Abstract
Removal of the 5′ cap on mRNA by the decapping enzyme Dcp2 is a critical step in 5′-to-3′ mRNA decay. Understanding the structural basis of Dcp2 activity has been a challenge because Dcp2 is dynamic and has weak affinity for the cap substrate. Here we present a 2.6-Å-resolution crystal structure of a heterotrimer of fission yeast Dcp2, its essential activator Dcp1, and the human NMD cofactor PNRC2, in complex with a tight-binding cap analog. Cap binding is accompanied by a conformational change in Dcp2, thereby forming a composite nucleotide-binding site comprising conserved residues in the catalytic and regulatory domains. Kinetic analysis of PNRC2 revealed that a conserved short linear motif enhances both substrate affinity and the catalytic step of decapping. These findings explain why Dcp2 requires a conformational change for efficient catalysis and reveals that coactivators promote RNA binding and the catalytic step of decapping, possibly through different conformational states.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Moore, M.J. From birth to death: the complex lives of eukaryotic mRNAs. Science 309, 1514–1518 (2005).
Topisirovic, I., Svitkin, Y.V., Sonenberg, N. & Shatkin, A.J. Cap and cap-binding proteins in the control of gene expression. Wiley Interdiscip. Rev. RNA 2, 277–298 (2011).
Parker, R. RNA degradation in Saccharomyces cerevisae. Genetics 191, 671–702 (2012).
Arribas-Layton, M., Wu, D., Lykke-Andersen, J. & Song, H. Structural and functional control of the eukaryotic mRNA decapping machinery. Biochim. Biophys. Acta 1829, 580–589 (2013).
Li, Y. & Kiledjian, M. Regulation of mRNA decapping. Wiley Interdiscip. Rev. RNA 1, 253–265 (2010).
Mildvan, A.S. et al. Structures and mechanisms of Nudix hydrolases. Arch. Biochem. Biophys. 433, 129–143 (2005).
Nagarajan, V.K., Jones, C.I., Newbury, S.F. & Green, P.J. XRN 5′ → 3′ exoribonucleases: structure, mechanisms and functions. Biochim. Biophys. Acta 1829, 590–603 (2013).
Geisler, S., Lojek, L., Khalil, A.M., Baker, K.E. & Coller, J. Decapping of long noncoding RNAs regulates inducible genes. Mol. Cell 45, 279–291 (2012).
Guo, H., Ingolia, N.T., Weissman, J.S. & Bartel, D.P. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 466, 835–840 (2010).
Jonas, S. & Izaurralde, E. Towards a molecular understanding of microRNA-mediated gene silencing. Nat. Rev. Genet. 16, 421–433 (2015).
Kervestin, S. & Jacobson, A. NMD: a multifaceted response to premature translational termination. Nat. Rev. Mol. Cell Biol. 13, 700–712 (2012).
Popp, M.W.-L. & Maquat, L.E. Organizing principles of mammalian nonsense-mediated mRNA decay. Annu. Rev. Genet. 47, 139–165 (2013).
Cho, H., Kim, K.M. & Kim, Y.K. Human proline-rich nuclear receptor coregulatory protein 2 mediates an interaction between mRNA surveillance machinery and decapping complex. Mol. Cell 33, 75–86 (2009).
Cho, H. et al. Staufen1-mediated mRNA decay functions in adipogenesis. Mol. Cell 46, 495–506 (2012).
Park, E. & Maquat, L.E. Staufen-mediated mRNA decay. Wiley Interdiscip. Rev. RNA 4, 423–435 (2013).
Brannan, K. et al. mRNA decapping factors and the exonuclease Xrn2 function in widespread premature termination of RNA polymerase II transcription. Mol. Cell 46, 311–324 (2012).
Presnyak, V. et al. Codon optimality is a major determinant of mRNA stability. Cell 160, 1111–1124 (2015).
Shukla, S. & Parker, R. Quality control of assembly-defective U1 snRNAs by decapping and 5′-to-3′ exonucleolytic digestion. Proc. Natl. Acad. Sci. USA 111, E3277–E3286 (2014).
