Abstract
Telomere length maintenance ensures self-renewal of human embryonic stem cells (hESCs) and induced pluripotent stem cells (hiPSCs); however, the mechanisms governing telomere length homeostasis in these cell types are unclear. Here, we report that telomere length is determined by the balance between telomere elongation, which is mediated by telomerase, and telomere trimming, which is controlled by XRCC3 and Nbs1, homologous recombination proteins that generate single-stranded C-rich telomeric DNA and double-stranded telomeric circular DNA (T-circles), respectively. We found that reprogramming of differentiated cells induces T-circle and single-stranded C-rich telomeric DNA accumulation, indicating the activation of telomere trimming pathways that compensate telomerase-dependent telomere elongation in hiPSCs. Excessive telomere elongation compromises telomere stability and promotes the formation of partially single-stranded telomeric DNA circles (C-circles) in hESCs, suggesting heightened sensitivity of stem cells to replication stress at overly long telomeres. Thus, tight control of telomere length homeostasis is essential to maintain telomere stability in hESCs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
de Lange, T. Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev. 19, 2100–2110 (2005).
Makarov, V.L., Hirose, Y. & Langmore, J.P. Long G tails at both ends of human chromosomes suggest a C strand degradation mechanism for telomere shortening. Cell 88, 657–666 (1997).
Griffith, J.D. et al. Mammalian telomeres end in a large duplex loop. Cell 97, 503–514 (1999).
de Lange, T. T-loops and the origin of telomeres. Nat. Rev. Mol. Cell Biol. 5, 323–329 (2004).
Doksani, Y., Wu, J.Y., de Lange, T. & Zhuang, X. Super-resolution fluorescence imaging of telomeres reveals TRF2-dependent T-loop formation. Cell 155, 345–356 (2013).
Olovnikov, A.M. A theory of marginotomy. The incomplete copying of template margin in enzymic synthesis of polynucleotides and biological significance of the phenomenon. J. Theor. Biol. 41, 181–190 (1973).
Watson, J.D. Origin of concatemeric T7 DNA. Nat. New Biol. 239, 197–201 (1972).
Wu, P., Takai, H. & de Lange, T. Telomeric 3′ overhangs derive from resection by Exo1 and Apollo and fill-in by POT1b-associated CST. Cell 150, 39–52 (2012).
Kim, N.W. et al. Specific association of human telomerase activity with immortal cells and cancer. Science 266, 2011–2015 (1994).
Wright, W.E., Piatyszek, M.A., Rainey, W.E., Byrd, W. & Shay, J.W. Telomerase activity in human germline and embryonic tissues and cells. Dev. Genet. 18, 173–179 (1996).
Cohen, S.B. et al. Protein composition of catalytically active human telomerase from immortal cells. Science 315, 1850–1853 (2007).
Bryan, T.M., Englezou, A., Dalla-Pozza, L., Dunham, M.A. & Reddel, R.R. Evidence for an alternative mechanism for maintaining telomere length in human tumors and tumor-derived cell lines. Nat. Med. 3, 1271–1274 (1997).
Londoño-Vallejo, J.A., Der-Sarkissian, H., Cazes, L., Bacchetti, S. & Reddel, R.R. Alternative lengthening of telomeres is characterized by high rates of telomeric exchange. Cancer Res. 64, 2324–2327 (2004).
Yeager, T.R. et al. Telomerase-negative immortalized human cells contain a novel type of promyelocytic leukemia (PML) body. Cancer Res. 59, 4175–4179 (1999).
Bryan, T.M., Englezou, A., Gupta, J., Bacchetti, S. & Reddel, R.R. Telomere elongation in immortal human cells without detectable telomerase activity. EMBO J. 14, 4240–4248 (1995).
Oganesian, L. & Karlseder, J. Mammalian 5′ C-rich telomeric overhangs are a mark of recombination-dependent telomere maintenance. Mol. Cell 42, 224–236 (2011).
Tokutake, Y. et al. Extra-chromosomal telomere repeat DNA in telomerase-negative immortalized cell lines. Biochem. Biophys. Res. Commun. 247, 765–772 (1998).
