Abstract
Multiple protein ubiquitination events at DNA double-strand breaks (DSBs) regulate damage recognition, signaling and repair. It has remained poorly understood how the repair process of DSBs is coordinated with the apoptotic response. Here, we identified the E4 ubiquitin ligase UFD-2 as a mediator of DNA-damage-induced apoptosis in a genetic screen in Caenorhabditis elegans. We found that, after initiation of homologous recombination by RAD-51, UFD-2 forms foci that contain substrate-processivity factors including the ubiquitin-selective segregase CDC-48 (p97), the deubiquitination enzyme ATX-3 (Ataxin-3) and the proteasome. In the absence of UFD-2, RAD-51 foci persist, and DNA damage-induced apoptosis is prevented. In contrast, UFD-2 foci are retained until recombination intermediates are removed by the Holliday-junction-processing enzymes GEN-1, MUS-81 or XPF-1. Formation of UFD-2 foci also requires proapoptotic CEP-1 (p53) signaling. Our findings establish a central role of UFD-2 in the coordination between the DNA-repair process and the apoptotic response.
Similar content being viewed by others
Main
DSBs are highly cytotoxic and require the assembly of DNA-damage signaling complexes and the DSB-repair machinery at DNA breaks1. In the C. elegans germ line, DSBs are repaired primarily by homologous recombination (HR)2. After initial processing of the damaged site, RAD-51 accumulates on single-stranded DNA (ssDNA) overhangs and mediates strand invasion into the undamaged template, thus facilitating recombination and repair. Ultimately, cruciform recombination intermediates called Holliday junctions (HJ) are formed3. HJs are processed by two major pathways: (i) HJ dissolution via the combined action of the Bloom's syndrome helicase and the topoisomerase TopoIIIα (ref. 4) or (ii) HJ resolution by nucleases acting as resolving enzymes5. Although HJ dissolution predominates in most systems6,7, in C. elegans the GEN-1 resolvase is needed for completion of HR repair of DSBs8. The resolution of HR intermediates is important for the apoptotic response to DSBs, because GEN-1 and HJ-processing factors are required for DNA-damage-induced programmed cell death. Although the mechanisms for such regulation are not yet known, the C-terminal noncatalytic domain of GEN-1 appears to be important for DNA-damage signaling8,9. The apoptotic response to persistent DSBs facilitates the removal of germ cells in C. elegans when DSBs or meiotic-recombination intermediates are not repaired, and it occurs in the meiotic pachytene zone of the nematode germ line10. DNA-damage-checkpoint signaling leads to the activation of the C. elegans p53 homolog CEP-1 and the subsequent induction of apoptosis11,12. CEP-1 (p53) becomes available in the late pachytene region of the germ line, thus leading to the apoptosis competency of these germ cells. CEP-1 expression in earlier stages of meiosis is translationally repressed by the conserved mRNA-binding protein GLD-1 (ref. 13). Thus, apoptosis is initiated only when aberrant meiotic-recombination intermediates or ionizing radiation (IR)-induced DSBs persist in late pachytene cells. However, it remains unclear how DNA-damage processing by recombination repair is coordinated with the apoptosis pathway, thereby allowing sufficient time to resolve HR intermediates.
To better understand how the apoptotic response to DSBs is regulated, we performed a genetic screen in C. elegans for defects in IR-induced germ-cell apoptosis. RNA interference (RNAi) knockdown and genetic mutation of ufd-2 resulted in a decreased apoptotic response. We found that after initiation of HR by the recombinase RAD-51, UFD-2 formed foci that we defined as ubiquitination hubs because they also contained substrate CDC-48, ATX-3 and proteasomes. In the absence of UFD-2 or its catalytic activity, RAD-51 foci persisted. Similarly to ufd-2 deficiency, elevated RAD-51 levels resulted in reduced apoptosis. When the resolution of HJs was hampered by the absence of GEN-1, MUS-81 or XPF-1, UFD-2 foci persisted. Formation of UFD-2 foci required not only RAD-51 but also proapoptotic signaling through CEP-1 (p53). We therefore propose that UFD-2-specific ubiquitination hubs link proapoptotic and DNA-repair signaling, thereby coordinating the apoptotic response with ongoing DSB-repair activity.
Results
Ligase activity of UFD-2 triggers DSB-induced apoptosis
To identify new regulators of the apoptotic response to DNA damage, we performed an RNAi screen targeting 770 genes whose transcription is enriched in the C. elegans germ line14 (Fig. 1a). We focused on those candidate genes because in C. elegans DNA-damage-induced apoptosis occurs only in germ cells10,15. We identified the E4 ubiquitin ligase UFD-2 as the most prominent hit resulting from our screen. RNAi against ufd-2 led to a dose-dependent decrease in IR-induced apoptosis (Fig. 1b), a phenotype confirmed by analysis of the two different null alleles ufd-2(tm1380) and ufd-2(hh1) (Fig. 1c,d). In contrast, neither developmental apoptosis, which occurs during the somatic development of the worm, nor physiological germ-cell apoptosis, a background germ-cell apoptosis that occurs independently of DNA damage, was defective in ufd-2 mutants (Supplementary Fig. 1a,b).
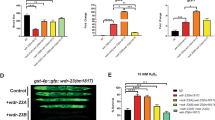
(a) Schematic illustration of the RNAi screen for identification of DNA-damage-induced apoptosis mediators. After RNAi treatment, worms were subjected to IR and scored for apoptotic corpses (indicated by filled arrowheads) 24 h later by differential interference contrast (DIC) microscopy. (b) Apoptotic corpses in worms treated with the indicated RNAi constructs and exposed to different IR doses 24 h after treatment. Data represent mean ± s.e.m., n = 2–11 animals (exact values in Supplementary Table 1); data from 3 independent experiments. (c) Representative images from 3 independent experiments showing late-pachytene cells of the C. elegans germ line 24 h after IR treatment (0 or 60 Gy). Arrowheads indicate apoptotic corpses. Scale bar, 5 μm. WT, wild type. (d) Analysis of DNA-damage-induced apoptosis 24 h after IR treatment (0, 30 or 60 Gy) of the indicated genotypes. Center lines, medians; box limits, 25th and 75th percentiles, as determined by R software; whiskers, 1.5 times the interquartile range (IQR) from the 25th and 75th percentiles; outliers, dots. The notches are defined as ±1.58× IQR/ and represent the 95% confidence interval for each median. Nonoverlapping notches indicate approximately 95% confidence that two medians differ; n = 69–80 animals (exact values in Supplementary Table 1); data from 5 independent experiments. (e) Immunoblot showing autoubiquitination of UFD-2 with UFD-2 (wild type) or UFD-2P951A ubiquitin ligases. Image is representative of results from 3 independent experiments. (f) Analysis of DNA-damage-induced apoptosis 24 h after IR treatment (0 or 60 Gy) of the indicated genotypes. Statistics are as in d; n = 36–63 animals (exact values in Supplementary Table 1); data from 3 independent experiments.
and represent the 95% confidence interval for each median. Nonoverlapping notches indicate approximately 95% confidence that two medians differ; n = 69–80 animals (exact values in Supplementary Table 1); data from 5 independent experiments. (e) Immunoblot showing autoubiquitination of UFD-2 with UFD-2 (wild type) or UFD-2P951A ubiquitin ligases. Image is representative of results from 3 independent experiments. (f) Analysis of DNA-damage-induced apoptosis 24 h after IR treatment (0 or 60 Gy) of the indicated genotypes. Statistics are as in d; n = 36–63 animals (exact values in Supplementary Table 1); data from 3 independent experiments.
UFD-2 participates in the ubiquitin fusion degradation (UFD) pathway, which was first identified in budding yeast16. Substrate ubiquitination involves E1 ubiquitin-activating enzymes, E2 ubiquitin-conjugating enzymes and E3 ubiquitin ligase enzymes. UFD-2 defines a class of so-called E4 enzymes, which further elongate preexisting ubiquitin chains and facilitate efficient proteasomal degradation17,18,19,20. UFD-2 preferentially targets lysine residues 29 and 48 of ubiquitin for autoubiquitination (Supplementary Fig. 1e). A P951A point mutation in the U-box domain completely blocks the ligase activity of UFD-2 (ref. 21) (Fig. 1e). To determine whether UFD-2 catalytic activity is required for DNA-damage-induced apoptosis, we transgenically expressed UFD-2::GFP or UFD-2P951A::GFP in the germ line of the wild-type or the ufd-2-deletion background. Importantly, UFD-2::GFP expression fully restored the apoptotic DNA-damage response in ufd-2(tm1380) mutant animals (Fig. 1f). In contrast, the catalytically dead mutant UFD-2P951A::GFP showed strongly reduced apoptosis after treatment with 60 Gy IR, to a level comparable to that of the ufd-2-deletion mutant. Overexpression of UFD-2P951A::GFP in the wild-type background also caused defective apoptosis, thus indicating that the inactive U-box mutant has a dominant-negative response to DNA damage (Fig. 1f).