Li, Y., Dai, J., Song, M., Fitzgerald-Bocarsly, P. & Kiledjian, M. Dcp2 decapping protein modulates mRNA stability of the critical interferon regulatory factor (IRF) IRF-7. Mol. Cell. Biol. 32, 1164–1172 (2012).
She, M. et al. Structural basis of dcp2 recognition and activation by dcp1. Mol. Cell 29, 337–349 (2008).
Dunckley, T., Tucker, M. & Parker, R. Two related proteins, Edc1p and Edc2p, stimulate mRNA decapping in Saccharomyces cerevisiae. Genetics 157, 27–37 (2001).
Borja, M.S., Piotukh, K., Freund, C. & Gross, J.D. Dcp1 links coactivators of mRNA decapping to Dcp2 by proline recognition. RNA 17, 278–290 (2011).
Fromm, S.A. et al. The structural basis of Edc3- and Scd6-mediated activation of the Dcp1:Dcp2 mRNA decapping complex. EMBO J. 31, 279–290 (2012).
Fromm, S.A. et al. In vitro reconstitution of a cellular phase-transition process that involves the mRNA decapping machinery. Angew. Chem. Int. Ed. Engl. 53, 7354–7359 (2014).
Harigaya, Y., Jones, B.N., Muhlrad, D., Gross, J.D. & Parker, R. Identification and analysis of the interaction between Edc3 and Dcp2 in Saccharomyces cerevisiae. Mol. Cell. Biol. 30, 1446–1456 (2010).
Chang, C.-T., Bercovich, N., Loh, B., Jonas, S. & Izaurralde, E. The activation of the decapping enzyme DCP2 by DCP1 occurs on the EDC4 scaffold and involves a conserved loop in DCP1. Nucleic Acids Res. 42, 5217–5233 (2014).
Lai, T. et al. Structural basis of the PNRC2-mediated link between mRNA surveillance and decapping. Structure 20, 2025–2037 (2012).
Chowdhury, A., Mukhopadhyay, J. & Tharun, S. The decapping activator Lsm1p-7p-Pat1p complex has the intrinsic ability to distinguish between oligoadenylated and polyadenylated RNAs. RNA 13, 998–1016 (2007).
Chowdhury, A. & Tharun, S. Activation of decapping involves binding of the mRNA and facilitation of the post-binding steps by the Lsm1-7-Pat1 complex. RNA 15, 1837–1848 (2009).
Coller, J. & Parker, R. General translational repression by activators of mRNA decapping. Cell 122, 875–886 (2005).
Sweet, T., Kovalak, C. & Coller, J. The DEAD-box protein Dhh1 promotes decapping by slowing ribosome movement. PLoS Biol. 10, e1001342 (2012).
She, M. et al. Crystal structure and functional analysis of Dcp2p from Schizosaccharomyces pombe. Nat. Struct. Mol. Biol. 13, 63–70 (2006).
Deshmukh, M.V. et al. mRNA decapping is promoted by an RNA-binding channel in Dcp2. Mol. Cell 29, 324–336 (2008).
He, F. & Jacobson, A. Control of mRNA decapping by positive and negative regulatory elements in the Dcp2 C-terminal domain. RNA 21, 1633–1647 (2015).
Piccirillo, C., Khanna, R. & Kiledjian, M. Functional characterization of the mammalian mRNA decapping enzyme hDcp2. RNA 9, 1138–1147 (2003).
Floor, S.N., Borja, M.S. & Gross, J.D. Interdomain dynamics and coactivation of the mRNA decapping enzyme Dcp2 are mediated by a gatekeeper tryptophan. Proc. Natl. Acad. Sci. USA 109, 2872–2877 (2012).
Floor, S.N., Jones, B.N., Hernandez, G.A. & Gross, J.D. A split active site couples cap recognition by Dcp2 to activation. Nat. Struct. Mol. Biol. 17, 1096–1101 (2010).
Jonas, S. & Izaurralde, E. The role of disordered protein regions in the assembly of decapping complexes and RNP granules. Genes Dev. 27, 2628–2641 (2013).
Valkov, E. et al. Structure of the Dcp2–Dcp1 mRNA-decapping complex in the activated conformation. Nat. Struct. Mol. Biol. 23, 574–579 (2016).
