Abstract
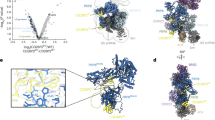
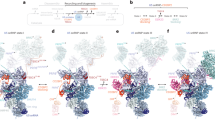
The spliceosome is a multimegadalton RNA-protein machine that removes noncoding sequences from nascent pre-mRNAs. Recruitment of the spliceosome to splice sites and subsequent splicing require a series of dynamic interactions among the spliceosome's component U snRNPs and many additional protein factors. These dynamics present several challenges for structural analyses, including purification of stable complexes to compositional homogeneity and assessment of conformational heterogeneity. We have isolated spliceosomes arrested before the second chemical step of splicing (C complex) in which U2, U5 and U6 snRNAs are stably associated. Using electron microscopy, we obtained images of C complex spliceosomes under cryogenic conditions and determined a three-dimensional structure of a core complex to a resolution of 30 Å. The structure reveals a particle of dimensions 27 × 22 × 24 nm with a relatively open arrangement of three primary domains.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Nilsen, T.W. The spliceosome: no assembly required? Mol. Cell 9, 8–9 (2002).
Jiang, J., Horowitz, D.S. & Xu, R.M. Crystal structure of the functional domain of the splicing factor Prp18. Proc. Natl. Acad. Sci. USA 97, 3022–3027 (2000).
Kambach, C. et al. Crystal structures of two Sm protein complexes and their implications for the assembly of the spliceosomal snRNPs. Cell 96, 375–387 (1999).
Kielkopf, C.L., Rodionova, N.A., Green, M.R. & Burley, S.K. A novel peptide recognition mode revealed by the X-ray structure of a core U2AF35/U2AF65 heterodimer. Cell 106, 595–605 (2001).
Oubridge, C., Ito, N., Evans, P.R., Teo, C.H. & Nagai, K. Crystal structure at 1.92 Å resolution of the RNA-binding domain of the U1A spliceosomal protein complexed with an RNA hairpin. Nature 372, 432–438 (1994).
Price, S.R., Evans, P.R. & Nagai, K. Crystal structure of the spliceosomal U2B″-U2A′ protein complex bound to a fragment of U2 small nuclear RNA. Nature 394, 645–650 (1998).
Vidovic, I., Nottrott, S., Hartmuth, K., Luhrmann, R. & Ficner, R. Crystal structure of the spliceosomal 15.5kD protein bound to a U4 snRNA fragment. Mol. Cell 6, 1331–2342 (2000).
Reuter, K., Nottrott, S., Fabrizio, P., Luhrmann, R. & Ficner, R. Identification, characterization and crystal structure analysis of the human spliceosomal U5 snRNP-specific 15 kD protein. J. Mol. Biol. 294, 515–525 (1999).
Stark, H., Dube, P., Luhrmann, R. & Kastner, B. Arrangement of RNA and proteins in the spliceosomal U1 small nuclear ribonucleoprotein particle. Nature 409, 539–542 (2001).
Golas, M.M., Sander, B., Will, C.L., Luhrmann, R. & Stark, H. Molecular architecture of the multiprotein splicing factor SF3b. Science 300, 980–984 (2003).
Jurica, M., Licklider, L., Gygi, S., Grigorieff, N. & Moore, M. Purification and characterization of native spliceosomes suitable for three-dimensional structural analysis. RNA 8, 426–439 (2002).
Stevens, S.W. et al. Composition and functional characterization of the yeast spliceosomal penta-snRNP. Mol. Cell 9, 31–44. (2002).
Van Heel, M. Angular reconstitution: a posteriori assignment of projection directions for 3D reconstruction. Ultramicroscopy 21, 111–123 (1987).
Radermacher, M., Wagenknecht, T., Verschoor, A. & Frank, J. Three-dimensional reconstruction from a single-exposure, random conical tilt series applied to the 50S ribosomal subunit of Escherichia coli. J. Microsc. 146 (Pt 2), 113–136 (1987).
