Abstract
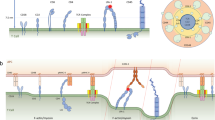
CD28 provides an essential costimulatory signal for T cell activation, and its function is critical in antitumor immunity. However, the molecular mechanism of CD28 transmembrane signaling remains elusive. Here we show that the conformation and signaling of CD28 are regulated by two counteractive charged factors, acidic phospholipids and Ca2+ ions. NMR spectroscopy analyses showed that acidic phospholipids can sequester CD28 signaling motifs within the membrane, thereby limiting CD28 basal signaling. T cell receptor (TCR) activation induced an increase in the local Ca2+ concentration around CD28, and Ca2+ directly disrupted CD28-lipid interaction, leading to opening and signaling of CD28. We observed that the TCR, Ca2+, and CD28 together form a dual-positive-feedback circuit that substantially amplifies T cell signaling and thus increases antigen sensitivity. This work unravels a new regulatory mechanism for CD28 signaling and thus contributes to the understanding of the dependence of costimulation signaling on TCR signaling and the high sensitivity of T cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Esensten, J.H., Helou, Y.A., Chopra, G., Weiss, A. & Bluestone, J.A. CD28 costimulation: from mechanism to therapy. Immunity 44, 973–988 (2016).
Chen, L. & Flies, D.B. Molecular mechanisms of T cell co-stimulation and co-inhibition. Nat. Rev. Immunol. 13, 227–242 (2013).
Rudd, C.E., Taylor, A. & Schneider, H. CD28 and CTLA-4 coreceptor expression and signal transduction. Immunol. Rev. 229, 12–26 (2009).
Boomer, J.S. & Green, J.M. An enigmatic tail of CD28 signaling. Cold Spring Harb. Perspect. Biol. 2, a002436 (2010).
Salomon, B. et al. B7/CD28 costimulation is essential for the homeostasis of the CD4+CD25+ immunoregulatory T cells that control autoimmune diabetes. Immunity 12, 431–440 (2000).
Tai, X., Cowan, M., Feigenbaum, L. & Singer, A. CD28 costimulation of developing thymocytes induces Foxp3 expression and regulatory T cell differentiation independently of interleukin 2. Nat. Immunol. 6, 152–162 (2005).
Zhang, R., Borges, C.M., Fan, M.Y., Harris, J.E. & Turka, L.A. Requirement for CD28 in effector regulatory T cell differentiation, CCR6 induction, and skin homing. J. Immunol. 195, 4154–4161 (2015).
Linterman, M.A. et al. CD28 expression is required after T cell priming for helper T cell responses and protective immunity to infection. eLife 3, e03180 (2014).
Liang, Y. et al. The lymphoid lineage-specific actin-uncapping protein Rltpr is essential for costimulation via CD28 and the development of regulatory T cells. Nat. Immunol. 14, 858–866 (2013).
Jain, N. et al. CD28 and ITK signals regulate autoreactive T cell trafficking. Nat. Med. 19, 1632–1637 (2013).
Martinez-Llordella, M. et al. CD28-inducible transcription factor DEC1 is required for efficient autoreactive CD4+ T cell response. J. Exp. Med. 210, 1603–1619 (2013).
Diekmann, F. Immunosuppressive TCR phosphorylation minimization with mTOR inhibitors and belatacept. Transpl. Int. 28, 921–927 (2015).
Westhovens, R. et al. Clinical efficacy and safety of abatacept in methotrexate-naive patients with early rheumatoid arthritis and poor prognostic factors. Ann. Rheum. Dis. 68, 1870–1877 (2009).
van der Stegen, S.J., Hamieh, M. & Sadelain, M. The pharmacology of second-generation chimeric antigen receptors. Nat. Rev. Drug Discov. 14, 499–509 (2015).
Kamphorst, A.O. et al. Rescue of exhausted CD8 T cells by PD-1-targeted therapies is CD28-dependent. Science 355, 1423–1427 (2017).
Hui, E. et al. T cell costimulatory receptor CD28 is a primary target for PD-1-mediated inhibition. Science 355, 1428–1433 (2017).
Tian, R. et al. Combinatorial proteomic analysis of intercellular signaling applied to the CD28 T-cell costimulatory receptor. Proc. Natl. Acad. Sci. USA 112, E1594–E1603 (2015).
