of blue (strong, dark blue; weak, light blue). Red boxes highlight the three main regions conserved throughout evolution (fungi display similar conservation patterns as those shown on this figure). Numbering above the sequences corresponds to the human protein.
b) Stereo view of the Fo–Fc simulated-annealing omit map contoured at 2.5 σ of H2A.Z elongated αC helix in the YL1–ZID–H2A.Z–H2B complex. Residues spanning the end of the canonical plus the extended part of the αC helix were removed prior to simulated-annealing.
Supplementary Figure 3 Human YL1, human ANP32E and yeast SWR1 all extend the αC helix, but only YL1 entirely covers the H2A.Z–H2B DNA-binding surface.
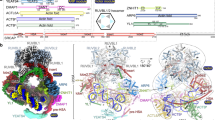
a) Superposition of (left) human YL1–ZID–H2A.Z–H2B (colored violet, orange and light gray, respectively) and human ANP32E–ZID–H2A.Z–H2B (colored blue, red and gray, respectively) complexes and of (right) human ANP32E–ZID–H2A.Z–H2B (colored blue, red and gray, respectively) and yeast SWR1–Z–H2A.Z–H2B (colored green, light red and dark gray, respectively) complexes. The common regions of interaction between YL1, ANP32E and SWR1 with the H2A.Z–H2B pair are boxed and labeled (ZIDM1 and ZIDM2). All H2A.Z histone chaperones extend H2A.Z αC helix, but only YL1 interacts extensively with the variant histone pair. In addition, YL1 binds to H2A.Z α3–αC region with a different conformation that ANP32E and SWR1, these latter chaperones showing high structural similarity.
b) Superposition of YL1 αN2 (violet) and ANP32E (blue) αN helices. The side chains of the hydrophobic residues of these helices that form hydrophobic interactions with the same set of hydrophobic residues from H2A.Z and H2B are shown as sticks. These hydrophobic residues from the histone chaperones occupy the same positions explaining that despite their difference in composition and the antiparallel orientation of the YL1 αN2 and ANP32E αN helices, they have the same effect in favoring H2A.Z αC helix extension. The only exception concerns YL1 Y30 that occupies a unique position and forms an YL1-specific hydrogen bonding network with residues from YL1 and H2A.Z.
c) Ribbon representation of (left) the superposed human YL1–ZID–H2A.Z–H2B (colored violet, orange and light gray, respectively) complex and H2A.Z–H2B histone pair (not shown) bound to a nucleosomal DNA region (green) and (right) the superposed human ANP32E–ZID–H2A.Z–H2B (colored blue, red and gray, respectively) complex and H2A.Z–H2B histone pair (not shown) bound to a nucleosomal DNA region (green). The C-terminal region of the YL1–ZID completely covers the H2A.Z–H2B DNA-binding surface.
d) Surface electrostatic potential of the DNA-binding region of the H2A.Z–H2B histone dimer (–5 kBT, 5 kBT; kB, Boltzmann constant; blue: positive, red: negative). Left: The H2A.Z–H2B dimer displays a highly positive electrostatic potential in its DNA binding groove. The YL1–ZID binding to the H2A.Z–H2B dimer is displayed as ribbon. Right panel: Same as left, but with YL1–ZID included in the surface electrostatic potential calculation. Upon binding, the YL1–ZID not only covers the DNA-binding region of the H2A.Z–H2B dimer, but completely changes the electrostatic potential displayed on this face of the complex, rendering it more negative and thus, repulsive for DNA interactions.
Supplementary Figure 4 YL1 ZIDN has the ability to interact with H2A.Z–H2B through an arginine anchor feature.
a) Dimer of the YL1–ZID–H2A.Z–H2B complex observed in the crystal, where dimerization occurs through stacking of YL1 αN1 helices. At this dimeric interface, the N-terminal region of the YL1–ZID (ZIDN) from one complex interacts with the H2A.Z–H2B pair from a symmetry-related YL1–ZID–H2A.Z–H2B complex. This interaction between the ZIDN and the H2A.Z–H2B pair includes an arginine anchor, where an YL1 arginine binds to the acidic patch pocket of H2A.Z. This interaction is reminiscent of interactions made by several transcription and epigenetic effectors when bound to the nucleosome. In addition, ZIDN conserved residues are involved in this interaction that is only possible due to the presence of H2A.Z-specific G92 (an asparagine in H2A), enabling YL1–ZIDN to lay onto H2A.Z.
b) Examples of arginine anchor features. Upper left: RingIB (from PCR1 complex) binding to the H2A nucleosome (PDB code 4R8P). Bottom: RCC1 binding to the H2A nucleosome (PDB code 3MVD). Upper right: YL1–ZIDN binding to H2A.Z (this study). The position of H2A N89 and equivalent H2A.Z G92 are displayed, showing that the presence of a bulkier side chain at this position affects the way of binding of the effectors to H2A/H2A.Z.
c) Stereo view of the Fo–Fc simulated-annealing omit map contoured at 2.5 σ of H2A.Z α2 helix C-terminus and L2 loop in the YL1–ZID–H2A.Z–H2B complex. Residues spanning this region were removed prior to simulated-annealing.
