Abstract
Histones are highly covalently modified, but the functions of many of these modifications remain unknown. In particular, it is unclear how histone marks are coupled to cellular metabolism and how this coupling affects chromatin architecture. We identified histone H3 Lys14 (H3K14) as a site of propionylation and butyrylation in vivo and carried out the first systematic characterization of histone propionylation. We found that H3K14pr and H3K14bu are deposited by histone acetyltransferases, are preferentially enriched at promoters of active genes and are recognized by acylation-state-specific reader proteins. In agreement with these findings, propionyl-CoA was able to stimulate transcription in an in vitro transcription system. Notably, genome-wide H3 acylation profiles were redefined following changes to the metabolic state, and deletion of the metabolic enzyme propionyl-CoA carboxylase altered global histone propionylation levels. We propose that histone propionylation, acetylation and butyrylation may act in combination to promote high transcriptional output and to couple cellular metabolism with chromatin structure and function.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Huang, H., Sabari, B.R., Garcia, B.A., Allis, C.D. & Zhao, Y. SnapShot: histone modifications. Cell 159, 458–458.e1 (2014).
Hanover, J.A., Krause, M.W. & Love, D.C. Bittersweet memories: linking metabolism to epigenetics through O-GlcNAcylation. Nat. Rev. Mol. Cell Biol 13, 312–321 (2012).
Bannister, A.J. & Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 21, 381–395 (2011).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Chen, Y. et al. Lysine propionylation and butyrylation are novel post-translational modifications in histones. Mol. Cell. Proteomics 6, 812–819 (2007).
Tan, M. et al. Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 146, 1016–1028 (2011).
Dai, L. et al. Lysine 2-hydroxyisobutyrylation is a widely distributed active histone mark. Nat. Chem. Biol. 10, 365–370 (2014).
Xie, Z. et al. Lysine succinylation and lysine malonylation in histones. Mol. Cell. Proteomics 11, 100–107 (2012).
Rousseaux, S. & Khochbin, S. Histone acylation beyond acetylation: terra incognita in chromatin biology. Cell J. 17, 1–6 (2015).
Vollmuth, F. & Geyer, M. Interaction of propionylated and butyrylated histone H3 lysine marks with Brd4 bromodomains. Angew. Chem. Int. Edn Engl. 49, 6768–6772 (2010).
Lin, H., Su, X. & He, B. Protein lysine acylation and cysteine succination by intermediates of energy metabolism. ACS Chem. Biol. 7, 947–960 (2012).
Kaelin, W.G. Jr. & McKnight, S.L. Influence of metabolism on epigenetics and disease. Cell 153, 56–69 (2013).
Wellen, K.E. et al. ATP-citrate lyase links cellular metabolism to histone acetylation. Science 324, 1076–1080 (2009).
Roth, S.Y., Denu, J.M. & Allis, C.D. Histone acetyltransferases. Annu. Rev. Biochem. 70, 81–120 (2001).
Candido, E.P., Reeves, R. & Davie, J.R. Sodium butyrate inhibits histone deacetylation in cultured cells. Cell 14, 105–113 (1978).
Strahl, B.D. & Allis, C.D. The language of covalent histone modifications. Nature 403, 41–45 (2000).
Zhou, V.W., Goren, A. & Bernstein, B.E. Charting histone modifications and the functional organization of mammalian genomes. Nat. Rev. Genet. 12, 7–18 (2011).
Kaczmarska, Z. et al. Structure of p300 in complex with acyl-CoA variants. Nat. Chem. Biol. 13, 21–29 (2017).
Ringel, A.E. & Wolberger, C. Structural basis for acyl-group discrimination by human Gcn5L2. Acta Crystallogr. D Struct. Biol. 72, 841–848 (2016).
Albaugh, B.N., Arnold, K.M. & Denu, J.M. KAT(ching) metabolism by the tail: insight into the links between lysine acetyltransferases and metabolism. ChemBioChem 12, 290–298 (2011).
Leemhuis, H., Packman, L.C., Nightingale, K.P. & Hollfelder, F. The human histone acetyltransferase P/CAF is a promiscuous histone propionyltransferase. ChemBioChem 9, 499–503 (2008).
Jin, Q. et al. Distinct roles of GCN5/PCAF-mediated H3K9ac and CBP/p300-mediated H3K18/27ac in nuclear receptor transactivation. EMBO J. 30, 249–262 (2011).
Bonnet, J. et al. The SAGA coactivator complex acts on the whole transcribed genome and is required for RNA polymerase II transcription. Genes Dev. 28, 1999–2012 (2014).