Dodson, L.F. et al. Targeted knock-in mice expressing mutations of CD28 reveal an essential pathway for costimulation. Mol. Cell. Biol. 29, 3710–3721 (2009).
Kong, K.F. et al. A motif in the V3 domain of the kinase PKC-θ determines its localization in the immunological synapse and functions in T cells via association with CD28. Nat. Immunol. 12, 1105–1112 (2011).
Dobbins, J. et al. Binding of the cytoplasmic domain of CD28 to the plasma membrane inhibits Lck recruitment and signaling. Sci. Signal. 9, ra75 (2016).
Diehn, M. et al. Genomic expression programs and the integration of the CD28 costimulatory signal in T cell activation. Proc. Natl. Acad. Sci. USA 99, 11796–11801 (2002).
Riley, J.L. et al. Modulation of TCR-induced transcriptional profiles by ligation of CD28, ICOS, and CTLA-4 receptors. Proc. Natl. Acad. Sci. USA 99, 11790–11795 (2002).
Li, L., Shi, X., Guo, X., Li, H. & Xu, C. Ionic protein-lipid interaction at the plasma membrane: what can the charge do? Trends Biochem. Sci. 39, 130–140 (2014).
Shi, X. et al. Ca2+ regulates T-cell receptor activation by modulating the charge property of lipids. Nature 493, 111–115 (2013).
Xu, C. et al. Regulation of T cell receptor activation by dynamic membrane binding of the CD3ɛ cytoplasmic tyrosine-based motif. Cell 135, 702–713 (2008).
Zhang, H., Cordoba, S.P., Dushek, O. & van der Merwe, P.A. Basic residues in the T-cell receptor ζ cytoplasmic domain mediate membrane association and modulate signaling. Proc. Natl. Acad. Sci. USA 108, 19323–19328 (2011).
DeFord-Watts, L.M. et al. The CD3 ζ subunit contains a phosphoinositide-binding motif that is required for the stable accumulation of TCR-CD3 complex at the immunological synapse. J. Immunol. 186, 6839–6847 (2011).
Deford-Watts, L.M. et al. The cytoplasmic tail of the T cell receptor CD3 ɛ subunit contains a phospholipid-binding motif that regulates T cell functions. J. Immunol. 183, 1055–1064 (2009).
Aivazian, D. & Stern, L.J. Phosphorylation of T cell receptor ζ is regulated by a lipid-dependent folding transition. Nat. Struct. Biol. 7, 1023–1026 (2000).
Gagnon, E., Schubert, D.A., Gordo, S., Chu, H.H. & Wucherpfennig, K.W. Local changes in lipid environment of TCR microclusters regulate membrane binding by the CD3ɛ cytoplasmic domain. J. Exp. Med. 209, 2423–2439 (2012).
Bettini, M.L. et al. Membrane association of the CD3ɛ signaling domain is required for optimal T cell development and function. J. Immunol. 193, 258–267 (2014).
Schwieters, C.D., Kuszewski, J.J., Tjandra, N. & Clore, G.M. The Xplor-NIH NMR molecular structure determination package. J. Magn. Reson. 160, 65–73 (2003).
Chen, X. et al. Acidic phospholipids govern the enhanced activation of IgG-B cell receptor. Nat. Commun. 6, 8552 (2015).
Wang, Y.H. et al. Divalent cation-induced cluster formation by polyphosphoinositides in model membranes. J. Am. Chem. Soc. 134, 3387–3395 (2012).
Yokosuka, T. et al. Spatiotemporal regulation of T cell costimulation by TCR-CD28 microclusters and protein kinase C θ translocation. Immunity 29, 589–601 (2008).
Chen, T.W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Michel, F., Attal-Bonnefoy, G., Mangino, G., Mise-Omata, S. & Acuto, O. CD28 as a molecular amplifier extending TCR ligation and signaling capabilities. Immunity 15, 935–945 (2001).
Oh-Hora, M. et al. Dual functions for the endoplasmic reticulum calcium sensors STIM1 and STIM2 in T cell activation and tolerance. Nat. Immunol. 9, 432–443 (2008).
Parekh, A.B. Ca2+ microdomains near plasma membrane Ca2+ channels: impact on cell function. J. Physiol. (Lond.) 586, 3043–3054 (2008).
Ma, H.T. et al. Canonical transient receptor potential 5 channel in conjunction with Orai1 and STIM1 allows Sr2+ entry, optimal influx of Ca2+, and degranulation in a rat mast cell line. J. Immunol. 180, 2233–2239 (2008).