UFD-2 forms focal accumulations after DSB induction
To determine in vivo localization, we raised polyclonal antibodies that specifically recognize UFD-2 and performed western blot analysis and immunofluorescence staining (Fig. 2a and Supplementary Fig. 2a). Using immunostaining, we found that under unperturbed conditions the protein was evenly distributed in the C. elegans germline syncytium (Supplementary Fig. 2b). Starting in the late pachytene, UFD-2 accumulated at the nuclear periphery, thus resulting in a ring-shaped staining pattern. After IR treatment, UFD-2 foci of varying size and number became detectable within the nucleoli (Fig. 2a,b and Supplementary Fig. 2b). We confirmed the pattern of antibody staining by GFP-tagged UFD-2 transgenes (Fig. 2c,d). These UFD-2 foci were present in the mitotic zone (data not shown) as well as in the mid-to-late pachytene zone of the germ line after IR (Supplementary Fig. 2b).
(a) Representative images (from 3 independent experiments) showing germ lines of worms of the indicated genotypes, stained with anti-UFD-2 antibody and nuclear stain (DAPI) 24 h after IR treatment (60 Gy). Filled arrowhead indicates nuclei with UFD-2 foci. Scale bar, 5 μm. (b) Quantification of UFD-2 foci in pachytene regions of germ lines, treated as in a. Data show means ± s.e.m. of n = 231 animals (WT, 0 Gy) and n = 280 animals (WT, 60 Gy), from 12 independent experiments. (c) Representative images (from 3 independent experiments) showing germ lines of worms of the indicated genotypes, stained with GFP-booster and DAPI 24 h after IR treatment (60 Gy). Filled arrowheads indicate nuclei with UFD-2 foci. Scale bar, 5 μm. (d) Quantification of UFD-2 foci in pachytene regions of germ lines, treated as in c. Data shown are means ± s.e.m., n = 34–51 animals (exact values in Supplementary Table 1); data from 3 independent experiments.
Given our interest in apoptosis, we focused on the formation of UFD-2 foci in the pachytene region. Pachytene cells elicit DNA-damage-induced apoptosis after DNA-damage-checkpoint activation, whereas mitotic nuclei in the distal germline compartment are subjected to cell-cycle arrest10. In contrast to the IR-induced apoptosis defect, we found that cell-cycle arrest was normally induced in ufd-2-mutant animals (this observation resulted from scoring the number of mitotic nuclei that were enlarged because of continuous growth of cellular and nuclear compartments in the absence of cell division; Supplementary Fig. 1c,d)10,22, thus suggesting that the DNA-damage checkpoint was functional in general. Unlike IR-induced RAD-51 repair foci, which accumulated immediately after damage induction, UFD-2 foci were not yet detectable 5 h after damage (Supplementary Fig. 2c). We therefore scored formation of UFD-2 foci 24 h after IR, a time point concurrent with full apoptosis activation10, by using both antibodies and GFP transgenes. The number of foci observed in pachytene cells increased from 0–5 foci per germ line to more than 15 foci after treatment with 60 Gy of IR (Fig. 2a–d and Supplementary Fig. 2b,c). Unexpectedly, the ubiquitin ligase mutant was equally efficient as the wild-type ligase in forming UFD-2 foci (Fig. 2c,d). Together, these data indicate that UFD-2 ligase activity is required to trigger DNA-damage-induced apoptosis (Fig. 1f), but is not necessary for the formation of UFD-2 foci (Fig. 2c,d).
Ubiquitin signaling fine-tunes the apoptotic response
Given that UFD-2 triggers protein degradation16,17,23, we examined whether factors associated with the ubiquitin–proteasome system (UPS) might associate with UFD-2 foci17,24,25. Hence, we analyzed ubiquitin localization 24 h after irradiation. In fact, an antibody recognizing conjugated mono- and polyubiquitin chains stained UFD-2 foci (Fig. 3a and Supplementary Fig. 3e). Additional staining experiments detected colocalization of the proteasome and the ubiquitin-selective segregase CDC-48 with UFD-2 foci (Fig. 3a). Among other processes, CDC-48 (p97) has been shown to coordinate the degradation of chromatin-associated proteins during DNA replication or DNA repair by extracting ubiquitinated substrate proteins from higher-order complexes26,27,28. Because CDC-48 has been shown to interact with UFD-2 (ref. 20), we asked whether the interaction might be necessary for UFD-2-dependent apoptotic signaling. Transgenic overexpression of UFD-2C448Y::GFP, a mutant that has ligase activity but is not able to interact with CDC-48 (ref. 29), led to the formation of UFD-2 foci even without IR treatment (Supplementary Fig. 3f–j). However, UFD-2C448Y::GFP did not rescue the apoptosis phenotype displayed by ufd-2-deletion worms (Fig. 3g), thus suggesting that, in addition to ligase activity, the interaction with CDC-48 is a prerequisite for the apoptotic function of UFD-2. CDC-48 has been demonstrated to guide ubiquitin-chain topology by coordinating different substrate-processing enzymes, such as UFD-2 and the deubiquitination enzyme ATX-3 (ref. 20). Intriguingly, we also found that ATX-3 localized to UFD-2 foci (Fig. 3a and Supplementary Fig. 3c,d), thus indicating an orchestrated action of UFD-2, ATX-3 and CDC-48 at ubiquitination hubs triggered by DNA damage. The ubiquitination activity of UFD-2 was dispensable for the recruitment of the proteasome, ATX-3 and CDC-48 (Fig. 3b). In contrast, apoptosis induction required the catalytic activity of UFD-2 as well as its interaction with CDC-48 (Figs. 1e,f and 3g).
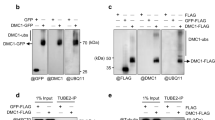
(a,b) Representative images from 3 independent experiments showing ufd-2(tm1380); UFD-2::GFP (a) and ufd-2(tm1380); UFD-2P951A::GFP (b) immunostained with antibodies to the indicated proteins 24 h after IR (Gy 60). The boxed area is shown in 3× zoom. Alpha SU, proteasome 20S alpha subunits. Scale bars, 5 μm. (c) Representative images of 3 independent experiments showing worm germ lines of indicated genotypes immunostained with anti-UFD-2 antibody and DAPI 24 h after IR treatment (60 Gy). Filled and empty arrowheads indicate nuclei positive or negative for UFD-2 foci, respectively. Scale bar, 5 μm. (d) Quantification of UFD-2 foci in pachytene regions of germ lines treated as in c. Data show means ± s.e.m., n = 36–107 animals (exact values in Supplementary Table 1); data from 3 independent experiments. (e–g) Analysis of DNA-damage-induced apoptosis 24 h after IR treatment (0 or 60 Gy) of the indicated genotypes. Statistics are as in Figure 1d; n = 39–52 animals (e), 38–51 animals (f) and 34–74 animals (g) (exact values in Supplementary Table 1); data from 3 independent experiments.
Given that in yeast and humans, Ufd2 and UBE4B, respectively, mediate the elongation of preformed ubiquitin chains, we tested whether UFD-2 cooperates with the E3 ligase HECD-1, the ortholog of budding yeast Ufd4 and human HECTD1 or TRIP12, thereby triggering DNA-damage-induced apoptosis17,30,31,32. Indeed, loss of HECD-1 prevented formation of UFD-2 foci, thus suggesting ubiquitin-dependent recruitment of UFD-2 (Fig. 3c,d). The decreased apoptosis in hecd-1 mutants suggested that UFD-2 focal accumulation may have a role in response to DNA damage (Fig. 3e). The apoptosis defect was even more pronounced in ufd-2 and hecd-1 double mutants, thus indicating that the activity of both enzymes is required to achieve apoptosis (Fig. 3e). In contrast, the deubiquitination enzyme ATX-3 counteracted UFD-2 recruitment, and both formation of UFD-2 foci and apoptosis were increased in atx-3 mutants (Fig. 3c,d,f). Accordingly, the excessive DNA-damage-induced apoptosis in atx-3 mutants was suppressed in ufd-2 and atx-3 double-mutant worms (Fig. 3f). The number of ubiquitin foci per germ line was decreased in hecd-1 mutants, whereas it was increased in atx-3 mutants (Fig. 3c,d and Supplementary Fig. 3k). This observation suggests that the formation of UFD-2 foci is ubiquitin dependent and determined by ubiquitin-mediated recruitment signals fine-tuned by HECD-1 and ATX-3. We therefore conclude that the apoptotic response to DNA damage is coordinated by ubiquitination signals defined by UFD-2 in cooperation with HECD-1 and ATX-3.