Pougovkina, O., Te Brinke, H., Wanders, R.J.A., Houten, S.M. & de Boer, V.C. Aberrant protein acylation is a common observation in inborn errors of acyl-CoA metabolism. J. Inherit. Metab. Dis. 37, 709–714 (2014).
Palladino, A.A., Chen, J., Kallish, S., Stanley, C.A. & Bennett, M.J. Measurement of tissue acyl-CoAs using flow-injection tandem mass spectrometry: acyl-CoA profiles in short-chain fatty acid oxidation defects. Mol. Genet. Metab. 107, 679–683 (2012).
Guenzel, A.J. et al. Generation of a hypomorphic model of propionic acidemia amenable to gene therapy testing. Mol. Ther. 21, 1316–1323 (2013).
Karmodiya, K., Krebs, A.R., Oulad-Abdelghani, M., Kimura, H. & Tora, L. H3K9 and H3K14 acetylation co-occur at many gene regulatory elements, while H3K14ac marks a subset of inactive inducible promoters in mouse embryonic stem cells. BMC Genomics 13, 424 (2012).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Xie, Z. et al. Metabolic regulation of gene expression by histone lysine β-hydroxybutyrylation. Mol. Cell 62, 194–206 (2016).
Sabari, B.R. et al. Intracellular crotonyl-CoA stimulates transcription through p300-catalyzed histone crotonylation. Mol. Cell 58, 203–215 (2015).
Koike, N. et al. Transcriptional architecture and chromatin landscape of the core circadian clock in mammals. Science 338, 349–354 (2012).
Taudt, A., Nguyen, M.A., Heinig, M., Johannes, F. & Colome-Tatche, M. chromstaR: Tracking combinatorial chromatin state dynamics in space and time. Preprint at https://www.biorxiv.org/content/early/2016/02/04/038612/ (2016).
McLean, C.Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Filippakopoulos, P. et al. Histone recognition and large-scale structural analysis of the human bromodomain family. Cell 149, 214–231 (2012).
Sanchez, R. & Zhou, M.-M.Therole of human bromodomains in chromatin biology and gene transcription. Curr. Opin. Drug Discov. Devel. 12, 659–665 (2009).
Vincentelli, R. et al. Quantifying domain-ligand affinities and specificities by high-throughput holdup assay. Nat. Methods 12, 787–793 (2015).
Fedorov, O. et al. Selective targeting of the BRG/PB1 bromodomains impairs embryonic and trophoblast stem cell maintenance. Sci. Adv. 1, e1500723 (2015).
Sanulli, S. et al. Jarid2 methylation via the PRC2 complex regulates H3K27me3 deposition during cell differentiation. Mol. Cell 57, 769–783 (2015).
An, W., Palhan, V.B., Karymov, M.A., Leuba, S.H. & Roeder, R.G. Selective requirements for histone H3 and H4 N termini in p300-dependent transcriptional activation from chromatin. Mol. Cell 9, 811–821 (2002).
Sabari, B.R., Zhang, D., Allis, C.D. & Zhao, Y. Metabolic regulation of gene expression through histone acylations. Nat. Rev. Mol. Cell Biol. 18, 90–101 (2017).
Struhl, K. Histone acetylation and transcriptional regulatory mechanisms. Genes Dev. 12, 599–606 (1998).
Taverna, S.D., Li, H., Ruthenburg, A.J., Allis, C.D. & Patel, D.J. How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nat. Struct. Mol. Biol. 14, 1025–1040 (2007).
Musselman, C.A., Lalonde, M.-E., Côté, J. & Kutateladze, T.G. Perceiving the epigenetic landscape through histone readers. Nat. Struct. Mol. Biol. 19, 1218–1227 (2012).
Jenuwein, T. & Allis, C.D. Translating the histone code. Science 293, 1074–1080 (2001).
Kadoch, C. et al. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat. Genet. 45, 592–601 (2013).
Hargreaves, D.C. & Crabtree, G.R. ATP-dependent chromatin remodeling: genetics, genomics and mechanisms. Cell Res. 396–420 (2011).
Kasten, M.M., Clapier, C.R. & Cairns, B.R. SnapShot: Chromatin remodeling: SWI/SNF. Cell 144, 310.e1 (2011).
Goudarzi, A. et al. Dynamic competing histone H4 K5K8 acetylation and butyrylation are hallmarks of highly active gene promoters. Mol. Cell 62, 169–180 (2016).
Shogren-Knaak, M. et al. Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science 311, 844–847 (2006).
Shchelochkov, O.A., Carrillo, N. & Venditti, C. Propionic acidemia. in GeneReviews (eds. Pagon, R.A. et al.) (Seattle, 1993).