Ziemniak, M. et al. Two-headed tetraphosphate cap analogs are inhibitors of the Dcp1/2 RNA decapping complex. RNA 22, 518–529 (2016).
Ziemniak, M. et al. Phosphate-modified analogues of m7GTP and m7Gppppm7G: synthesis and biochemical properties. Bioorg. Med. Chem. 23, 5369–5381 (2015).
Marcotrigiano, J., Gingras, A.-C., Sonenberg, N. & Burley, S.K. Cocrystal structure of the messenger RNA 5′ cap-binding protein (eIF4E) bound to 7-methyl-GDP. Cell 89, 951–961 (1997).
Mazza, C., Segref, A., Mattaj, I.W. & Cusack, S. Large-scale induced fit recognition of an m7GpppG cap analogue by the human nuclear cap-binding complex. EMBO J. 21, 5548–5557 (2002).
Bailey, S. et al. The crystal structure of diadenosine tetraphosphate hydrolase from Caenorhabditis elegans in free and binary complex forms. Structure 10, 589–600 (2002).
Svensson, L.M. et al. Crystal structure of human MTH1 and the 8-oxo-dGMP product complex. FEBS Lett. 585, 2617–2621 (2011).
Jones, B.N., Quang-Dang, D.-U., Oku, Y. & Gross, J.D. A kinetic assay to monitor RNA decapping under single-turnover conditions. Methods Enzymol. 448, 23–40 (2008).
Smith, N.C. & Matthews, J.M. Mechanisms of DNA-binding specificity and functional gene regulation by transcription factors. Curr. Opin. Struct. Biol. 38, 68–74 (2016).
von Hippel, P.H. From “simple” DNA-protein interactions to the macromolecular machines of gene expression. Annu. Rev. Biophys. Biomol. Struct. 36, 79–105 (2007).
Kalodimos, C.G. et al. Structure and flexibility adaptation in nonspecific and specific protein-DNA complexes. Science 305, 386–389 (2004).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Dolinsky, T.J. et al. PDB2PQR: expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res. 35, W522–W525 (2007).
Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & McCammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Acknowledgements
The authors thank X. Liu, J. Binning, and C. Waddling at UCSF for valuable help and advice on crystallography experiments, and J. Holton and G. Meigs at Lawrence Berkeley National Laboratory, Advanced Light Source beamline 8.3.1, for help with X-ray data collection. We also thank J. Kowalska for helpful discussions on the design and synthesis of the two-headed cap analog. This work was supported by the US National Institutes of Health (R01 GM078360 to J.D.G. and NRSA fellowship F32 GM105313 to J.S.M.) and the National Science Centre, Poland (grant no. UMO-2012/05/E/ST5/03893 to J.J. and fellowship no. UMO-2014/12/T/NZ1/00528 to M.Z.). The Advanced Light Source is supported by the US Department of Energy under contract no. DE-AC02-05CH11231.
Author information
Authors and Affiliations
Contributions
J.S.M. designed and purified all protein constructs, carried out crystallization experiments, collected and refined crystallographic data, carried out decapping kinetics experiments, wrote the manuscript, and prepared the figures. M.Z. and J.J. designed and synthesized the two-headed cap analog. J.D.G. supervised the project and experimental design and guided manuscript preparation and editing. All authors read and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Human PNRC2(91–121) activates fission yeast Dcp1–Dcp2.
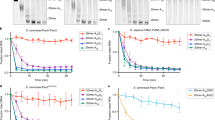
(a) Complexes of Hs PNRC2(91-121)–Sp Dcp1–Dcp2(1-243) (purple) decap a 355-mer RNA substrate faster than Sp Dcp1–Dcp2 alone (green). kobs is plotted against protein concentration to determine kinetic constants. Errors are s.d. on individual fits to determine kobs. (b) Bar graph showing kmax (red) and KM (blue) values determined from fits of kinetic data shown in (a). For Hs PNRC2(91-121)–Sp Dcp1–Dcp2(1-243) kmax = 1.22 ± 0.09 min-1 and KM = 27 ± 4 µM, for Sp Dcp1–Dcp2(1-243) kmax = 0.15 ± 0.02 min-1 and KM = 26 ± 5 µM. This corresponds to an 8-fold activation in catalysis by PNRC2(91-121). Errors are s.d. of fits to determine kmax and KM. (c) Single chain constructs where PNRC2(91-121) is tethered to the N-terminus of Dcp2 by a flexible GGGGS linker (Dcp1–scPNRC2Dcp2) are more active than co-purified complexes of PNRC2(91-121)–Dcp1–Dcp2. Plots of fraction m7GDP product versus time are shown for decapping reactions for both complexes under saturating protein concentrations (25 µM). (d) Bar graph showing kobs values determined from the decapping time course shown in (c). For the co-purified complex PNRC2(91-121)–Dcp1–Dcp2, kobs = 0.72 ± 0.08 min-1; for the single chain Dcp1–scPNRC2Dcp2 complex, kobs = 1.33 ± 0.07 min-1. Errors are s.d. of the individual exponential fits to obtain kobs shown in (c).