Cesare, A.J. & Griffith, J.D. Telomeric DNA in ALT cells is characterized by free telomeric circles and heterogeneous T-loops. Mol. Cell. Biol. 24, 9948–9957 (2004).
Henson, J.D. et al. DNA C-circles are specific and quantifiable markers of alternative-lengthening-of-telomeres activity. Nat. Biotechnol. 27, 1181–1185 (2009).
Sharpless, N.E. & DePinho, R.A. How stem cells age and why this makes us grow old. Nat. Rev. Mol. Cell Biol. 8, 703–713 (2007).
Aubert, G. Telomere dynamics and aging. Prog. Mol. Biol. Transl. Sci. 125, 89–111 (2014).
Zeng, S. et al. Telomerase-mediated telomere elongation from human blastocysts to embryonic stem cells. J. Cell Sci. 127, 752–762 (2014).
Batista, L.F. et al. Telomere shortening and loss of self-renewal in dyskeratosis congenita induced pluripotent stem cells. Nature 474, 399–402 (2011).
Raices, M. et al. C. elegans telomeres contain G-strand and C-strand overhangs that are bound by distinct proteins. Cell 132, 745–757 (2008).
Pickett, H.A., Cesare, A.J., Johnston, R.L., Neumann, A.A. & Reddel, R.R. Control of telomere length by a trimming mechanism that involves generation of t-circles. EMBO J. 28, 799–809 (2009).
Pickett, H.A., Henson, J.D., Au, A.Y., Neumann, A.A. & Reddel, R.R. Normal mammalian cells negatively regulate telomere length by telomere trimming. Hum. Mol. Genet. 20, 4684–4692 (2011).
Nabetani, A. & Ishikawa, F. Unusual telomeric DNAs in human telomerase-negative immortalized cells. Mol. Cell. Biol. 29, 703–713 (2009).
Wang, R.C., Smogorzewska, A. & de Lange, T. Homologous recombination generates T-loop-sized deletions at human telomeres. Cell 119, 355–368 (2004).
O'Sullivan, R.J. et al. Rapid induction of alternative lengthening of telomeres by depletion of the histone chaperone ASF1. Nat. Struct. Mol. Biol. 21, 167–174 (2014).
Marion, R.M. et al. Telomeres acquire embryonic stem cell characteristics in induced pluripotent stem cells. Cell Stem Cell 4, 141–154 (2009).
Agarwal, S. et al. Telomere elongation in induced pluripotent stem cells from dyskeratosis congenita patients. Nature 464, 292–296 (2010).
Wang, F. et al. Molecular insights into the heterogeneity of telomere reprogramming in induced pluripotent stem cells. Cell Res. 22, 757–768 (2012).
Panopoulos, A.D. et al. Rapid and highly efficient generation of induced pluripotent stem cells from human umbilical vein endothelial cells. PLoS One 6, e19743 (2011).
Panopoulos, A.D. et al. The metabolome of induced pluripotent stem cells reveals metabolic changes occurring in somatic cell reprogramming. Cell Res. 22, 168–177 (2012).
Poole, L.A. et al. SMARCAL1 maintains telomere integrity during DNA replication. Proc. Natl. Acad. Sci. USA 112, 14864–14869 (2015).
Rizzo, A. et al. Stabilization of quadruplex DNA perturbs telomere replication leading to the activation of an ATR-dependent ATM signaling pathway. Nucleic Acids Res. 37, 5353–5364 (2009).
Lin, W. et al. Mammalian DNA2 helicase/nuclease cleaves G-quadruplex DNA and is required for telomere integrity. EMBO J. 32, 1425–1439 (2013).
Thangavel, S. et al. DNA2 drives processing and restart of reversed replication forks in human cells. J. Cell Biol. 208, 545–562 (2015).
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009).
Oganesian, L. & Karlseder, J. 5′ C-rich telomeric overhangs are an outcome of rapid telomere truncation events. DNA Repair (Amst.) 12, 238–245 (2013).