Collins, A.V. et al. The interaction properties of costimulatory molecules revisited. Immunity 17, 201–210 (2002).
Weiss, J.M., Morgan, P.H., Lutz, M.W. & Kenakin, T.P. The cubic ternary complex receptor-occupancy model. III. Resurrecting efficacy. J. Theor. Biol. 181, 381–397 (1996).
Lever, M., Maini, P.K., van der Merwe, P.A. & Dushek, O. Phenotypic models of T cell activation. Nat. Rev. Immunol. 14, 619–629 (2014).
Chakraborty, A.K. & Das, J. Pairing computation with experimentation: a powerful coupling for understanding T cell signalling. Nat. Rev. Immunol. 10, 59–71 (2010).
Chakraborty, A.K. & Weiss, A. Insights into the initiation of TCR signaling. Nat. Immunol. 15, 798–807 (2014).
van den Bogaart, G. et al. Membrane protein sequestering by ionic protein-lipid interactions. Nature 479, 552–555 (2011).
Bashour, K.T. et al. Cross talk between CD3 and CD28 is spatially modulated by protein lateral mobility. Mol. Cell. Biol. 34, 955–964 (2014).
Marinari, B., Costanzo, A., Marzano, V., Piccolella, E. & Tuosto, L. CD28 delivers a unique signal leading to the selective recruitment of RelA and p52 NF-κB subunits on IL-8 and Bcl-xL gene promoters. Proc. Natl. Acad. Sci. USA 101, 6098–6103 (2004).
Porciello, N. & Tuosto, L. CD28 costimulatory signals in T lymphocyte activation: emerging functions beyond a qualitative and quantitative support to TCR signalling. Cytokine Growth Factor Rev. 28, 11–19 (2016).
Thaker, Y.R., Schneider, H. & Rudd, C.E. TCR and CD28 activate the transcription factor NF-κB in T-cells via distinct adaptor signaling complexes. Immunol. Lett. 163, 113–119 (2015).
Takeda, K. et al. CD28 stimulation triggers NF-κB activation through the CARMA1-PKCθ-Grb2/Gads axis. Int. Immunol. 20, 1507–1515 (2008).
Camperio, C. et al. CD28 ligation in the absence of TCR stimulation up-regulates IL-17A and pro-inflammatory cytokines in relapsing-remitting multiple sclerosis T lymphocytes. Immunol. Lett. 158, 134–142 (2014).
Muscolini, M., Sajeva, A., Caristi, S. & Tuosto, L. A novel association between filamin A and NF-κB inducing kinase couples CD28 to inhibitor of NF-κB kinase α and NF-κB activation. Immunol. Lett. 136, 203–212 (2011).
Piccolella, E. et al. Vav-1 and the IKKα subunit of IκB kinase functionally associate to induce NF-κB activation in response to CD28 engagement. J. Immunol. 170, 2895–2903 (2003).
Wertek, F. & Xu, C. Digital response in T cells: to be or not to be. Cell Res. 24, 265–266 (2014).
Huang, J. et al. A single peptide-major histocompatibility complex ligand triggers digital cytokine secretion in CD4+ T cells. Immunity 39, 846–857 (2013).
O'Donoghue, G.P., Pielak, R.M., Smoligovets, A.A., Lin, J.J. & Groves, J.T. Direct single molecule measurement of TCR triggering by agonist pMHC in living primary T cells. eLife 2, e00778 (2013).
Davis, M.M. et al. T cells as a self-referential, sensory organ. Annu. Rev. Immunol. 25, 681–695 (2007).
Feinerman, O., Veiga, J., Dorfman, J.R., Germain, R.N. & Altan-Bonnet, G. Variability and robustness in T cell activation from regulated heterogeneity in protein levels. Science 321, 1081–1084 (2008).
Das, J. et al. Digital signaling and hysteresis characterize Ras activation in lymphoid cells. Cell 136, 337–351 (2009).
Cordoba, S.P. et al. The large ectodomains of CD45 and CD148 regulate their segregation from and inhibition of ligated T-cell receptor. Blood 121, 4295–4302 (2013).
James, J.R. & Vale, R.D. Biophysical mechanism of T-cell receptor triggering in a reconstituted system. Nature 487, 64–69 (2012).
Tan, Y.X. et al. Inhibition of the kinase Csk in thymocytes reveals a requirement for actin remodeling in the initiation of full TCR signaling. Nat. Immunol. 15, 186–194 (2014).