Supplementary Figure 2 Crystal structure of the apo Dcp1–scPNRC2Dcp2 complex.
(a) Asymmetric unit of Dcp1–scPNRC2Dcp2. The Dcp1–scPNRC2Dcp2 complex crystallizes as a nearly symmetric domain-swapped dimer, in which the PNRC2 peptide tethered to the N-terminus of Dcp2 binds in trans to Dcp1 in the adjacent protomer. PNRC2 peptides are colored the same as the Dcp2 regulatory domain to which they are tethered; a dotted line shows where the flexible GGGGS linker connects the C-terminus of PNRC2(91-121) to the N-terminus of Dcp2. (b) Alignment of apo Dcp1–scPNRC2Dcp2 structure (colored, PNRC2 not shown) from this study, with closed, ATP-bound Sp Dcp1–Dcp2 structure from PDB 2QKM20 (Dcp1–Dcp2 in light gray, ATP in dark gray, Dcp2 residues 1-243 are shown). Dcp1–Dcp2(1-90) was used to align the structures; backbone RMSD is 0.7 Å over 213 residues. (c) Alignment of apo Dcp1–scPNRC2Dcp2 structure (colored, PNRC2 not shown) from this study, with Sp Dcp1–Dcp2 structure from PDB 5J3Y39 (light gray). Dcp1–Dcp2(1-90) was used to align the structures; backbone RMSD is 1.2 Å over 214 residues.
Supplementary Figure 3 Two-headed cap analog binds both molecules of the Dcp1–scPRNC2Dcp2 asymmetric unit in a different conformation.
(a) Asymmetric unit of Dcp1–scPNRC2Dcp2 with bound cap analogs. One Dcp1–Dcp2 protomer undergoes a conformational change to bind cap analog at a composite active site (colored in blues; this binding mode and conformational change are described in the main text). The second protomer of Dcp1–Dcp2 has the same conformation as in the apo structure, but with cap analog bound to exclusively the Nudix domain of Dcp2 (colored in reds). (b) Cap analog binds exclusively to the Nudix domain of Dcp2 by bridging Y220 and W117. Fo-Fc omit map shown at 1.5σ. (c) Although the electron density for the cap analog bound to only the Nudix domain is weak and discontinuous, likely due to weak binding and flexibility in the phosphate chain, this binding mode agrees perfectly with previous NMR chemical shift mapping experiments in which cap analog was titrated into the isolated Nudix domain.40 The cap analog-bound Nudix domain is shown here with previously determined NMR chemical shift perturbations (green) and resonance broadening (cyan) mapped to the surface.
Supplementary Figure 4 Cap-binding residues are occluded or separated by large distances in previous structures of Dcp1–Dcp2.