Compton, S.A., Choi, J.H., Cesare, A.J., Ozgür, S. & Griffith, J.D. Xrcc3 and Nbs1 are required for the production of extrachromosomal telomeric circles in human alternative lengthening of telomere cells. Cancer Res. 67, 1513–1519 (2007).
Cesare, A.J. & Reddel, R.R. Alternative lengthening of telomeres: models, mechanisms and implications. Nat. Rev. Genet. 11, 319–330 (2010).
Niida, H. et al. Telomere maintenance in telomerase-deficient mouse embryonic stem cells: characterization of an amplified telomeric DNA. Mol. Cell. Biol. 20, 4115–4127 (2000).
Wang, Y. et al. An increase in telomere sister chromatid exchange in murine embryonic stem cells possessing critically shortened telomeres. Proc. Natl. Acad. Sci. USA 102, 10256–10260 (2005).
Zalzman, M. et al. Zscan4 regulates telomere elongation and genomic stability in ES cells. Nature 464, 858–863 (2010).
Sexton, A.N. et al. Genetic and molecular identification of three human TPP1 functions in telomerase action: recruitment, activation, and homeostasis set point regulation. Genes Dev. 28, 1885–1899 (2014).
Winkler, T. et al. Defective telomere elongation and hematopoiesis from telomerase-mutant aplastic anemia iPSCs. J. Clin. Invest. 123, 1952–1963 (2013).
Moon, D.H. et al. Poly(A)-specific ribonuclease (PARN) mediates 3′-end maturation of the telomerase RNA component. Nat. Genet. 47, 1482–1488 (2015).
Gu, B.W. et al. Impaired telomere maintenance and decreased canonical WNT signaling but normal ribosome biogenesis in induced pluripotent stem cells from X-linked dyskeratosis congenita patients. PLoS One 10, e0127414 (2015).
Verdun, R.E. & Karlseder, J. The DNA damage machinery and homologous recombination pathway act consecutively to protect human telomeres. Cell 127, 709–720 (2006).
Zhong, Z.H. et al. Disruption of telomere maintenance by depletion of the MRE11/RAD50/NBS1 complex in cells that use alternative lengthening of telomeres. J. Biol. Chem. 282, 29314–29322 (2007).
Takata, M. et al. Chromosome instability and defective recombinational repair in knockout mutants of the five Rad51 paralogs. Mol. Cell. Biol. 21, 2858–2866 (2001).
Dumon-Jones, V. et al. Nbn heterozygosity renders mice susceptible to tumor formation and ionizing radiation-induced tumorigenesis. Cancer Res. 63, 7263–7269 (2003).
Gilson, E. & Géli, V. How telomeres are replicated. Nat. Rev. Mol. Cell Biol. 8, 825–838 (2007).
Martínez, P. et al. Increased telomere fragility and fusions resulting from TRF1 deficiency lead to degenerative pathologies and increased cancer in mice. Genes Dev. 23, 2060–2075 (2009).
Bétous, R. et al. SMARCAL1 catalyzes fork regression and Holliday junction migration to maintain genome stability during DNA replication. Genes Dev. 26, 151–162 (2012).
Ahuja, A.K. et al. A short G1 phase imposes constitutive replication stress and fork remodelling in mouse embryonic stem cells. Nat. Commun. 7, 10660 (2016).
Göhring, J., Fulcher, N., Jacak, J. & Riha, K. TeloTool: a new tool for telomere length measurement from terminal restriction fragment analysis with improved probe intensity correction. Nucleic Acids Res. 42, e21 (2014).
Ludwig, T.E. et al. Derivation of human embryonic stem cells in defined conditions. Nat. Biotechnol. 24, 185–187 (2006).
Wong, J.M. & Collins, K. Telomerase RNA level limits telomere maintenance in X-linked dyskeratosis congenita. Genes Dev. 20, 2848–2858 (2006).
O'Sullivan, R.J., Kubicek, S., Schreiber, S.L. & Karlseder, J. Reduced histone biosynthesis and chromatin changes arising from a damage signal at telomeres. Nat. Struct. Mol. Biol. 17, 1218–1225 (2010).