UFD-2 supports RAD-51 dissociation from DNA-repair sites
Next we analyzed whether UFD-2 also affects the DNA-repair process in addition to apoptosis. In contrast to DSB induction by IR, UV irradiation did not result in the formation of UFD-2 foci, a result consistent with a specific role of UFD-2 in responding to DSBs (Supplementary Fig. 3a). We also found that, in line with this observation, RPA-1::GFP and BRD-1::GFP HR fusion proteins33,34 accumulated in UFD-2 foci 24 h after IR treatment (Fig. 4a,b). Furthermore, IR of L4-stage ufd-2-mutant larvae resulted in reduced embryonic survival in the subsequent generation (Supplementary Fig. 3b).
(a) Schematic illustration of DNA DSB repair by HR in C. elegans. After DSB induction, RPA binds resected ssDNA, BRD-1 acts together with BRCA-1 at the DSB site, RPA is exchanged for RAD-51, which mediates strand invasion, and Gen-1 resolves the HJ, thus resulting in a repaired DSB. Names in brackets indicate human homologs. (b) Representative images (from 3 independent experiments) showing BRD-1::GFP and RPA-1::GFP germ lines stained with anti-UFD-2 and DAPI 24 h after IR treatment (60 Gy). Scale bar, 5 μm. (c) Representative images (from 3 independent experiments) showing wild-type and ufd-2(tm1380) germ lines isolated 16 h after IR treatment (20 Gy), stained with anti-RAD-51 and DAPI. Filled arrowheads indicate nuclei positive for RAD-51 staining. Scale bar, 10 μm. (d) Quantification of germ cells positive for RAD-51 staining of wild-type and ufd-2(tm1380) worms treated with IR (0 Gy) and isolated after 1 h or treated with IR (20 Gy) and isolated after 1, 7, 16 or 48 h. Data shown are means ± s.e.m. n = 35–43 animals (exact values in Supplementary Table 1); data from 3 independent experiments. ***P ≤ 0.001 by two-tailed Student's t test.
To establish whether ufd-2 promotes the processing of DNA-repair intermediates, we analyzed the kinetics of RAD-51 foci. Whereas both wild-type and ufd-2 mutants accumulated an equal amount of RAD-51 positive nuclei 1 h after IR, twice as many RAD-51-stained nuclei persisted 16 h later in ufd-2 mutants (Fig. 4c,d). This delay in RAD-51 foci dissociation that temporally coincided with formation of UFD-2 foci suggested that UFD-2 contributes to the resolution of repair intermediates.
UFD-2 acts downstream of proapoptotic signaling
We next sought to further investigate the role of the DSB-repair process in formation of UFD-2 foci (Fig. 4a). Impairment of HR in rad-51-deletion-mutant worms blocked formation of UFD-2 foci (Fig. 5b). In contrast, rad-54-deletion mutants, which were defective in removal of RAD-51 from DNA during HR repair35, exhibited an accumulation of UFD-2 foci (Fig. 5b). The nucleases GEN-1, MUS-81 and XPF-1 are required for the resolution of HJs in order to complete the HR repair process of IR-induced DSBs8,36,37. Deletion of the gen-1, mus-81 and/or xpf-1 HJ-processing enzymes also led to focal accumulation of UFD-2 (Fig. 5c and Supplementary Fig. 4a). Notably, mus-81- and xpf-1-mutant animals also showed elevated numbers of UFD-2 foci in the absence of IR-induced DSBs, a result consistent with the function of MUS-81 and XPF-1 in meiotic HJ resolution36,37. These results indicate that HR must commence in order for UFD-2 foci to form, and the foci persist until HR is completed (Fig. 5b,c).
(a) Schematic illustration of the apoptosis pathway in C. elegans. Names in brackets indicate human homologs. (b–d) Quantification of UFD-2 foci in pachytene regions of germ lines of worms of the indicated genotypes, isolated 24 h after irradiation (60 Gy). Data show means ± s.e.m., n = 30–113 animals (b), 34–92 animals (c) and 20–202 animals (d) (exact values in Supplementary Table 1); data from 3 independent experiments.
Because ufd-2-mutant worms displayed decreased apoptosis, we assessed whether apoptotic signaling might be affected in ufd-2-mutant worms. The apoptotic core machinery is conserved between C. elegans and mammals. The p53 homolog CEP-1 induces transcription of the two BH3-only proteins EGL-1 and CED-13 (refs. 13,38), which bind to the only Bcl2-like protein, CED-9. As a consequence, the inhibitory effect of CED-9 on the Apaf1-like CED-4 is alleviated, and CED-4 activates the caspase CED-3, which in turn executes cell death39 (Fig. 5a). In view of the ubiquitin ligase activity, we tested whether CEP-1 protein might accumulate after DNA damage in the absence of UFD-2. However, in wild-type and ufd-2-mutant worms, CEP-1 protein was equally expressed after 60-Gy irradiation (Supplementary Fig. 4b–d). Additional evaluation of mRNA transcripts of the CEP-1-target gene egl-1 showed a comparable transcriptional regulation in both genotypes 4 and 24 h after damage infliction (Supplementary Fig. 4d).
Having established that CEP-1 activation occurs independently of ufd-2, we wondered whether the formation of UFD-2 foci might be dependent on CEP-1. Strikingly, loss of CEP-1 prevented the formation of UFD-2 foci after IR (Fig. 5d), whereas UFD-2 protein expression remained unaffected (Supplementary Fig. 5b). Moreover, a double mutant of the two proapoptotic CEP-1 effectors, egl-1 and ced-13, which has defects in DNA-damage-induced apoptosis that are similar to those of cep-1 mutants40, mimicked the cep-1 defect in formation of UFD-2 foci after DNA damage (Fig. 5d and Supplementary Fig. 5a). To further confirm a direct role of CEP-1 in the formation of UFD-2 foci, we enhanced CEP-1 activity by using a gld-1(op236) mutation, which has previously been shown to increase CEP-1 levels13. The gld-1 mutants indeed displayed strongly elevated UFD-2 foci, thus supporting the idea that CEP-1 promotes UFD-2 focal accumulation. The cep-1 and gld-1 double mutant and wild-type germ cells displayed a similar number of UFD-2 foci (Fig. 5d and Supplementary Fig. 5a). One potential explanation for the failure of cep-1 to completely suppress foci formation in gld-1 might be the numerous additional target mRNAs of GLD-1 (refs. 41,42). Notably, the failure of cep-1 to initiate apoptosis does not affect repair activity, because IR-induced embryonic lethality has previously been shown to remain unaffected13. Furthermore, we found that the disassembly of RAD-51 foci 16 h after damage induction was as efficient in cep-1 and gld-1 mutants as in wild type (Supplementary Fig. 5c). In contrast to the loss of CEP-1 signaling, formation of UFD-2 foci was unaltered in apoptosis-deficient ced-3- and ced-4-mutant worms (Fig. 5d), thus emphasizing the necessity of CEP-1 activity, rather than the apoptotic process in general, in the formation of UFD-2 foci. These results suggest that UFD-2 acts downstream of the proapoptotic signaling cascade.
Resolution of RAD-51 is linked to apoptotic signaling
We next sought to investigate the role of UFD-2 in the removal of RAD-51 foci and its effect on apoptosis. Germline-specific expression of UFD-2::GFP in transgenic ufd-2-deletion mutants rescued the delay of RAD-51 removal from DNA (Fig. 6a). Increased RAD-51 retention occurred in worms moderately overexpressing RAD-51::GFP after 24 h of IR, compared with wild-type worms (Fig. 6a). Importantly, the retention of RAD-51 filaments either by loss of ufd-2 or by RAD-51 overexpression strictly correlated with decreased apoptosis levels (Fig. 6b). Despite the elevated RAD-51 protein levels, the GFP transgenic line possessed a normal repair capacity, as assessed on the basis of embryonic survival after IR, thus suggesting that the decreased apoptosis is not related to enhanced removal of DSBs (Supplementary Fig. 6a). In contrast, the atx-3 mutant, which displayed increased UFD-2 foci and apoptosis after DNA damage, showed decreased RAD-51 retention 16 h after IR (Supplementary Fig. 6b). To test whether elevated RAD-51 levels might directly account for the decreased apoptosis observed in ufd-2 mutants or after overexpression of RAD-51, we depleted RAD-51 by RNAi knockdown. Indeed, rad-51 RNAi in ufd-2-mutant or RAD-51::GFP-expressing worms reverted the apoptosis defect after IR treatment (Fig. 6c). Importantly, rad-51 RNAi also resulted in decreased embryonic survival after IR in wild-type and rad-51-mutant worms (Supplementary Fig. 6c). We further validated the role of RAD-51 filaments in suppressing the apoptotic response by inhibition of RAD-51-filament formation with the RAD-51 inhibitor B02 (ref. 43). Similarly to the effects of decreased RAD-51 levels, treatment with B02 reverted the apoptosis phenotype of ufd-2-deletion mutants or the RAD-51-overexpression line (Fig. 6d), thus suggesting that RAD-51 accumulation directly antagonizes apoptotic signaling. Moreover, rad-51-heterozygous mutants with decreased RAD-51 levels showed reversion of the apoptosis defect of ufd-2 mutants (Fig. 6e). In summary, these observations support the idea that UFD-2 contributes to the resolution of DNA-repair sites and that retention of RAD-51 filaments leads to inhibition of apoptosis (Fig. 6f).