Schmitt, C.A., McCurrach, M.E., de Stanchina, E., Wallace-Brodeur, R.R. & Lowe, S.W. INK4a/ARF mutations accelerate lymphomagenesis and promote chemoresistance by disabling p53. Genes Dev. 13, 2670–2677 (1999).
Nugent, T. & Jones, D.T. Transmembrane protein topology prediction using support vector machines. BMC Bioinformatics 10, 159 (2009).
Li, F.Y. et al. Second messenger role for Mg2+ revealed by human T-cell immunodeficiency. Nature 475, 471–476 (2011).
Hogan, P.G., Lewis, R.S. & Rao, A. Molecular basis of calcium signaling in lymphocytes: STIM and ORAI. Annu. Rev. Immunol. 28, 491–533 (2010).
Szymczak, A.L. et al. Correction of multi-gene deficiency in vivo using a single 'self-cleaving' 2A peptide-based retroviral vector. Nat. Biotechnol. 22, 589–594 (2004).
Johnson, B.A. & Blevins, R.A. NMR View: a computer program for the visualization and analysis of NMR data. J. Biomol. NMR 4, 603–614 (1994).
Kobayashi, N. et al. KUJIRA, a package of integrated modules for systematic and interactive analysis of NMR data directed to high-throughput NMR structure studies. J. Biomol. NMR 39, 31–52 (2007).
Chylek, L.A. et al. Rule-based modeling: a computational approach for studying biomolecular site dynamics in cell signaling systems. Wiley Interdiscip. Rev. Syst. Biol. Med. 6, 13–36 (2014).
Mukhopadhyay, H., Cordoba, S.P., Maini, P.K., van der Merwe, P.A. & Dushek, O. Systems model of T cell receptor proximal signaling reveals emergent ultrasensitivity. PLoS Comput. Biol. 9, e1003004 (2013).
Huse, M. et al. Spatial and temporal dynamics of T cell receptor signaling with a photoactivatable agonist. Immunity 27, 76–88 (2007).
Acknowledgements
The NMR spectroscopy and imaging experiments were performed at the National Center for Protein Science Shanghai, Core Facility for Molecular Biology of SIBCB, and Core Facility for Cell Biology of SIBCB. We thank Z. Liu (NMR section, NCPSS) for help in setting up filter NOESY experiments. We thank Y. Yu (Integrated Laser Microscopy section, NCPSS) for help in setting up imaging experiments. We thank K. Wucherpfennig and E. Gagnon for helpful discussions. The CD28-deficient cell line was a generous gift from O. Acuto (Oxford University, Oxford, UK). This work was supported by NSFC (grants 31370860, 31425009, 31530022, and 31621003 to C.X.; grants 31470734 and 31670751 to H.L.), CAS (Strategic Priority Research Program grant XDB08020100 to C.X.). MOST (grant 2014CB541903 to H.L.), and the Wellcome Trust and the Royal Society (Sir Henry Dale Fellowship, grant 098363 to O.D.). N.T acknowledges support from Systems Biology DTC supported by the Engineering and Physical Science Research Council (UK).
Author information
Authors and Affiliations
Contributions
C.X. conceived the project and wrote the manuscript. C.X., H.L., O.D., and H.S. designed the experiments. W.P. and H.L. performed the NMR experiments. W.Y., W.P., and S.J. performed the biochemical and immunological experiments. S.C. and W.Y. performed the imaging experiments. N.T. and O.D. performed the mathematical modeling. F.X., W.W., and S.C. performed the bioinformatics analysis. H.W., X. Liu and H. Ji provided extensive discussion. Z.P. and M.X. helped with the protein purification. X. Li and H. Jiang generated BBN-14 cells. All authors contributed to revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 The pI-value distribution of single-pass membrane proteins at the plasma membrane in immune cells and in total cells.
The PI values were measured for the first 10 amino acids in the cytoplasmic domains of human single-pass transmembrane proteins that localized at the plasma membrane. Source data are available in Source Data 3.
Supplementary Figure 2 CD28 cytoplasmic domain specifically binds to acidic phospholipids.
(a) Alignment of CD28CD from different species. Most residues are highly conserved, especially the two important YxxM and PYAP signaling motifs (marked in red) and basic residues (marked in green). The residue conservation is visualized by the sequence logo of CD28CD generated from the Skylign web server http://www.skylign.org.