Scopes, R.K. Measurement of protein by spectrophotometry at 205 nm. Anal. Biochem. 59, 277–282 (1974).
Tropberger, P. et al. Regulation of transcription through acetylation of H3K122 on the lateral surface of the histone octamer. Cell 152, 859–872 (2013).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Rumbaugh, G. & Miller, C.A. Epigenetic changes in the brain: measuring global histone modifications. Methods Mol. Biol. 670, 263–274 (2011).
Demény, M.A. et al. Identification of a small TAF complex and its role in the assembly of TAF-containing complexes. PLoS One 2, e316 (2007).
Chaya, D. & Zaret, K.S. Sequential chromatin immunoprecipitation from animal tissues. Methods Enzymol. 376, 361–372 (2004).
Arrigoni, L. et al. Standardizing chromatin research: a simple and universal method for ChIP-seq. Nucleic Acids Res. 44, e67 (2016).
Di Cerbo, V. et al. Acetylation of histone H3 at lysine 64 regulates nucleosome dynamics and facilitates transcription. eLife 3, e01632 (2014).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B.A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Thorvaldsdóttir, H., Robinson, J.T. & Mesirov, J.P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X.S. Identifying ChIP-seq enrichment using MACS. Nat. Protoc. 7, 1728–1740 (2012).
Zang, C. et al. A clustering approach for identification of enriched domains from histone modification ChIP-Seq data. Bioinformatics 25, 1952–1958 (2009).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res. 44 W1, W3–W10 (2016).
Oliveros, J.C. Venny: an interactive tool for comparing lists with Venn's diagrams. http://bioinfogp.cnb.csic.es/tools/venny/index.html (2007).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Ye, T. et al. seqMINER: an integrated ChIP-seq data interpretation platform. Nucleic Acids Res. 39, e35 (2011).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Anders, S., Pyl, P.T. & Huber, W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Love, M.I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Dignam, J.D., Lebovitz, R.M. & Roeder, R.G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 11, 1475–1489 (1983).
Vermeulen, M. Identifying chromatin readers using a SILAC-based histone peptide pull-down approach. Methods Enzymol. 512, 137–160 (2012).
Luger, K., Rechsteiner, T.J. & Richmond, T.J. Expression and purification of recombinant histones and nucleosome reconstitution. Methods Mol. Biol. 119, 1–16 (1999).
An, W. & Roeder, R.G. Reconstitution and transcriptional analysis of chromatin in vitro. Methods Enzymol. 377, 460–474 (2004).
Ito, T., Bulger, M., Pazin, M.J., Kobayashi, R. & Kadonaga, J.T. ACF, an ISWI-containing and ATP-utilizing chromatin assembly and remodeling factor. Cell 90, 145–155 (1997).
Margueron, R. et al. Ezh1 and Ezh2 maintain repressive chromatin through different mechanisms. Mol. Cell 32, 503–518 (2008).
Acknowledgements
We thank K. Bloom and M. Bennett (Children's Hospital Philadelphia) for Acads knockout tissues; A. Guenzel and M. Barry (Mayo Clinic) for Pcca knockout and gene-therapy-treated tissues; L. Arrigoni and J. Pospisilik for help with liver chromatin preparation; L. Tora (Institut de Genetique et de Biologie Moleculaire et Cellulaire, IGBMC) for baculoviruses expressing GCN5 and PCAF and for anti-GCN5 antibody; P. Laurette and I. Davidson (IGBMC) for antibodies against (P)BAF complex subunits; P. Eberling for peptide synthesis; S. Knapp (University of Frankfurt) for PFI-3 inhibitor; D. Widmann for initial analyses of data; and members of the Schneider laboratory for helpful discussions and reagents. M.C.-T. acknowledges support from the Helmholtz Association's Initiative and Networking Fund and from the University of Groningen (Rosalind Franklin Fellowship). Work by G. Meszaros and R.R. was supported by a European Research Council (ERC) starting grant (ERC-2011-StG, 281271-STRESSMETABOL). Work in the Schneider laboratory was supported by the Agence Nationale de Recherche (CoreAc), the DFG through SFB 1064, the Epigenomics of Common Diseases EpiTrio project and the Helmholtz Gesellschaft. Sequencing was performed by the IGBMC Microarray and Sequencing platform, a member of the 'France Génomique' consortium (ANR-10-INBS-0009).