(a) Quantification of positive RAD-51 staining in germ cells of worms of the indicated genotypes treated with IR (0 or 20 Gy) and isolated after 24 h. Data show means ± s.e.m., n = 34–63 animals (exact values in Supplementary Table 1); data from 3 independent experiments. ***P ≤ 0.001 by two-tailed Student's t test. (b) Analysis of DNA-damage-induced apoptosis 24 h after IR treatment (0, 60 Gy) of worms of the indicated genotypes. Statistics are as in Figure 1d. n = 61–82 animals (exact values in Supplementary Table 1); data from 3 independent experiments. (c–e) Analysis of DNA-damage-induced apoptosis 24 h after IR treatment (0, 60 Gy) of ufd-2 and RAD-51::GFP worms treated with rad-51 RNAi or control RNAi (c), wild-type, ufd-2 and RAD-51::GFP worms treated with RAD51 inhibitor B02 (200 mM) (d) or the indicated genotypes (e). Statistics are as in Figure 1d; n = 30–47 animals (c), 28–69 animals (d) and 29–40 animals (e) (exact values in Supplementary Table 1); data from 3 independent experiments. (f) Model for the coordination between HR and apoptotic signaling by UFD-2. DSB repair triggers RAD-51 accumulation at ssDNA, thus facilitating homology pairing. After efficient strand invasion, RAD-51 is removed, and HJs are resolved by HJ-processing enzymes (GEN-1, MUS-81 and XPF-1). UFD-2 supports RAD-51 dissociation from DSBs at advanced time points. Ongoing repair is reflected by the presence of UFD-2-containing hubs late after IR. These ubiquitination hubs contain processivity factors such as CDC-48 and proteasomes (not shown). Interaction between UFD-2 and CDC-48 is necessary to transduce a proapoptotic signal. UFD-2 hub formation is fine-tuned by the E3 ligase HECD-1, the DUB ATX-3 and proapoptotic CEP-1 (p53) signaling. Het, heterozygous.
Discussion
In this study, we uncovered a ubiquitin-dependent process that facilitates the communication between DNA repair and the apoptotic response. We identified the E4 ubiquitin ligase UFD-2 as a central regulator of the spatiotemporal coordination of both processes. Our data suggest that defects preventing the timely progression of HR either by failure to resolve HJs, as previously demonstrated8,9, or by aberrant retention of RAD-51 foci at the chromatin, caused by loss of UFD-2, as shown here, halt the apoptotic response. In contrast, RAD-51-filament assembly and proapoptotic signaling by the p53 tumor-suppressor homolog CEP-1 are both required for the formation of UFD-2-specific hubs, which we defined on the basis of the presence of proteolytic factors of the UPS machinery (Fig. 6f). We propose that these degradation hubs calibrate the progression of the DNA-repair machinery with apoptotic activity via modulation of ubiquitin signaling. Such a calibration might provide additional time for ongoing HR repair when CEP-1-dependent apoptotic signaling has already been triggered. Indeed, CEP-1 activity can be detected within the first hour after IR treatment40, whereas the rapidly formed RAD-51 foci are turned over in the course of 24 h. To prevent the premature demise of cells that are engaged in the process of repairing DSBs, the simultaneous presence of proapoptotic signaling and ongoing HR requires coordination, which we propose is orchestrated at the HR-repair sites through the UFD-2 ubiquitination hubs, which might provide regulatory feedback to the apoptotic signaling, depending on the status of the damage removal. The fine-tuning of ubiquitin-chain topology by the concerted action of UFD-2, the E3 ligase HECD-1 and the hydrolase ATX-3 at HR sites might constitute a versatile signaling tool that enables communication between the apoptotic response and DNA damage (Fig. 6f). Because the E3 ligase HECD-1 is required for UFD-2-hub formation and for apoptosis, we propose that E4 activity17,31,44 provides an additional layer of regulation by modifying ubiquitin-chain topology.
The human E4 homolog UBE4B cooperates similarly with the HECT domain E3 ligase TRIP12 in substrate ubiquitination, thus suggesting the existence of a conserved signaling pathway30. In support of this possibility, TRIP12 fine-tunes ubiquitin-controlled events at DSBs45, and recent reports have linked UBE4B to different cancer types, thereby highlighting the relevance of ubiquitin signaling in the decision between DNA-damage and apoptosis responses46,47,48,49. Disassembly of RAD-51 filaments might involve the ubiquitin-selective segregase CDC-48 (p97), which has recently been implicated in the degradation of chromatin-associated proteins26,27. Moreover, Cdc48 has been shown to limit RAD51 occupancy on DNA50. In agreement with this notion, CDC-48 binding is required for UFD-2 to trigger DNA-damage-induced cell death (Fig. 3g). Defects in DNA repair and apoptosis are particularly relevant in tumor formation. Thus, understanding the conserved role of UFD-2 and UBE4B in response to IR-induced DNA damage might provide new therapeutic directions for drug development and cancer treatment.
Methods
C. elegans strains.
C. elegans strains were cultured at 20 °C on nematode growth medium (NGM) and fed with Escherichia coli strain OP50 according to standard procedures51. The Bristol strain N2 was used as the wild type. Mutants and transgenic animals used in this study were as follows: mus-81(tm1937) I, rad-54&snx-3(ok615) I/hT2 [bli-4(e937) let-?(q782) qIs48] (I;III), cep-1(lg12501)I, ced-1(e1735)I, gld-1(op236)I, ufd-2(tm1380)II, ufd-2(hh1)II, xpf-1(tm2842) II, gen-1(tm2940)III, ced-4(n1162) III, hecd-1(tm2371)IV, rad-51(ok2218) IV/nT1[qIs51](IV;V), ced-3(n717) IV, atx-3(gk193)V, egl-1(n1084n3082)V; ced-13(tm536)X,, Is[rad-51::GFP:3xFLAG], gla-3(op216)I, hus-1(op241)I, unc-119(ed3)III; gtIs[unc-119(+), Ppie-1::GFP::rpa-1::pie-1-3′UTR], hhIs121[unc-119(+), Pmex-5::ufd-2::GFP::tbb-2 3′UTR], hhIs135[unc-119(+), Pmex-5 (w/o ATG)::ufd-2 (w/o TAA, P951A)::(Gly)5Ala::gfp F64LS65T(w introns/stop)::tbb-2 3′UTR], hhIs134[unc-119(+), Pmex-5::ufd-2 (C448Y)::GFP::tbb-2 3′UTR].
The transgenic lines hhIs121, hhIs134 and hhIs135 were generated for this study. Briefly, the fosmid WRM0621dE05 was used as a template to obtain the genomic sequence of ufd-2 that was cloned together with ppJA252, pJA257 into pCG150 containing the unc-119(+) marker for selection of transgenic worms52. ufd-2 was modified by site-directed mutagenesis to create ufd-2P951A or ufd-2C448Y. The constructs were bombarded into unc-119(ed4)III mutants as described previously53.
No statistical method was used to predetermine sample size. The experiments were not randomized and were not performed with blinding to the conditions of the experiments. All n values are specified in Supplementary Table 1 (for data in Figs. 1,2,3,4,5,6) and Supplementary Table 2 (for data in Supplementary Figs. 1,2,3,4,5,6).
Ionizing radiation.
Synchronized hermaphrodites were grown until the L4 stage and irradiated with the corresponding dose (radiation source: 120-kV X-rays (25 mA; 0.5-mm Alu filter; ISOVOLT 160 M1/10-55, GE Sensing & Inspection Technologies or Biobeam 8000, with Cs137 as the radiation source).
RNAi treatment.
RNA interference was performed with the feeding method54. Three P0 worms were placed on IPTG- and ampicillin-containing NGM plates seeded with E. coli HT115(DE3) expressing double-stranded RNA (dsRNA) and incubated at 15 °C for 72 h. Three single F1 worms were each transferred to a new freshly seeded plate and allowed to lay eggs for approximately 20 h. F1 worms were removed, and F2 worms were allowed to grow until the L4 stage, treated with ionizing radiation and analyzed for radiation-induced apoptosis. Clones in RNAi feeding vectors were provided by M. Vidal (Dana Farber Cancer Center).