Supplementary Figure 5 YL1 ZID–H2A.Z–H2B is monomeric in solution, and the YL1 and SWC2 N-terminal regions (ZIDN) are required for proper binding to human H2A.Z but not to yeast HTZ1.
a) Size exclusion chromatography purification profile on a Superdex 200 16/60 (left) and analytical ultracentrifugation (AUC) data of the YL1–ZID–H2A.Z–H2B complex. This analysis shows that the YL1–ZID–H2A.Z–H2B complex is monomeric in solution.
b) Ribbon representation of the YL1–ZID–H2A.Z–H2B complex as observed in the structure (left; ZIDN colored violet in a position interacting with a symmetry-related H2A.Z–H2B dimer) and of the model of the YL1–ZID–H2A.Z–H2B complex (right; ZIDN colored green interacting with the same H2A.Z–H2B dimer than the rest of the YL1–ZID domain). The model was created by modeling the ZIDN onto the H2A.Z–H2B dimer recognized by the rest of the YL1–ZID and using the mode of binding observed when it binds to a symmetry-related H2A.Z–H2B dimer. The model was then refined to obtain correct stereochemistry. This procedure showed that only the position of YL1–ZID residues E24, E25 and E26 (colored blue), which are part of the hinge region between YL1–ZID αN1 and αN2 helices, need to be moved to adapt to the new ZIDN conformation.
c) Sequence of yeast SWC2–ZID from alignment in figure 2b. Stars below the alignment indicate the residues mutated in either mutant m1 (dark blue) or mutant m2 (light blue). Mutant SWC2-ΔN was made by deletion of the N-terminal region of SWC2 (deletion of amino acids 1–45), including the αN1 helix. Mutant SWC2-m1 (I63A L65A L66A F67A) targeted the H2A.Z αC binding region of YL1 and mutant SWC2-m2 (D73A D75A F76A) targeted the first DNA-binding region of SWC2.
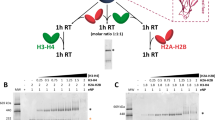
d) His-pulldowns with yeast His–SWC2–ZID and HTZ1/HTB1 complexes using WT HTZ1 and the indicated SWC2 mutants. The interaction between SWC2 and HTZ1–HTB1 is strongly affected by the m1 and m2 mutations but not by deletion of the N-terminal region (ΔN).
e) GST-pulldowns with human GST–YL1–ZID (1–69) or GST–YL1–ZID–ΔN (25–69) and human H2A.Z–H2B or yeast HTZ1–HTB1 dimers. The interaction between human YL1 and histones is dependent of its N-terminal region (ZIDN) for proper binding to human H2A.Z–H2B but not to yeast HTZ1–HTB1.‘*’, degradation products.
f) His-pulldowns with yeast His–SWC2–ZID (1–101) or His–SWC2–ZID–ΔN (46–101) and human H2A.Z–H2B or yeast HTZ1–HTB1 dimers. The interaction between yeast SWC2 and histones is dependent of its N-terminal region (ZIDN) for proper binding to human H2A.Z–H2B but not to yeast HTZ1–HTB1.
Supplementary Figure 6 ITC data.
ITC profiles of the titration of the human YL1–ZID (YL1 residues 1–69) by human H2A.Z–H2B and H2A–H2B pairs. The name of the proteins and mutants used are indicated underneath the data. For all experiments: top panel: raw titration data of YL1 (1–69) injected into H2A(.Z)–H2B. Bottom panel: integrated heat measurements for the titration of YL1 with H2A(.Z)–H2B.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Table 1 (PDF 3471 kb)
Supplementary Data Set 1
Complete gels (PDF 5193 kb)
Rights and permissions
About this article
Cite this article
Latrick, C., Marek, M., Ouararhni, K. et al. Molecular basis and specificity of H2A.Z–H2B recognition and deposition by the histone chaperone YL1. Nat Struct Mol Biol 23, 309–316 (2016). https://doi.org/10.1038/nsmb.3189
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3189
This article is cited by
-
Mammalian PERIOD2 regulates H2A.Z incorporation in chromatin to orchestrate circadian negative feedback
Nature Structural & Molecular Biology (2022)
-
Tip60 activates Hoxa9 and Meis1 expression through acetylation of H2A.Z, promoting MLL-AF10 and MLL-ENL acute myeloid leukemia
Leukemia (2021)
-
Znhit1 controls intestinal stem cell maintenance by regulating H2A.Z incorporation
Nature Communications (2019)
-
MacroH2A1 chromatin specification requires its docking domain and acetylation of H2B lysine 20
Nature Communications (2018)
-
Histone chaperone networks shaping chromatin function
Nature Reviews Molecular Cell Biology (2017)