Large rotations of the Nudix domain are required for all previously identified closed conformations of Dcp1–Dcp2 to access the cap-binding conformation described in this study. Dcp1 is yellow, Dcp2 regulatory domain is purple, Dcp2 Nudix domain is green, Dcp2 dorsal RNA binding helix is blue, cap-binding residues are red. (a) cap analog-bound conformation of Sp Dcp1–Dcp2 from this study (top; PNRC2 not shown), and close-up view of W43 / Y220 residues (bottom). Y92 points toward the interior of the protein to bind cap, W43 and Y220 from the regulatory and Nudix domains of Dcp2 are positioned close to one another and are surface exposed in order to sandwich cap. (b) Closed, ATP-bound conformation of Sp Dcp1–Dcp2 from PDB 2QKM (top),20 and close-up view of W43 / Y220 residues (bottom). Y92 points toward solvent, W43 is occluded by residue R167 and unavailable for cap binding, and Y220 binds ATP but is not aligned with W43 for cap binding. A 30° rotation of the Nudix domain is required to bind cap with the composite active site depicted in (a). (c) Structure of Sp Edc1–Dcp1–Dcp2 from PDB 5J3T (top)39 and close-up of W43 / Y220 residues (bottom); Sp Edc1 peptide is colored orange. Y92 points toward solvent, W43 is buried in the interface with Sp Edc1 and the Nudix domain of Dcp2 and is unavailable to bind cap, and W43 and Y220 are separated by nearly 20 Å. A 90° rotation of the Nudix domain is required to form the composite active site as in (a).
Supplementary Figure 5 Interaction diagram showing contacts between two-headed cap analog and Dcp2 residues.
Residues on the Dcp2 regulatory domain (1-95) are shown in purple, residues on the Dcp2 Nudix domain (96-243) are shown in green. All distances are in Angstroms, and are heteroatom-heteroatom distances. Curved bold lines denote aromatic stacking interactions. Charges on protein sidechains and the ligand phosphate backbone are omitted in this schematic diagram.
Supplementary Figure 6 Alternative RNA binding path on Dcp1–Dcp2.
An alternative pathway for RNA binding to Dcp1–Dcp2 in the cap analog-bound conformation is shown on the electrostatic surface (a), or on a cartoon with RNA binding residues highlighted in blue (b). In this putative RNA binding path, RNA binds the dorsal helix of the Nudix domain (Box B motif), follows a flexible, lysine-rich loop (K206-K216) along the exterior of the protein, Y92 binds the first transcribed nucleotide, and m7G of cap is positioned near the center of the enzyme and bound by W43/Y220/D47.
Supplementary Figure 7 Comparison of decapping activation kinetics by Hs PNRC2 and Sp Edc1, and purification of PNRC2–Dcp1–Dcp2 complexes for kinetics experiments.
(a) Sp Edc1 strongly activates mRNA decapping by Dcp1–Dcp2. Comparison of decapping activation kinetics for complexes of Hs PNRC2(91-121) or Sp Edc1 with Sp Dcp1–Dcp2. Sp Edc1 is a stronger activator than Hs PNRC2 at 25 µM enzyme complex concentration; decapping kinetics with Sp Edc1 are too fast to extract kobs under these conditions by manual pipetting. (b) Size exclusion chromatography of the WT Hs PNRC2(1-121) – Sp Dcp1–Dcp2 complex; Hs PNRC2 binds to Sp Dcp1–Dcp2 and the proteins co-elute (top), as visualized by SDS-PAGE (bottom). (c) SDS-PAGE of purified Hs PNRC2(1-121) – Sp Dcp1–Dcp2 complexes with PNRC2 point mutants in the YAGxxF Dcp2-activating motif used in kinetic experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Table 1 (PDF 1815 kb)
Supplementary Data Set 1
Kinetic data associated with Figure 4 and Supplementary Figure 1a,b. (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Mugridge, J., Ziemniak, M., Jemielity, J. et al. Structural basis of mRNA-cap recognition by Dcp1–Dcp2. Nat Struct Mol Biol 23, 987–994 (2016). https://doi.org/10.1038/nsmb.3301
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3301
This article is cited by
-
YBX1 mediates alternative splicing and maternal mRNA decay during pre-implantation development
Cell & Bioscience (2022)
-
Non-canonical features of microRNAs: paradigms emerging from cardiovascular disease
Nature Reviews Cardiology (2022)
-
miR-4293 upregulates lncRNA WFDC21P by suppressing mRNA-decapping enzyme 2 to promote lung carcinoma proliferation
Cell Death & Disease (2021)
-
Structure of the activated Edc1-Dcp1-Dcp2-Edc3 mRNA decapping complex with substrate analog poised for catalysis
Nature Communications (2018)
-
Structural and molecular mechanisms for the control of eukaryotic 5′–3′ mRNA decay
Nature Structural & Molecular Biology (2018)