Sarbassov, D.D., Guertin, D.A., Ali, S.M. & Sabatini, D.M. Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 307, 1098–1101 (2005).
Ruiz, S. et al. A high proliferation rate is required for cell reprogramming and maintenance of human embryonic stem cell identity. Curr. Biol. 21, 45–52 (2011).
Cesare, A.J. et al. Spontaneous occurrence of telomeric DNA damage response in the absence of chromosome fusions. Nat. Struct. Mol. Biol. 16, 1244–1251 (2009).
Karlseder, J., Smogorzewska, A. & de Lange, T. Senescence induced by altered telomere state, not telomere loss. Science 295, 2446–2449 (2002).
Zellinger, B., Akimcheva, S., Puizina, J., Schirato, M. & Riha, K. Ku suppresses formation of telomeric circles and alternative telomere lengthening in Arabidopsis. Mol. Cell 27, 163–169 (2007).
Acknowledgements
We are grateful to K. Collins (University of California, Berkeley) for generously sharing the pBABE-hTR plasmid and R. Alvarez Rodriguez (the Salk Institute) and members of J.C.I. Belmonte's laboratory (the Salk Institute) for sharing reagents. Packaging plasmids pCMV-gag-pol-PA and pCMV-VSVg were provided by G. Pao (the Salk Institute); pBABE-U3-hTR was provided by K. Collins (University of California, Berkeley); third-generation lentiviral vector was provided by R.A. Rodriguez; packaging plasmids pMDL, Rev and VSVg were provided by O. Singer (the Salk Institute). We thank the members of the Salk Institute's Stem Cell Core for expert advice and members of J.K.'s laboratory for comments. T.R. was supported by the Glenn Center for Research on Aging and CIRM training grant TG2-01158. J.K. is supported by the Salk Institute Cancer Center Core Grant (P30CA014195), the NIH (R01GM087476, R01CA174942), the Donald and Darlene Shiley Chair, the Highland Street Foundation, the Fritz B. Burns Foundation, the Emerald Foundation and the Glenn Center for Aging Research.
Author information
Authors and Affiliations
Contributions
T.R. designed and performed the experiments and wrote the manuscript. C.H. carried out experiments. S.C. provided RHPS4. J.K. designed experiments, supervised the work and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Analysis of extrachromosomal T-circles, 5’ C-rich telomeric DNA and APBs in hESCs.
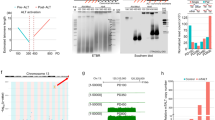
(a) Native and denatured 2D gels of genomic DNA from hES cell lines (H1 and H9), hybridized with radioactively labeled (CCCTAA)5 and (TTAGGG)4 oligonucleotides. Red arrows indicate the arc resulting from T-circles.
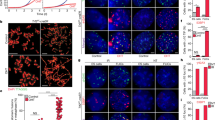
(b) Immunofluorescence for PML (red) and TRF2 (green) in hESCs (H1, H9 and HUES6). U2OS was included as a positive control for APB. DNA was stained with DAPI (blue). Scale bar, 10 μm.
Supplementary Figure 2 Self-renewal and differentiation potential of hESCs with elongated telomeres.
(a) Relative gene expression analysis of the indicated genes following the transduction of vector control or hTR. Mean values represent fold change compared to parental HUES6, n=3. Error bars represent s.e.m. ***P <0.001 (two-tailed Student’s t-test).
(b) TRAP assay of untreated, vector control and hTR transduced HUES6 cells. Quantification of telomerase activity relative to untreated cells is shown on the right, n=3. Data represent means ± s.d. **P <0.01 (two-tailed Student’s t-test).
(c) Expression of pluripotent markers in HUES6 with elongated telomeres. Immunofluorescence analysis of Sox2 (red), SSEA-4 (green), top panels; Oct4 (red), TRA-1-60 (green), bottom panels of parental, vector control and hTR expressing HUES6 cells. DNA is stained with DAPI. Scale bar, 20 μm.
(d,e) Embryoid body (EB)-mediated differentiation assay in HUES6, vector and hTR cells.