(b) Net charges of human and mouse CD28 cytoplasmic domains at pH 7.0.
(c) The protein sequences of mouse CD28CD and three mutants used in (d-g). Two polybasic regions (PBR) are marked in red, and mutant sites are marked in blue.
(d-g) Binding of CD28CD WT and mutants to acidic phospholipids was measured by the equilibrium-based microdialysis assay.
(d) A cartoon illustration of the equilibrium-based microdialysis assay. See Online Methods for details.
(e) Binding of CD28CD WT to different acidic phospholipids measured by the microdialysis assay. LUVs in the experimental chamber were composed of either 100% of the acidic phospholipids POPG, POPS, PI, PA, or a lipid mixture (40% POPC, 20% POPS, 20% POPE, 10% POPG and 10% PI). LUVs in the control chamber were composed of 100% of the zwitterionic lipid POPC. CD28CD-Alexa 488 was used at a concentration of 10 nM and LUVs at 5 mM. Two to three independent samples were measured for each condition and the results are plotted as the percentage of the fluorescence intensity in the experimental and control chambers.
(f-g) Binding of CD28CD WT and mutants to acidic phospholipids measured by the equilibrium-based microdialysis assay. Two to three independent samples were measured for each condition and the results are plotted as the percentage of the fluorescence intensity in the experimental and control chambers (F), and are further converted to the binding efficiencies of the mutants to POPG (G). Data were analyzed by unpaired t-test. **P < 0.01; ***P < 0.001; ns: no significant difference.
Data are representative of three (e) or seven (f, g) independent experiments. The center value and error bar in e-g denote mean and range. Source data for e-g are available in Source Data 4.
Supplementary Figure 3 Measurement of membrane binding of CD28CD by the aromatic fluorescence emission (AFE) assay.
(a) Incubation of 2 μM CD28CD with 0.2 mM acidic POPG LUVs led to significant increase of AFE value but such an increase was not observed in that with 0.2 mM zwitterionic POPC LUVs.
(b-c) Titration of acidic POPG LUVs with the indicated concentrations into 2 μM CD28CD sample led to the gradient increase of AFE value (at 310nm).
For each condition, three independent samples were measured in one experiment. Data are representative of three independent experiments. Unpaired t-test was used for comparing each two groups. One-Way ANOVA was used to test whether POPG treatment could cause significant change of the AFE value, P < 0.0001 (C).
The center value and error bar in c denote mean and s.e.m.. Source data are available in Source Data 5.
Supplementary Figure 4 Membrane binding of Notch1 juxtamembrane polybasic region.
(a) Sequence information and charge property of human Notch1 juxtamembrane domain.
(b) Incubation of 2 μM Notch1_JM with 0.1 mM acidic POPG LUVs led to significant increase of AFE value but such an increase was not observed in that with 0.1 mM zwitterionic POPC LUVs.
(c-d) Titration of acidic POPG LUVs with the indicated concentrations into 2 μM Notch1_JM sample led to the gradient increase of AFE value (at approximate 350nm). Three independent samples were measured for each condition. Unpaired t-test was used for the statistical analysis of each two groups. One-Way ANOVA was used to test whether POPG treatment could cause significant change of the AFE value, P < 0.0001.
(e-f) The de-quenching FRET was used to measure the interaction of Notch1 juxtamembrane polybasic region with the plasma membrane in live Jurkat T cells. Representative cells are shown in e. Data in f are representative of three independent experiments. Bars in e represent 2 μm. N = 31, 32, 35 for the three conditions (from left to right). Each dot represents the FRET value from one individual cell. One-Way ANOVA was used to analyze difference among three groups, the p-values of these three panels from left to right were <0.0001, 0.2074 and 0.1555. Unpaired t-test was used to analyze difference between each two groups.
The center value and error bar in d,f denote mean and s.e.m.. Source data for b-d, and f are available in Source Data 6.
Supplementary Figure 5 Intrinsic dynamics of membrane-bound CD28CD.
15N backbone spin relaxation measurements of CD28CD in POPG bicelles. (a) 15N longitudinal relaxation rates (R1), (b) 15N transverse relaxation rates (R2) and (c) the heteronuclear 1H-15N NOEs. Disappeared or overlapped residues are marked by asterisks. Prolines that lack backbone amides are marked by P.
Supplementary Figure 6 Disruption of CD28CD-membrane binding promoted the protein’s basal signaling.