Apoptotic corpses.
For physiological apoptosis analysis, synchronized L1 larvae were grown until the L4 stage. Apoptotic corpses were scored 24 h later. Worms were mounted on 3% agar pads, paralyzed with 60 nM NaN3 and analyzed via DIC microscopy55. For DNA-damage-induced apoptosis, worms were subjected to IR at the L4 stage before apoptosis was evaluated 24 h later. Developmental apoptosis was assessed in L1 larvae. Therefore, worms were grown until day one of adulthood. 100 worms were transferred to an NGM agar plate without E. coli, allowed to lay eggs and removed after 1 h. Freshly hatched L1 larvae were scored for apoptotic corpses56.
UFD-2 foci.
Synchronized worms were grown until the L4 larval stage and irradiated with 0 or 60 Gy. 24 h later, germ lines were isolated and immunostained. Numbers of UFD-2 foci were scored in all focal planes in pachytene germ cells. One germ line per worm was scored.
Protein expression and purification.
cDNAs encoding ufd-2b, ufd-2bC448Y and ufd-2bP951A were cloned into the pET-21d expression vector (Novagen) and pGex4T1 (GE Healthcare). Recombinant proteins were expressed in E. coli strain BL21 Codon Plus (Novagen) and purified with an ÄKTA purifier system (GE Healthcare).
Antibody production.
Histidine-tagged purified proteins (UFD-2 and ATX-3 (ref. 20)) were used for immunization of rabbits, and antisera were affinity purified with the respective GST-tagged recombinant proteins (BioGenes). Antibody validation is shown in Supplementary Figures 2a and 3d, respectively.
Preparation of worm lysates.
Synchronized L1 larvae were grown on NGM agar plates with OP50 bacteria until they reached adulthood. Worm lysates used for SDS–PAGE were prepared either from a distinct number of worms (n = 150) or by washing worms from NGM agar plates and then performing multiple washing steps with M9 buffer (3 g/l KH2PO4, 6 g/l Na2HPO4, 5 g/l NaCl and 1 mM Mg SO4 (added after sterilization)), until bacteria were removed. The samples were heated to 95 °C for 5 min and subsequently shock-frozen in liquid nitrogen. After being thawed, samples were subjected to sonication (two times for 15 s, on ice; 50% power; Sonopuls UW 2200, Bandelin), mixed with 4× SDS sample buffer and centrifuged at 15,000 r.p.m. for 10 min.
Immunotechniques.
Immunostaining of isolated germ lines was performed according to the 'freeze-crack' protocol. Worms were dissected onto polylysine-coated slides (Thermo Scientific) in 60 nM NaN3 to isolate germ lines and fixed in fixation buffer (3.7% formaldehyde, 0.2% Tween 20) for 10 min with subsequent shock freezing in liquid nitrogen. This was followed by incubation in a 1:1 mixture of methanol and acetone at –20 °C for 10 min. Germ lines were permeabilized three times in 1% PBS–Triton X-100 for 20 min, washed in 0.1% PBS-Tween 20 (PBS-T) for 10 min and blocked in 10% goat serum in 0.1% PBS-T. A specific staining protocol was followed for GFP-expressing lines, avoiding freezing. Isolated germ lines were fixed with fixation buffer for 10 min in PCR tubes, then immediately permeabilized and blocked as described above. Germ lines were incubated with primary antibody overnight at 4 °C (anti-UFD-2, described above, 1:3,000; anti-CDC-48, 1:12,000 (ref. 57), anti-RAD-51 1:350 (14B4, cat. no. NB100-148, Novus Biologicals58), anti-FK2-ubiquitin 1:100 (AB_612093, cat. no. 04-263, Millipore; validation on manufacturer's website), anti-proteasome 20S alpha 1+2+3+5+6+7, 1:300 (MCP231, cat. no. ab22674, Abcam59) and anti-ATX-3, described above, 1:700). Incubation with the fluorescently labeled secondary antibodies (cat. nos. A-11037 and R37117, Life Technologies, 1:200) or GFP-booster (cat. no. GBA-488, ChromoTek, 1:400 (ref. 60)) was carried out at room temperature for 1 h. Germ lines were mounted in DAPI Fluoromount-G medium (SouthernBiotech). For western blotting, worm lysates were separated by SDS–PAGE) and transferred to nitrocellulose membranes (Whatman, Protran). Membranes were blocked in 1× Roti-Block (Roth) and incubated with the primary antibodies overnight at 4 °C in Roti-Block (Roth; anti-ATX-3, 1:10,000; anti-CDC-48, 1:50,000 (ref. 57); anti-UFD-2, 1:20,000; anti-CEP, 1:15,000 (ref. 13) and anti-α-tubulin 1:5,000 (clone DM1A, Sigma-Aldrich61). Incubation with fluorescently labeled secondary antibodies (Li-Cor IRDye 680, cat. nos. 926-32222 and 926-32223, Li-Cor IRDye 800, cat. nos. 926-32212 and 926-32213, 1:10,000) was carried out at room temperature, before detection of signals with a Li-Cor Odyssey scanner. Quantification of signal intensities was performed with the Odyssey V4.0 software (Li-Cor). The uncropped versions of western blots that have been used to assemble the main figures are shown in Supplementary Figure 7.
Immunoprecipitation in vivo and in vitro.
Worm lysates were prepared as described above, and the protein concentration was determined by measuring absorption at 260 nm with a NanoDrop 800 UV/vis spectrometer. 250 μl of Dynabeads (Invitrogen) was used per reaction, washed twice with conjugation buffer (100 mM sodium phosphate and 0.15 M NaCl) and resuspended in 1 ml 5 mM cross-linking reagent BS3 (Thermo Scientific). Cross-linking was performed on a rotation wheel for 30 min at room temperature. The reaction was stopped by addition of 50 μl quenching buffer (1 M Tris-HCL, pH 7.5), then incubated 15 min at room temperature. The beads were washed three times with 0.5× PBS before 50 μg anti-UFD-2 antibody was added and incubated for 20 min under constant rotation at room temperature. After repeated washing with 0.5× PBS, 2.5 mg of corresponding worm protein lysate was added to the beads and incubated overnight at 4 °C with rotation. For in vitro immunoprecipitation, antigen (UFD-2) was incubated with antibody-coupled beads for 4 h before addition of the putative binding partner (CDC-48) for an additional incubation for 4 h. Both proteins were added in an equimolar ratio (1 mM). Elution was performed as described in the manufacturer's manual.
Microscopy and image acquisition.
Immunostained germ lines were imaged with an AxioImager.M1/Z1 microscope with Apoptome equipped with an AxioCam MRm camera (Carl Zeiss). To allow direct comparison of signal intensities, images were recorded under identical conditions. Processing of selected pictures was carried out in ZEN2011 and ImageJ.
In vitro ubiquitination assay.
UFD-2b::His, UFD-2bC448Y::GST and UFD-2bP951A::His fusion proteins were expressed in E. coli strain BL21-AI and lysed in buffer A (50 mM Tris, pH 7.5, 250 mM NaCl, 5 mM DTT, 1% Triton X-100, 2 mM PMSF and protease-inhibitor mix (Roche)). 10 μg of the aforementioned bacterial lysate was mixed with E1 (25 ng), E2 (Let-70; 400 ng), 2 μg of FLAG::ubiquitin, energy-regenerating solution (Boston Biochemicals) and ubiquitin-conjugation reaction buffer (Enzo Life Sciences). Samples were incubated at 30 °C for 1.5 h, terminated by boiling for 5 min with SDS sample buffer, resolved by SDS–PAGE and immunoblotted with anti-UFD-2 antibodies to monitor ubiquitination of UFD-2.
Persistence of RAD-51 foci after IR.
Synchronized worms were grown until the L4 larvae stage and irradiated with 0 and 20 Gy. 1 to 48 h later, germ lines were isolated and immunostained. Z-stacks were taken of late-pachytene cells of the germ line. Two focal planes covering the upper and lower parts of the germ line were subjected to analysis by scoring each plane for RAD-51-positive cells in the last 25 nuclei of pachytene germ cells before they entered diakinesis.
RNA isolation and real-time PCR.