(d) Gene expression analysis of differentiation markers for the mesodermal (ACTA2, MSX1), endodermal (GATA6, AFP, ALB, FOXA2) and ectodermal (PAX6, beta-III tubulin, FGF5) germ layers in the indicated cells. Data represent means ± s.e.m., n=3.
(e) Immunofluorescence analysis of the three embryonic germ layers, AFP (endoderm), SMA (mesoderm) and TUJ1 (ectoderm) 14 days following differentiation of HUES6, vector and hTR cells. DNA was stained with DAPI (blue). Scale bar, 40 μm.
Supplementary Figure 3 IMR90 iPSC lines demonstrate pluripotency in vitro.
(a) Karyotype analysis demonstrating that IMR90-iPS cell lines maintain normal chromosomal integrity.
(b) Immunofluorescence analysis of pluripotent markers in IMR90-iPSCs. Cells were stained for Sox2 (red) and TRA-1-60 (green) (left panels) or Oct4 (red) and SSEA-4 (green) (right panels) and DAPI (blue). Scale bar, 20 μm.
(c) Gene expression analysis of pluripotent markers (Nanog, Oct4, Sox2) in IMR90-iPSCs and EB derived IMR90-iPSCs. Data represent relative expression to hESC (HUES6) of three independent experiments (mean ± s.d).
(d) IMR90-iPSCs were used in embryoid body (EB)-mediated differentiation assays and stained by immunofluorescence for AFP (endoderm), SMA (mesoderm), TUJ1 (ectoderm) and Cdx2 (trophectodermal) markers of the embryonic germ layers. DNA was stained with DAPI. Scale bar, 40 μm.
(e) Gene expression analysis of differentiation markers for the mesodermal (ACTA2, MSX1), endodermal (GATA6, AFP, ALB, FOXA2) and ectodermal (PAX6, beta-III tubulin, FGF5) germ layers in EB from IMR90-iPSCs. Data represent relative expression to EB derived from hESCs (HUES6), n=3 (mean ± s.e.m.).
Supplementary Figure 4 Extrachromosomal T-circle formation and telomere elongation is induced after cell reprogramming.
(a) T-circle assay from IMR90, IMR90-iPSCs and HUES6 cells. The presence or absence of ϕ29 DNA polymerase is indicated. Black arrow indicates T-circle products.
(b) Absence of APBs in IMR90-iPSCs. Immunofluorescence for PML (red) and TRF2 (green) in IMR90-iPSCs. DNA was stained with DAPI. Scale bar, 10 μm.
(c) Gene expression analysis of TERT and pluripotent markers (Nanog, Sox2) in IMR90-iPSCs, HUVEC-iPSCs and Keratinocytes-iPSCs. Data represent mean ± s.e.m. relative expression to hESC (HUES6), n=3.
(d) TRF analysis of IMR90-iPSCs, HUVEC-iPSCs, Keratinocytes-iPSCs and HUES6 cells.
Supplementary Figure 5 Telomere elongation in IMR90 cells induces accumulation of ECTRs.
(a) TRAP assay of untreated, vector control and hTERT transduced IMR90 cells.
(b) TRF analysis of vector control and hTERT transduced IMR90 cells at different population doublings (PD) following transduction.
(c) T-circle assay from IMR90 cells transduced as in (a). The presence or absence of ϕ29 DNA polymerase is indicated. Black arrow indicates T-circle products.
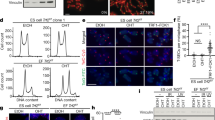
(d) C-circle assay from untreated, vector control and hTERT transduced IMR90 cells at PD40 and PD135. Quantification of C-circle levels (bottom) is shown relative to untreated IMR90 cells. Data represent means ± s.e.m of three experiments. *P <0.05; ns, not significant (two-tailed Student’s t-test).
Supplementary Figure 6 Replication stress induced in HUES6+hTR cells.