(a) The de-quenching FRET was used to measure the membrane binding of CD28 WT and CD28 linker mutant (n = 21, 38 (from left to right), each dot represents the FRET value from one individual cell).
(b-e) TFP-T2A-HA-CD28 WT or linker mutant construct was transduced into mouse CD28−/− T cells by retrovirus, respectively. After protein translation of the fusion construct, TFP and HA-CD28 were separated due to the self-cleaving property of T2A. Basal CD28 signaling was measured in transduced cells without stimulation. IL-2 production was measured in transduced cells under TCR and/or CD28 stimulation.
(b-c) The cells without stimulation were lysed and the immunoprecipitation assay was performed to detect basal phosphorylation of CD28 and its interactions with signaling proteins. The bands were quantitated by ImageJ. The pCD28/HA, P85/HA, Lck/HA and Grb2/HA ratios were obtained and further normalized to the value of WT condition of each strip. Average results of 4 independent samples are shown in panel C.
(d-e) The cells were stimulated for 4 hours with plated-bound α-CD3 (0.5 μg/ml) alone or α-CD3 (0.5 μg/ml) + α-CD28 (2 μg/ml), and IL-2 production was measured by intracellular staining and flow cytometric analysis. CD4+ TFP+ cells were gated for the analysis of IL-2 level. TFP levels of CD28 WT cells and CD28 linker cells were matched, reflecting the comparable expression level of CD28 WT and CD28 linker. Percentage of IL-2 positive cells and median fluorescence intensity (MFI) of all cells are shown (n = 3).
Data are representative of three (a, c) or four (e) independent experiments, and were analyzed by unpaired t-test. The center value and error bar in a, c and e denote mean and s.e.m.. The original gel image of b can be found in Supplementary Data Set 2. Source data for a, c and e are available in Source Data 7.
Supplementary Figure 7 Ionomycin-induced Ca2+ influx enhances the openness of CD28.
The de-quenching FRET method was used to measure the effect of Ca2+ on CD28CD-membrane binding in live Jurkat T cells. Ionomycin (1 μM) was used to trigger Ca2+ influx in T cells. Ca2+ influx led to the significant decrease of the FRET value of CD28-TFP (a), indicating the dissociation of CD28CD from the membrane. In contrast, the FRET values of the 3aa-TFP and 50aa-TFP control constructs were insensitive to Ca2+ influx (b, c). The FRET efficiency was measured and plotted as mean +/- s.e.m. (n = 12, 15 in (A), n = 17, 16 in (B), n = 22, 19 in (C) (from left to right), each dot represents the FRET value from one individual cell). The surface mTFP level and R18 level were comparable.
Data are representative of two independent experiments. All the data were analyzed by unpaired t-test except the left sub-panels in (a) and (c) where Mann-Whitney test were used because the FERT efficiency data sets of the CD28 group and the 50aa-TFP group do not fit the normal distribution. The center value and error bar denote mean and s.e.m.. Source data are available in Source Data 8.
Supplementary Figure 8 Ca2+ influx enhances the openness and signaling of CD28.
(a-b) To further demonstrate the effect of Ca2+ on CD28CD-membrane binding, we use different concentrations of α-CD3 to generate different levels of Ca2+ influx. We also used BAPTA pre-treatment to chelate the intracellular Ca2+ (a). The de-quenching FRET was used to detect the effect of TCR-induced Ca2+ influx on the CD28CD-membrane interaction. Different doses of α-CD3 induced different levels of Ca2+ influx (a). Treatment of 10 μM BAPTA led to partial chelation of intracellular Ca2+. N = 64, 28, 22, 36 (from left to right) for each condition. Each dot represents the FRET value from one individual cell. Mouse IgG (3 μg/ml) was used as the isotype control antibody for α-CD3.
(c) To exam the effect of Ca2+ influx on CD28 signaling in primary T cell, the mouse CTL were generated and stimulated with α-CD3 (0.5 μg/ml) + α-CD28 (2 μg/ml) with indicated times in the Ca2+/Mg2+-free Ringer’s buffer containing 1 mM Ca2+ or not at 37 oC. After stimulation, cells were lysed for immunoprecipitation and immunoblotting. The bands were quantitated by ImageJ. The pCD28/CD28, P85/CD28, Lck/CD28 and Grb2/CD28 ratios were obtained and further normalized to the value of no stimulation condition of each strip.