Author information
Authors and Affiliations
Contributions
A.F.K. and R.S. conceived the project. A.F.K. characterized antibodies, performed most of the ChIP, knockdown and peptide pulldown experiments and analyzed data. A.N. contributed to antibody characterization and knockdown experiments, and performed GAL4 recruitment ChIP and luciferase assays. L.Z.S. performed in vitro transcription experiments with the help of R.M. S.L.G. and D.A.G. performed analyses of ChIP and RNA-seq data. F.R. and G. Mittler performed and analyzed histone-modification mass spectrometry experiments. M.P.B. and M.V. performed and analyzed peptide-pulldown mass spectrometry experiments. G. Meszaros, H.F.M. and R.R. contributed to animal fasting and liver cross-linking experiments. A.T. and M.C.-T. performed ChIP–seq data analyses. A.F.K., S.D. and R.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Mass Spectrometric identification and analysis of histone propionylation and butyrylation.
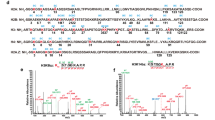
Acid-extracted histones from HeLa cells were separated by C8 reversed phase chromatography, run on a 4-20% gradient Tris-glycine gel and bands cut out and digested either with trypsin or ArgC. Peptides were then analyzed via LC-MS. (A) Tandem spectra (MS/MS) of H3 peptide (10-17) showing the presence of propionylation (mass shift: 56.03 Da) at K14 (H3K14pr). (B) MS/MS of H3 peptide (9-17) showing butyrylated (mass shift = 70.04 Da) K14 (H3K14bu). (C) Tandem spectra (MS/MS) of double modified H3 peptide (9-17) acetylated (42.01 Da) at K9 and propionylated at K14 (H3K9acK14pr). (D) same as C but with butyrylated K14 (H3K9acK14bu).. Note: In C, prominent unannotated peaks (in black) correspond to fragment ions originating from a co-eluting isobaric peptide isomer, which contains the alternative combinatorial modification pattern – H3K9prK14ac. Supporting spectra are available upon request.
Supplementary Figure 2 Characterization of antibodies specific for histone propionylation and butyrylation.
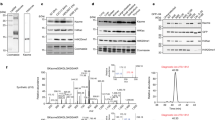
(A) Immunoblotting using H3K14pr (left), and H3K14bu (right) antibodies on acid-extracted histones from untreated and 10 mM sodium butyrate (NaBu) – treated HeLa cells. (B-C) Peptide competition with anti-H3K14pr (B) and anti-H3K14bu (C) antibodies. Antibodies were pre-incubated with either with un, ac, pr, or bu H3K14 peptides before being used to probe blots with acid extracted histones. For (A-C) Ponceau is shown as loading control and the position of each core histone is indicated. (D-E) Peptide dot blot assays with anti-H3K14pr (D), anti-H3K14bu (E) antibodies and histone H3 peptides unmodified (un), acetylated (ac), propionylated (pr) or butyrylated (bu) at lysines 9, 14, 18 and 23. Peptide sequences are given in Supplementary Table S1. (F) Immunoblotting for H3K14pr and H3K14bu in different human and mouse cell lines. Anti-H3K14bu and anti-H3K14pr antibodies were used to probe acid extracts from indicated cell lines. (G) Immunoblotting of H3K14pr and H3K14bu using acid-extracted histones from five different mouse tissues. For (F,G) Ponceau is shown as loading control. (H) Immunofluorescence staining of mouse fibroblasts (MEFs) using anti-H3K14bu antibody. DAPI was used to stain the DNA. Scale bar represents 5 μm.
Supplementary Figure 3 Histone acetyltransferases can propionylate and butyrylate histones.
(A) In vitro histone acylation assay using full length Flag-tagged GCN5 or PCAF (purified from insect cells), unlabeled acyl-coA donors (acetyl-, propionyl- and butyryl-coA) and calf thymus histones as substrate. Histones were resolved by SDS-PAGE. The indicated antibodies used to probe the membranes ponceau staining as loading control are shown. (B) Time course histone propionylation assay on histone octamers using Flag-tagged GCN5. Same volumes of reaction were spotted on nitrocellulose membrane at each time point and immunoblotting done with anti-H3K14pr antibody. (C) Quantification of the H3K14pr signal in (B). (D) siRNA knockdown of GCN5 and PCAF in HeLa cells. Anti-PCAF and anti-GCN5 antibodies are used to confirm depletion and anti-tubulin as loading control. (E) Immunoblotting to verify the depletion of p300 upon siRNA-mediated knockdown with tubulin as loading control. *Non-specific band. (F) Quantification of H3K9ac, H3K14pr and H3K14bu signals normalized to H4 in GCN5 and/or PCAF knockdown samples (see Fig.2B). Error bars represent standard deviation of three independent knockdown experiments. (G) Quantification of H3K18ac, H3K14pr and H3K14bu signals normalized to H4 in p300 and/or CBP knockdown samples (see Fig.2C). Error bars represent standard deviation of at two independent knockdown experiments. (H-I) RT-qPCR results assessing the efficiency of siRNA-mediated knockowns for p300/CBP (H), GCN5/PCAF (I) respectively. Error bars indicate standard deviation of two technical replicates of a representative experiment. (J) Immunoblotting for GCN5 and PCAF following their double knockdown in HeLa cells for ChIP experiment in Fig.2E. (K) Quantification of GCN5 and PCAF signals in (J) normalized to stain-free total protein staining. Error bars represent standard deviation of three technical replicates. (L) Quantification of H3K14pr signals from wild type (Wt), Pcca−/− and Pcca−/− gene therapy treated mice livers (see Fig. 3D). Error bars represent standard deviation of three different livers. (M) Quantification of H3K14bu signals from Wt and Acads−/− livers (see Fig. 3E). Error bars represent standard deviation of three technical replicates.