Total RNA was isolated with TRIzol (Invitrogen) and a Qiagen RNeasy kit. Briefly, worms were washed off the plates with M9 buffer (3 g/l KH2PO4, 6 g/l Na2HPO4, 5 g/l NaCl and 1 mM Mg SO4 (added after sterilization)) and 600 μl TRIzol. Silica beads (1 mm diameter) were added to the samples and homogenized with a Precellys tissue homogenizer. Chloroform was added, and samples were vortexed vigorously before phase separation through centrifugation. The aqueous phase was transferred to Qiagen RNeasy Mini spin columns, and RNA was isolated according to the manufacturer's instructions. cDNA was synthesized with 200 ng total RNA with a High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems). Gene expression levels were determined by real-time PCR with Brilliant III Ultra-Fast SYBR Green QPCR Master Mix (Agilent Technologies) and Bio-Rad CFX96 Real-Time PCR Detection System. Relative gene expression was normalized to tbg-1 (F58A4.8) mRNA levels. In the experiment, three biological and three technical replicate samples were analyzed. The primer sequences used in the RT–PCR reactions were:
tbg-1 forward, 5′-gtacactccactgatctctgctgacaag-3′
tbg-1 reverse, 5′-CTCTGTACAAGAGGCAAACAGCCATG-3′62
egl-1 forward, 5′-TACTCCTCGTCTCAGGACTT-3′
egl-1 reverse, 5′- CATCGAAGTCATCGCACAT-3′
Embryonic survival.
To determine the radiation sensitivity, L4-stage hermaphrodites were irradiated with a single dose of IR, as indicated. After 12 h, worms were transferred to fresh plates (three worms per plate, five plates in total) and allowed to lay eggs for 5 h. After this period, adults were removed, and 24 h later the number of hatched and unhatched embryos was scored (number of hatched larvae normalized to results after mock treatment). As a control for embryonic survival, a heterozygous deletion mutant lacking rad-51 on one chromosome was used.
B02 treatment.
Synchronized L1 larvae were grown in liquid culture in S medium with heat-inactivated (3 × 5 min at 65 °C, with vortexing in between) E. coli strain OP50 (ref. 63), containing 200 mM B02.
Cell-cycle arrest of mitotic germ cells after IR.
Worms were irradiated with 0 or 60 Gy at the late L4 larval stage, as described previously10. 16 h after irradiation, worms were mounted on 3% agar pads and paralyzed with 60 nM NaN3 for DIC microscopy, and the distal region of the germ line was scored for the number of nuclei in all focal planes within a defined area of 2 μm × 6 μm.
Statistical analysis.
Statistical analysis was performed with Excel (Microsoft). Statistical significance was calculated with two-tailed paired Student's t tests. Box plots were generated with BoxPlotR64. Center lines show the medians; box limits indicate the 25th and 75th percentiles, as determined by R software; whiskers extend 1.5× the interquartile range from the 25th and 75th percentiles; and outliers are represented by dots. The notches are defined as ±1.58× IQR/ and represent the 95% confidence interval for each median. Nonoverlapping notches indicate a roughly 95% confidence that that two medians differ.
and represent the 95% confidence interval for each median. Nonoverlapping notches indicate a roughly 95% confidence that that two medians differ.
References
Hoeijmakers, J.H. Genome maintenance mechanisms for preventing cancer. Nature 411, 366–374 (2001).
Clejan, I., Boerckel, J. & Ahmed, S. Developmental modulation of nonhomologous end joining in Caenorhabditis elegans. Genetics 173, 1301–1317 (2006).
Lemmens, B.B. & Tijsterman, M. DNA double-strand break repair in Caenorhabditis elegans. Chromosoma 120, 1–21 (2011).
Bizard, A.H. & Hickson, I.D. The dissolution of double Holliday junctions. Cold Spring Harb. Perspect. Biol. 6, a016477 (2014).
Matos, J. & West, S.C. Holliday junction resolution: regulation in space and time. DNA Repair (Amst.) 19, 176–181 (2014).
Schwartz, E.K. & Heyer, W.D. Processing of joint molecule intermediates by structure-selective endonucleases during homologous recombination in eukaryotes. Chromosoma 120, 109–127 (2011).
West, S.C. et al. Resolution of recombination intermediates: mechanisms and regulation. Cold Spring Harb. Symp. Quant. Biol. 80, 103–109 (2015).
Bailly, A.P. et al. The Caenorhabditis elegans homolog of Gen1/Yen1 resolvases links DNA damage signaling to DNA double-strand break repair. PLoS Genet. 6, e1001025 (2010).
Silva, N., Adamo, A., Santonicola, P., Martinez-Perez, E. & La Volpe, A. Pro-crossover factors regulate damage-dependent apoptosis in the Caenorhabditis elegans germ line. Cell Death Differ. 20, 1209–1218 (2013).
Gartner, A., Milstein, S., Ahmed, S., Hodgkin, J. & Hengartner, M.O. A conserved checkpoint pathway mediates DNA damage--induced apoptosis and cell cycle arrest in C. elegans. Mol. Cell 5, 435–443 (2000).
Schumacher, B., Hofmann, K., Boulton, S. & Gartner, A. The C. elegans homolog of the p53 tumor suppressor is required for DNA damage-induced apoptosis. Curr. Biol. 11, 1722–1727 (2001).
Derry, W.B., Putzke, A.P. & Rothman, J.H. Caenorhabditis elegans p53: role in apoptosis, meiosis, and stress resistance. Science 294, 591–595 (2001).
Schumacher, B. et al. Translational repression of C. elegans p53 by GLD-1 regulates DNA damage-induced apoptosis. Cell 120, 357–368 (2005).
Reinke, V. et al. A global profile of germline gene expression in C. elegans. Mol. Cell 6, 605–616 (2000).
Vermezovic, J., Stergiou, L., Hengartner, M.O. & d'Adda di Fagagna, F. Differential regulation of DNA damage response activation between somatic and germline cells in Caenorhabditis elegans. Cell Death Differ. 19, 1847–1855 (2012).
Johnson, E.S., Ma, P.C.M., Ota, I.M. & Varshavsky, A. A proteolytic pathway that recognizes ubiquitin as a degradation signal. J. Biol. Chem. 270, 17442–17456 (1995).
Koegl, M. et al. A novel ubiquitination factor, E4, is involved in multiubiquitin chain assembly. Cell 96, 635–644 (1999).
Saeki, Y., Tayama, Y., Toh-e, A. & Yokosawa, H. Definitive evidence for Ufd2-catalyzed elongation of the ubiquitin chain through Lys48 linkage. Biochem. Biophys. Res. Commun. 320, 840–845 (2004).
Hoppe, T. Multiubiquitylation by E4 enzymes: 'one size' doesn't fit all. Trends Biochem. Sci. 30, 183–187 (2005).
Kuhlbrodt, K. et al. The Machado-Joseph disease deubiquitylase ATX-3 couples longevity and proteostasis. Nat. Cell Biol. 13, 273–281 (2011).
Okumura, F., Hatakeyama, S., Matsumoto, M., Kamura, T. & Nakayama, K.I. Functional regulation of FEZ1 by the U-box-type ubiquitin ligase E4B contributes to neuritogenesis. J. Biol. Chem. 279, 53533–53543 (2004).
Hodgkin, J., Horvitz, H.R. & Brenner, S. Nondisjunction mutants of the nematode Caenorhabditis elegans. Genetics 91, 67–94 (1979).
Janiesch, P.C. et al. The ubiquitin-selective chaperone CDC-48/p97 links myosin assembly to human myopathy. Nat. Cell Biol. 9, 379–390 (2007).
Rape, M. et al. Mobilization of processed, membrane-tethered SPT23 transcription factor by CDC48(UFD1/NPL4), a ubiquitin-selective chaperone. Cell 107, 667–677 (2001).
Richly, H. et al. A series of ubiquitin binding factors connects CDC48/p97 to substrate multiubiquitylation and proteasomal targeting. Cell 120, 73–84 (2005).
Meerang, M. et al. The ubiquitin-selective segregase VCP/p97 orchestrates the response to DNA double-strand breaks. Nat. Cell Biol. 13, 1376–1382 (2011).
Acs, K. et al. The AAA-ATPase VCP/p97 promotes 53BP1 recruitment by removing L3MBTL1 from DNA double-strand breaks. Nat. Struct. Mol. Biol. 18, 1345–1350 (2011).
Dantuma, N.P. & Hoppe, T. Growing sphere of influence: Cdc48/p97 orchestrates ubiquitin-dependent extraction from chromatin. Trends Cell Biol. 22, 483–491 (2012).
Baek, G.H., Kim, I. & Rao, H. The Cdc48 ATPase modulates the interaction between two proteolytic factors Ufd2 and Rad23. Proc. Natl. Acad. Sci. USA 108, 13558–13563 (2011).
Park, Y., Yoon, S.K. & Yoon, J.-B.B. TRIP12 functions as an E3 ubiquitin ligase of APP-BP1. Biochem. Biophys. Res. Commun. 374, 294–298 (2008).