Data are representative of three independent experiments. In (b), for the left sub-panel, One-Way ANOVA was used to analyze difference among four groups (P<0.0001) and unpaired t-test was used to analyze difference between two groups. For the middle and right sub-panels, the R18 level of the α-CD3 (3μg/ml) group and the mTFP level of the isotype control group do not fit the normal distribution. To compare each two groups, Mann Whitney test was used for data sets that do not fit the normal distribution and unpaired t-test was used for the rest. The center value and error bar in b denote mean and s.e.m.. The original gel image of b can be found in Supplementary Data Set 2. Source data for b are available in Source Data 9.
Supplementary Figure 9 CD28 ligation caused the dissociation of CD28 cytoplasmic domain from the membrane but was insufficient to induce CD28 phosphorylation.
(a) The de-quenching FRET method was used to measure whether CD28 ligation could induce CD28 cytoplasmic domain to dissociate from the membrane in live Jurkat T cells. The stimulating buffer was Ca2+/Mg2+-free Ringer’s buffer. Compared with mock treatment (no antibody), addition of 5 μg/ml α-CD28 induced dissociation of CD28CD from the membrane. To test whether this induction was dependent on the phosphorylation of CD28, 100 μM PP2 was pre-incubated with T cells for 10 min to inhibit Src family kinase and then the cells were stimulated by 5 μg/ml a-CD28. The result indicated CD28 phosphorylation was not required for the dissociation of CD28CD from the membrane. N = 22, 11, 12 for each condition (from left to right). Each dot represents the FRET value from one individual cell. Data are representative of three independent experiments. Unpaired t-test was used to analyze difference between two groups.
(b) CD28-deficient Jurkat T cells expressing HA-mCD28 were stimulated by 1 μg/ml a-CD3 alone, 2 μg/ml a-CD28 alone or 2 μg/ml a-CD28 and 1 μg/ml a-CD3 in normal Ringer’s buffer (containing 1 mM Ca2+) for the indicated time at 37 oC. Either CD3 ligation alone or CD28 ligation alone could not trigger CD28 phosphorylation, but CD28 ligation and TCR ligation together could trigger CD28 phosphorylation. The bands were quantitated by Image J. The pCD28 band intensity was divided by the corresponding CD28 band intensity to obtain the pCD28/CD28 ratio, which were further normalized to the value of 0 min of α-CD3 condition.
The center value and error bar in a denote mean and s.e.m.. The original gel image of b can be found in Supplementary Data Set 2. Source data for a are available in Source Data 10.
Supplementary Figure 10 Low surface expression of CD28 ligands in tumor cells.
CD80 and CD86 surface levels were detected on CT26 murine colon carcinoma cell line, EL4 murine lymphoma cell line, Eμ-Myc p19Afr−/− mouse B cell lymphoma cell line and BBN-14 murine bladder carcinoma cell line. Activated dendritic cell (activated by 100 ng/ml LPS at 37°C for 24 h) was used as a positive control.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Table 1 and Supplementary Note 1 (PDF 2880 kb)
Supplementary Data Set 1
Single-pass transmembrane proteins in immune cells that contain intracellular juxtamembrane polybasic regions (length of 10 amino acids; pI ≥ 11). (XLSX 22 kb)
Supplementary Data Set 2
Original gel images for Figure 5e and Supplementary Figures 6b, 8c, and 9b. (PDF 35585 kb)
Source data
Rights and permissions
About this article
Cite this article
Yang, W., Pan, W., Chen, S. et al. Dynamic regulation of CD28 conformation and signaling by charged lipids and ions. Nat Struct Mol Biol 24, 1081–1092 (2017). https://doi.org/10.1038/nsmb.3489
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3489
This article is cited by
-
Single cell RNA sequencing reveals differentiation related genes with drawing implications in predicting prognosis and immunotherapy response in gliomas
Scientific Reports (2022)
-
Identification of a differentiation-related prognostic nomogram based on single-cell RNA sequencing in clear cell renal cell carcinoma
Scientific Reports (2022)
-
Identification of stem cell-related subtypes and risk scoring for gastric cancer based on stem genomic profiling
Stem Cell Research & Therapy (2021)
-
PD-L1 degradation is regulated by electrostatic membrane association of its cytoplasmic domain
Nature Communications (2021)
-
Structural understanding of T cell receptor triggering
Cellular & Molecular Immunology (2020)