Frank, J. & Radermacher, M. Three-dimensional reconstruction of single particles negatively stained or in vitreous ice. Ultramicroscopy 46, 241–262 (1992).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Grigorieff, N. Structure of the respiratory NADH:ubiquinone oxidoreductase (complex I). Curr. Opin. Struct. Biol. 9, 476–483 (1999).
Ajuh, P. et al. Functional analysis of the human CDC5L complex and identification of its components by mass spectrometry. EMBO J. 19, 6569–6581. (2000).
Neubauer, G. et al. Mass spectrometry and EST-database searching allows characterization of the multi-protein spliceosome complex. Nat. Genet. 20, 46–50. (1998).
Ohi, M.D. et al. Proteomics analysis reveals stable multiprotein complexes in both fission and budding yeasts containing Myb-related Cdc5p/Cef1p, novel pre-mRNA splicing factors, and snRNAs. Mol. Cell. Biol. 22, 2011–2024. (2002).
Rappsilber, J., Ryder, U., Lamond, A.I. & Mann, M. Large-scale proteomic analysis of the human spliceosome. Genome Res. 12, 1231–1245. (2002).
Zhou, Z., Licklider, L.J., Gygi, S.P. & Reed, R. Comprehensive proteomic analysis of the human spliceosome. Nature 419, 182–185. (2002).
Staley, J.P. & Guthrie, C. Mechanical devices of the spliceosome: motors, clocks, springs, and things. Cell 92, 315–326 (1998).
Kastner, B., Bach, M. & Luhrmann, R. Electron microscopy of small nuclear ribonucleoprotein (snRNP) particles U2 and U5: evidence for a common structure-determining principle in the major U snRNP family. Proc. Natl. Acad. Sci. USA 87, 1710–1714 (1990).
Makarov, E.M. et al. Small nuclear ribonucleoprotein remodeling during catalytic activation of the spliceosome. Science 298, 2205–2208 (2002).
Behrens, S.E., Tyc, K., Kastner, B., Reichelt, J. & Luhrmann, R. Small nuclear ribonucleoprotein (RNP) U2 contains numerous additional proteins and has a bipartite RNP structure under splicing conditions. Mol. Cell. Biol. 13, 307–319 (1993).
Mindell, J.A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Crowther, R.A., Henderson, R. & Smith, J.M. MRC image processing programs. J. Struct. Biol. 116, 9–16 (1996).
Matthews, B.W. Solvent content of protein crystals. J. Mol. Biol. 33, 491–497 (1968).
Acknowledgements
We thank T. Walz and M. Ohi for discussion and H. Stark for advice on sample preparation. N.G. is an assistant investigator and M.J.M is an associate investigator with the Howard Hughes Medical Institute; M.S.J. is a Paul Sigler/Agouron Institute fellow of the Helen Hay Whitney Foundation; D.R.S. is supported by a US National Science Foundation integrative graduate education and research training grant. This work was supported by US National Institutes of Health grant 1 P01 GM-62580 (N.G.) and GM 53007 (M.J.M.) and funding from the Keck Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Jurica, M., Sousa, D., Moore, M. et al. Three-dimensional structure of C complex spliceosomes by electron microscopy. Nat Struct Mol Biol 11, 265–269 (2004). https://doi.org/10.1038/nsmb728
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb728
This article is cited by
-
Mechanistic insights into precursor messenger RNA splicing by the spliceosome
Nature Reviews Molecular Cell Biology (2017)
-
Cryo-electron microscopy snapshots of the spliceosome: structural insights into a dynamic ribonucleoprotein machine
Nature Structural & Molecular Biology (2017)
-
Cryo-EM structure of a human spliceosome activated for step 2 of splicing
Nature (2017)
-
Exon, intron and splice site locations in the spliceosomal B complex
The EMBO Journal (2009)
-
A protein-based EM label for RNA identifies the location of exons in spliceosomes
Nature Structural & Molecular Biology (2008)