Liu, G., Rogers, J., Murphy, C.T. & Rongo, C. EGF signalling activates the ubiquitin proteasome system to modulate C. elegans lifespan. EMBO J. 30, 2990–3003 (2011).
Shaye, D.D. & Greenwald, I. OrthoList: a compendium of C. elegans genes with human orthologs. PLoS One 6, e20085 (2011).
Maréchal, A. & Zou, L. RPA-coated single-stranded DNA as a platform for post-translational modifications in the DNA damage response. Cell Res. 25, 9–23 (2015).
Boulton, S.J. et al. BRCA1/BARD1 orthologs required for DNA repair in Caenorhabditis elegans. Curr. Biol. 14, 33–39 (2004).
Solinger, J.A., Kiianitsa, K. & Heyer, W.D. Rad54, a Swi2/Snf2-like recombinational repair protein, disassembles Rad51:dsDNA filaments. Mol. Cell 10, 1175–1188 (2002).
Agostinho, A. et al. Combinatorial regulation of meiotic holliday junction resolution in C. elegans by HIM-6 (BLM) helicase, SLX-4, and the SLX-1, MUS-81 and XPF-1 nucleases. PLoS Genet. 9, e1003591 (2013).
O'Neil, N.J. et al. Joint molecule resolution requires the redundant activities of MUS-81 and XPF-1 during Caenorhabditis elegans meiosis. PLoS Genet. 9, e1003582 (2013).
Hofmann, E.R. et al. Caenorhabditis elegans HUS-1 is a DNA damage checkpoint protein required for genome stability and EGL-1-mediated apoptosis. Curr. Biol. 12, 1908–1918 (2002).
Horvitz, H.R. Genetic control of programmed cell death in the nematode Caenorhabditis elegans. Cancer Res. 59 (Suppl.), 1701s–1706s (1999).
Schumacher, B. et al. C. elegans ced-13 can promote apoptosis and is induced in response to DNA damage. Cell Death Differ. 12, 153–161 (2005).
Lee, M.H. & Schedl, T. Identification of in vivo mRNA targets of GLD-1, a maxi-KH motif containing protein required for C. elegans germ cell development. Genes Dev. 15, 2408–2420 (2001).
Doh, J.H., Jung, Y., Reinke, V. & Lee, M.-H.H. RNA-binding protein GLD-1 recognizes its multiple targets using sequence, context, and structural information to repress translation. Worm 2, e26548 (2013).
Huang, F. & Mazin, A.V. A small molecule inhibitor of human RAD51 potentiates breast cancer cell killing by therapeutic agents in mouse xenografts. PLoS One 9, e100993 (2014).
Park, Y., Yoon, S.K. & Yoon, J.-B.B. The HECT domain of TRIP12 ubiquitinates substrates of the ubiquitin fusion degradation pathway. J. Biol. Chem. 284, 1540–1549 (2009).
Gudjonsson, T. et al. TRIP12 and UBR5 suppress spreading of chromatin ubiquitylation at damaged chromosomes. Cell 150, 697–709 (2012).
Krona, C. et al. Screening for gene mutations in a 500 kb neuroblastoma tumor suppressor candidate region in chromosome 1p; mutation and stage-specific expression in UBE4B/UFD2. Oncogene 22, 2343–2351 (2003).
Carén, H., Holmstrand, A., Sjöberg, R.M. & Martinsson, T. The two human homologues of yeast UFD2 ubiquitination factor, UBE4A and UBE4B, are located in common neuroblastoma deletion regions and are subject to mutations in tumours. Eur. J. Cancer 42, 381–387 (2006).
Zage, P.E. et al. UBE4B levels are correlated with clinical outcomes in neuroblastoma patients and with altered neuroblastoma cell proliferation and sensitivity to epidermal growth factor receptor inhibitors. Cancer 119, 915–923 (2013).
Kloppsteck, P., Ewens, C.A., Förster, A., Zhang, X. & Freemont, P.S. Regulation of p97 in the ubiquitin-proteasome system by the UBX protein-family. Biochim. Biophys. Acta 1823, 125–129 (2012).
Bergink, S. et al. Role of Cdc48/p97 as a SUMO-targeted segregase curbing Rad51-Rad52 interaction. Nat. Cell Biol. 15, 526–532 (2013).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Zeiser, E., Frøkjær-Jensen, C., Jorgensen, E. & Ahringer, J. MosSCI and gateway compatible plasmid toolkit for constitutive and inducible expression of transgenes in the C. elegans germline. PLoS One 6, e20082 (2011).
Praitis, V., Casey, E., Collar, D. & Austin, J. Creation of low-copy integrated transgenic lines in Caenorhabditis elegans. Genetics 157, 1217–1226 (2001).
Timmons, L. & Fire, A. Specific interference by ingested dsRNA. Nature 395, 854 (1998).
Gumienny, T.L., Lambie, E., Hartwieg, E., Horvitz, H.R. & Hengartner, M.O. Genetic control of programmed cell death in the Caenorhabditis elegans hermaphrodite germline. Development 126, 1011–1022 (1999).
Hengartner, M.O., Ellis, R.E. & Horvitz, H.R. Caenorhabditis elegans gene ced-9 protects cells from programmed cell death. Nature 356, 494–499 (1992).
Franz, A. et al. CDC-48/p97 coordinates CDT-1 degradation with GINS chromatin dissociation to ensure faithful DNA replication. Mol. Cell 44, 85–96 (2011).
Lans, H. et al. DNA damage leads to progressive replicative decline but extends the life span of long-lived mutant animals. Cell Death Differ. 20, 1709–1718 (2013).
Stout, G.J. et al. Insulin/IGF-1-mediated longevity is marked by reduced protein metabolism. Mol. Syst. Biol. 9, 679 (2013).
Pourkarimi, E., Greiss, S. & Gartner, A. Evidence that CED-9/Bcl2 and CED-4/Apaf-1 localization is not consistent with the current model for C. elegans apoptosis induction. Cell Death Differ. 19, 406–415 (2012).
Gönczy, P. et al. Dissection of cell division processes in the one cell stage Caenorhabditis elegans embryo by mutational analysis. J. Cell Biol. 144, 927–946 (1999).
Hoogewijs, D., Houthoofd, K., Matthijssens, F., Vandesompele, J. & Vanfleteren, J.R. Selection and validation of a set of reliable reference genes for quantitative sod gene expression analysis in C. elegans. BMC Mol. Biol. 9, 9 (2008).
Lewis, J.A. & Fleming, J.T. Basic culture methods. Methods Cell Biol. 48, 3–29 (1995).
Spitzer, M., Wildenhain, J., Rappsilber, J. & Tyers, M. BoxPlotR: a web tool for generation of box plots. Nat. Methods 11, 121–122 (2014).
Acknowledgements
We thank Y. Kohara (National Institute of Genetics, Shizuoka, Japan), the Caenorhabditis Genetics Center (funded by the NIH National Center for Research Resources), the Bloomington Stock Center, M. Vidal (Dana-Farber Cancer Institute), Addgene and Geneservice Ltd for antibodies, plasmids, cDNAs and strains. We thank K. Gödderz, A. Lisowski and E. Stellbrink for technical help. We thank A. Franz and A. Gutschmidt for critical reading of the manuscript. We thank K. Ramadan (CRVK/MRC Oxford Institute for Radiation Oncology) and Y. Shiloh (David and Inez Myers Laboratory for Cancer Genetics) for insightful discussions on the project and for exchange of unpublished results. This work was supported by a Wellcome Trust Senior Research award (090944/Z/09/Z) to A.G.; grants from the German-Israeli Foundation (GIF 1104-68.11/2010), the Deutsche Forschungsgemeinschaft (EXC 229, SFB 829, SFB 670 and KFO 286), the European Research Council (ERC Starting Grant 260383), Marie Curie Actions (FP7 ITN CodeAge 316354, aDDRess 316390 and MARRIAGE 316964) and the Bundesministerium für Forschung und Bildung (Sybacol FKZ0315893A-B) to B.S.; and the Deutsche Forschungsgemeinschaft (EXC 229, HO 2541/8-1 and KFO 286) and the European Research Council (consolidator grant 616499) to T.H. In addition, this work was supported by COST Action (PROTEOSTASIS BM1307), supported by COST (European Cooperation in Science and Technology).
Author information
Authors and Affiliations
Contributions
L.A. and M.S. designed, performed and analyzed the experiments. W.P. performed in vitro ubiquitination assays. É.K. generated anti-ATX-3. A.G. and B.S. designed and performed the RNAi screen. B.S. and T.H. supervised the design and data interpretation. L.A., B.S. and T.H. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Cell-cycle arrest and developmental and physiological apoptosis are functional in ufd-2-deleted worms.