Supplementary Figure 4 H3K14 is propionylated and butyrylated in mouse livers.
Dialyzed (PBS) acid-extracted histones from mouse livers were separated on a 16% Tris-glycine gel and bands cut out and digested with trypsin. Peptides were then analyzed via LC-MS. (A) Tandem spectra (MS/MS) of H3 peptide (10-17) showing the presence of propionylation (mass shift: 56.03 Da) at K14 (H3K14pr). (B) MS/MS of H3 peptide (9-17) showing butyrylation (mass shift = 70.04 Da) at K14 (H3K14bu).
Supplementary Figure 5 H3K14pr and H3K14bu are histone marks linked to active transcription.
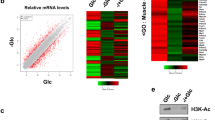
(A). Overlap of H3K14pr and H3K14bu with H3K4me3 in refed and fasted livers. Target genes were defined with the presence of a peak ±1kb of a TSS. (B) Immunoprecipitation of purified native HeLa mononucleosomes with indicated antibodies followed by immunoblotting to detect co-occuring modifications. Input represents 2%. (C) Top five enriched Gene Ontology (GO) biological process terms for H3K14pr-specific genes in livers from refed mice. (D) Top five enriched GO biological process terms for H3K14bu-specific genes in livers from refed mice. (E) Spearman correlation heatmap of histone acylations with other active (H3K4me3, H3K9bhb) (Xie, Z., et al. Mol Cell, 2016. 62(2)) and repressive (H3K9me3, H3K27me3) (Sugathan, A. and D.J. Waxman. Mol Cell Biol, 2013. 33(18)) marks as well as RNA pol II (ser5phospho) (Koike, N., et al., Science, 2012. 338(6105)). AL, ad libitum; ST, starved; R, Refed; F, Fasted (F-G) Top ten GO biological process terms associated with "triple-acylated" genes in the refed (F) and fasted (G) state. See Fig. 5B-C.
Supplementary Figure 6 Histone acylations bind the BAF complex differentially
(A) Euclidean clustering heatmap of proteins identified by mass spectrometry as enriched in affinity purifications with H3K14 peptides. Purifications were performed using HeLa nuclear extract and biotinylated H3 peptides unmodified (un), acetylated (ac), propionylated (pr) or butyrylated (bu) at K14. BAF/PBAF complex subunits binding to H3K14ac and H3K14pr are marked on the right. (B) Coomassie-stained gel of purified GST-tagged 2nd bromodomain of PBRM1 (GST-PBRM1(2)) used for the direct binding experiment in Fig. 6D-E.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–4 (PDF 1267 kb)
Supplementary Data Set 1
Uncropped blots and membranes from key data within main figures. (PDF 2577 kb)
Rights and permissions
About this article
Cite this article
Kebede, A., Nieborak, A., Shahidian, L. et al. Histone propionylation is a mark of active chromatin. Nat Struct Mol Biol 24, 1048–1056 (2017). https://doi.org/10.1038/nsmb.3490
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3490
This article is cited by
-
Histone butyrylation in the mouse intestine is mediated by the microbiota and associated with regulation of gene expression
Nature Metabolism (2024)
-
Metabolic waypoints during T cell differentiation
Nature Immunology (2024)
-
A glimpse into novel acylations and their emerging role in regulating cancer metastasis
Cellular and Molecular Life Sciences (2024)
-
Roles of protein post-translational modifications in glucose and lipid metabolism: mechanisms and perspectives
Molecular Medicine (2023)
-
Epigenetic meets metabolism: novel vulnerabilities to fight cancer
Cell Communication and Signaling (2023)