(a) HUES6+hTR cells were treated with 0.2 mM or 3 mM hydroxyurea (HU), 5 μM aphidicolin (Aph) and 0.5 μM RHPS4 for 24 hours, and cell cycle profiles were analyzed by flow cytometry. Untreated and DMSO treated cells were used as a control. The percentages of cells in G1, S and G2 phases are shown in the upper left corner in each plot. Cells were incubated with 10 μM BrdU for 30 minutes prior to the analysis. BrdU incorporation was analyzed by flow cytometry and represented in each condition. A representative experiment quantifying ≥ 10,000 cells per condition is shown.
(b) Relative gene expression of the indicated genes in HUES6+hTR cells 4 days after transfection of the indicated siRNAs. Values of gene expression are shown as fold change relative to the value of the sample control. Data represent mean ± s.e.m., n=3.
(c) Relative gene expression of the indicated genes in HUES6+vector or HUES6+hTR cells 10 days post-transduction with shScramble or shTERT. Values of gene expression are shown as fold change relative to the value of the sample control. Data represent mean ± s.e.m., n=3.
(d) Quantification of metaphase chromosomes with MTS in cells depicted in (c). Data represent means ± s.e.m of three independent experiments; ns, not significant (two-tailed Student’s t-test).
(e) TRF1 and TRF2 ChIP of parental, vector or hTR transduced HUES6 cells. Quantification of the ChIP assay relative to untreated cells is shown on the right.
Supplementary Figure 7 Contribution of XRCC3 and Nbs1 to ECTR formation.
(a) Expression of XRCC3 as fold change relative to untreated cells (HUES6) at 72 h following siControl or siXRCC3 transfection. Data represent mean ± s.d. of three independent experiments **P <0.01 (two-tailed Student’s t-test).
(b) T-circle assay from untreated, siControl and siXRCC3 HUES6 cells depicted in (a). The presence or absence of ϕ29 DNA polymerase is indicated. The quantification of T-circle levels relative to untreated cells is shown on the right. Data represent means ± s.e.m. of three independent experiments; ns, not significant (two-tailed Student’s t-test).
(c) Expression of Nbs1 as fold change relative to untreated cells (HUES6) 5 days post-transduction with shScramble or shRNAs (sh1 and sh2) against Nbs1. Data represent mean ± s.d. of three independent experiments **P <0.01, *P <0.05 (two-tailed Student’s t-test).
(d) C-circle assay from untreated, siControl or siXRCC3 HUES6 cells. Quantification of C-circle levels (bottom) is shown relative to untreated HUES6. Data represent means ± s.d of four experiments; ns, not significant (two-tailed Student’s t-test).
(e) C-circle analysis of HUES6 cells following the knockdown of Nbs1 using two different shRNAs. Quantification represents mean values relative to untreated HUES6 cells from three experiments. Error bars represent s.d., ns, not significant (two-tailed Student’s t-test).
(f) Expression of Nbs1, XRCC3 and Oct4 in XRCC3 and Nbs1 knockdown cells. Quantification represents relative gene expression to control depleted shScramble cells normalized against GAPDH expression from two experiments (mean ± s.d.).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Table 1 (PDF 1964 kb)
Supplementary Data Set 1
Uncropped gels and blots (PDF 3197 kb)
Rights and permissions
About this article
Cite this article
Rivera, T., Haggblom, C., Cosconati, S. et al. A balance between elongation and trimming regulates telomere stability in stem cells. Nat Struct Mol Biol 24, 30–39 (2017). https://doi.org/10.1038/nsmb.3335
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3335
This article is cited by
-
Impacts of radiation exposure, hindlimb unloading, and recovery on murine skeletal muscle cell telomere length
npj Microgravity (2023)
-
The complementarity of DDR, nucleic acids and anti-tumour immunity
Nature (2023)
-
Gene amplifications and extrachromosomal circular DNAs: function and biogenesis
Molecular Biology Reports (2023)
-
Connecting telomere maintenance and regulation to the developmental origin and differentiation states of neuroblastoma tumor cells
Journal of Hematology & Oncology (2022)
-
Extrachromosomal circular DNA: biogenesis, structure, functions and diseases
Signal Transduction and Targeted Therapy (2022)