(a) Developmental apoptosis in head region of newly hatched L1 larvae of indicated genotypes. Data show means ± s.e.m. of 3 independent experiments. n = 38 animals for ced-1(e1735) and n = 33 animals for ced-1(e1735);ufd-2(tm1380). n.s. not significant in two-tailed Student’s t-test. (b) Physiological apoptosis in pachytene region of day 1 adults of indicated genotypes. Data show means ± s.e.m. of 3 independent experiments. n varied from 44-45 animals, see Supplementary Table 2. The triple asterisk indicates P values of ≤ 0.0001 in two-tailed Student’s t-test, n.s. not significant. (c) Representative DIC images of cell cycle arrest in mitotic region of germlines of indicated genotypes 16 hrs after IR treatment(60 Gy). Filled arrowheads mark enlarged arrested mitotic cells. Representative images of 3 independent experiments. (d) Quantification of cell cycle arrest in mitotic region of germlines 16 hrs after IR treatment (60 Gy). Data show means ± s.e.m. of 3 independent experiments. n varied from 23-33 animals, see Supplementary Table 2. The triple asterisk indicates P value of ≤ 0.0001 in two-tailed Student’s t-test, n.s. not significant. (e) Autoubiquitylation of UFD-2 using UFD-2 (wild-type), UFD-2P951A, or UFD-2C448Y as ubiquitin ligases and ubiquitin (wild-type), or ubiquitin mutants that contain only one lysine for conjugation (ubiquitinonly K29, ubiquitinonly K48). Representative immunoblot of 3 independent experiments.
Supplementary Figure 2 UFD-2 forms foci late after damage induction.
(a) α-UFD-2 and α-TBG-1 western blot. Samples are lysates of wild-type, ufd-2(tm1380) and ufd-(hh1) worms. Representative immunoblot of 3 independent experiments. (b) Schematic illustration of C. elegans germline and illustrative images of a IR treated (60 Gy) germline stained with α-UFD-2 and DAPI. Two slides from a z-stack (z-slide 1 and z-slide b), one from upper and one from lower part of the germline are shown. Filled arrowheads mark nuclei positive for UFD-2 foci. Scale bar, 10 µm. (c) Quantification of UFD-2 foci in pachytene region of wild-type germlines treated with IR (0, 60 Gy) isolated 5 and 24 hrs later. Data show means of ± s.e.m. of 3 experiments. n varied from 37-61 animals, see Supplementary Table 2.
Supplementary Figure 3 UFD-2 forms IR-dependent foci and interacts with CDC-48.
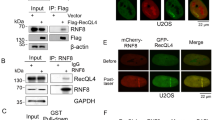
(a) Quantification of UFD-2 foci in pachytene region of wild-type germlines 24 hrs after treatment with IR (0, 60 Gy) or UV (600 J/m2). Data show means ± s.e.m. of 3 experiments. n varied from 69-96 animals, see Supplementary Table 2. (b) Embryonic survival after IR treatment (0, 30 or 60 Gy). Data show means ± s.e.m. of five experiments. n varied from 75-147 animals, see Supplementary Table 2. The single, double and triple asterisks indicate P values of ≤ 0.05, 0.001, 0.0001 in two-tailed Student’s t-test. (c) Representative images of worm germlines of indicated genotypes stained with α-ATX-3 antibody and DAPI 24 hrs after IR treatment (60 Gy). Scale bar, 5 µm. Representative images of 3 independent experiments. (d) α-ATX-3 and α-TBG-1 western blot. Samples are lysates of wild-type and atx-3(gk193) worms. Representative immunoblot of 3 independent experiments. (e) Quantification of number of UFD-2 foci that co-localize with ubiquitin in wild-type germlines 24 hrs after IR treatment (60 Gy). Data show means ± s.e.m. of 3 experiments. n = 32 animals. (f) Sequence alignment shows UFD-2 from C. elegans and other species, the conserved residue C448 is highlighted. (g) in vivo co-immunoprecipitation of CDC-48 with UFD-2 from indicated worm lysates. Representative immunoblot of 3 independent experiments. (h) in vitro co-immunoprecipitation of CDC-48 with UFD-2 with purified recombinantly expressed protein. Representative immunoblot of 3 independent experiments. (i) Autoubiquitylation of UFD-2 using UFD-2 (wild-type), UFD-2P951A or UFD-2C448Y as ubiquitin ligases. Representative immunoblot of 3 independent experiments. (j) Quantification of UFD-2 foci in pachytene region of indicated germlines 24 hrs after IR treatment (0, 60 Gy). Data show means of ± s.e.m. of three experiments. n varied from 40-43 animals, see Supplementary Table 2. (k) Quantification of ubiquitin foci in pachytene region of indicated germlines 24 hrs after IR treatment (60 Gy). Data show means of ± s.e.m. of three experiments. n varied from 41-45 animals, see Supplementary Table 2.
Supplementary Figure 4 Apoptosis induction in ufd-2-deleted worms is functional.
(a) Representative images of worm germlines of indicated genotypes stained with α-UFD-2 antibody and DAPI 24 hrs after IR treatment (60 Gy). Filled and empty arrowhead indicated nuclei positive or negative for UFD-2 foci, respectively. Scale bar, 5 µm. Representative images of 3 independent experiments. (b) α-CEP-1 and α-TBG-1 western blot of worm lysates of indicated genotypes. Representative immunoblot of 3 independent experiments and (c) corresponding quantification. (d) Relative expression levels of egl-1 target gene in wild-type and ufd-2(tm1380) worms at indicated time points after IR treatment (0 or 60 Gy). mRNA levels were normalized to 0 Gy samples. Data show means ± s.e.m. of 3 independent experiments.
Supplementary Figure 5 UFD-2 foci are dependent on apoptosis and DNA-damage signaling.
(a) Representative images of worm germlines of indicated genotypes stained with α-UFD-2 antibody and DAPI 24 hrs after IR treatment (60 Gy). Filled and empty arrowhead indicated nuclei positive or negative for UFD-2 foci, respectively. Scale bar, 5 µm. Representative images of 3 independent experiments. (b) α-UFD and α-TBG-1 western blot of worm lysates of indicated genotypes 24 hrs after IR treatment (0, 60 Gy). Representative immunoblot of 3 independent experiments. (c) Quantification of germ cells positive for RAD-51 staining of indicated genotypes 24 hrs after IR treatment (0 or 20 Gy). Data show means ± s.e.m. of 3 independent experiments. n varied from 30-33 animals, see Supplementary Table 2. n.s. not significant in two-tailed Student’s t-test.
Supplementary Figure 6 RAD-51 in DSB repair.
(a) Embryonic survival after IR treatment (0, 30 or 60 Gy) of indicated genotypes. Data show means ± s.e.m. of 3 independent experiments. n varied from 90-95 animals, see Supplementary Table 2. The single asterisk indicates P value of ≤ 0.05 in two-tailed Student’s t-test. (b) Quantification of germ cells positive for RAD-51 staining of indicated genotypes 24 hrs after IR treatment (0 or 20 Gy). Data show means ± s.e.m. of 3 independent experiments. n varied from 35-48 animals, see Supplementary Table 2. The single and double asterisks indicate P values of ≤ 0.05 and 0.001 in two-tailed Student’s t-test. (c) Embryonic survival after IR treatment (0, 30 or 60 Gy) of indicated genotypes treated with rad-51 or control RNAi. Data show means ± s.e.m. of one experiment, n varied from 15-25 animals, see Supplementary Table 2. The double asterisk indicates P value of ≤ 0.001 in two-tailed Student’s t-test.
Supplementary Figure 7 Full-sized images of immunoblots in figures.
(a) Presented in Figure 1e. (b) Presented in Supplementary Figure 1e. (c) Presented in Supplementary Figure 2a. (d) Presented in Supplementary Figure 3d. (e) Presented in Supplementary Figure 3g. (f) Presented in Supplementary Figure 3h. (g) Presented in Supplementary Figure 3i. (h) Presented in Supplementary Figure 4b. (i) Presented in Supplementary Figure 5b.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 1616 kb)
Supplementary Table 1
n values for Figures 1–6 (XLSX 13 kb)
Supplementary Table 2
n values for Supplementary Figures 1–6 (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Ackermann, L., Schell, M., Pokrzywa, W. et al. E4 ligase–specific ubiquitination hubs coordinate DNA double-strand-break repair and apoptosis. Nat Struct Mol Biol 23, 995–1002 (2016). https://doi.org/10.1038/nsmb.3296
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3296
This article is cited by
-
DNA damage response revisited: the p53 family and its regulators provide endless cancer therapy opportunities
Experimental & Molecular Medicine (2022)
-
UFD-2 is an adaptor-assisted E3 ligase targeting unfolded proteins
Nature Communications (2018)