Abstract
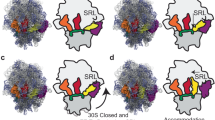
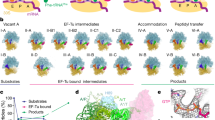
Using single-molecule methods we observed the stepwise movement of aminoacyl-tRNA (aa-tRNA) into the ribosome during selection and kinetic proofreading using single-molecule fluorescence resonance energy transfer (smFRET). Intermediate states in the pathway of tRNA delivery were observed using antibiotics and nonhydrolyzable GTP analogs. We identified three unambiguous FRET states corresponding to initial codon recognition, GTPase-activated and fully accommodated states. The antibiotic tetracycline blocks progression of aa-tRNA from the initial codon recognition state, whereas cleavage of the sarcin-ricin loop impedes progression from the GTPase-activated state. Our data support a model in which ribosomal recognition of correct codon-anticodon pairs drives rotational movement of the incoming complex of EF-Tu–GTP–aa-tRNA toward peptidyl-tRNA during selection on the ribosome. We propose a mechanistic model of initial selection and proofreading.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Ramakrishnan, V. Ribosome structure and the mechanism of translation. Cell 108, 557–572 (2002).
Hazlett, T.L., Johnson, A.E. & Jameson, D.M. Time-resolved fluorescence studies on the ternary complex formed between bacterial elongation factor Tu, guanosine 5′-triphosphate, and phenylalanyl-tRNAPhe. Biochemistry 28, 4109–4117 (1989).
Gavrilova, L.P., Perminova, I.N. & Spirin, A.S. Elongation factor Tu can reduce translation errors in poly(U)-directed cell-free systems. J. Mol. Biol. 149, 69–78 (1981).
Gavrilova, L.P., Kostiashkina, O.E., Koteliansky, V.E., Rutkevitch, N.M. & Spirin, A.S. Factor-free (“non-enzymic”) and factor-dependent systems of translation of polyuridylic acid by Escherichia coli ribosomes. J. Mol. Biol. 101, 537–552 (1976).
Rodnina, M.V. & Wintermeyer, W. Ribosome fidelity: tRNA discrimination, proofreading and induced fit. Trends Biochem. Sci. 26, 124–130 (2001).
Gromadski, K.B. & Rodnina, M.V. Kinetic determinants of high-fidelity tRNA discrimination on the ribosome. Mol. Cell 13, 191–200 (2004).
Hamel, E., Koka, M. & Nakamoto, T. Requirement of an Escherichia coli 50S ribosomal protein component for effective interaction of the ribosome with T and G factors and with guanosine triphosphate. J. Biol. Chem. 247, 805–814 (1972).
Liljas, A. & Gudkov, A.T. The structure and dynamics of ribosomal protein L12. Biochimie 69, 1043–1047 (1987).
Rodnina, M.V. & Wintermeyer, W. Fidelity of aminoacyl-tRNA selection on the ribosome: kinetic and structural mechanisms. Annu. Rev. Biochem. 70, 415–435 (2001).
Rodnina, M.V. et al. GTPases mechanisms and functions of translation factors on the ribosome (Review). Biol. Chem. 381, 377–387 (2000).
Gale, E.F., Cundliffe, E., Reynolds, P.E., Richmond, M.H. & Waring, M.J. The Molecular Basis of Antibiotic Action (Wiley, London, 1981).
Parker, J. Errors and alternatives in reading the universal genetic code. Microbiol. Rev. 53, 273–298 (1989).
Grosjean, H.J., Henau, S. & Crothers, D.M. On the physical basis for ambiguity in genetic coding interactions. Proc. Natl. Acad. Sci. USA 75, 610–614 (1978).
Hopfield, J.J. Kinetic proofreading: a new mechanism for reducing errors in biosynthetic processes requiring high specificity. Proc. Natl. Acad. Sci. USA 71, 4135–4139 (1974).
Ninio, J. Kinetic amplification of enzyme discrimination. Biochimie 57, 587–595 (1975).
Thompson, R.C. & Dix, D.B. Accuracy in protein synthesis. A kinetic study of the reaction of poly(U)-programmed ribosomes with a leucyl-tRNA2 elongation factor Tu-GTP complex. J. Biol. Chem. 257, 6677–6682 (1982).
Pape, T., Wintermeyer, W. & Rodina, M.W. Induced fit in initial selection and proofreading of aminoacyl-tRNA on the ribosome. Eur. J. Mol. Biol. 18, 3800–3807 (1999).
Fourmy, D., Recht, M.I., Blanchard, S.C. & Puglisi, J.D. Structure of the A site of E. coli 16S ribosomal RNA complexed with an aminoglycoside antibiotic. Science 274, 1367–1371 (1996).
Yoshizawa, S., Fourmy, D. & Puglisi, J.D. Recognition of the codon-anticodon helix by ribosomal RNA. Science 285, 1722–1725 (1999).
Ogle, J.M. et al. Recognition of cognate transfer RNA by the 30S ribosomal subunit. Science 292, 897–902 (2001).
Ogle, J.M., Carter, A.P. & Ramakrishnan, V. Insights into the decoding mechanism from recent ribosome structures. Trends Biochem. Sci. 28, 259–266 (2003).
Piepenburg, O. et al. Intact aminoacyl-tRNA is required to trigger GTP hydrolysis by elongation factor Tu on the ribosome. Biochemistry 39, 1734–1738 (2000).
Powers, T. & Noller, H.F. The 530 loop of 16S rRNA: a signal to EF-Tu? Trends Genet. 10, 27–31 (1994).
Valle, M. et al. Incorporation of aminoacyl-tRNA into the ribosome as seen by cryo-electron microscopy. Nat. Struct. Biol. 10, 899–906 (2003).
Watson, B.S. et al. Macromolecular arrangement in the aminoacyl-tRNA.elongation factor Tu.GTP ternary complex. A fluorescence energy transfer study. Biochemistry 34, 7904–7912 (1995).
Blanchard, S.C., Kim, H.D., Gonzalez, R.L. Jr., Puglisi, J.D. & Chu, S. tRNA dynamics on the ribosome during translation. Proc. Natl. Acad. Sci. USA 101, 12893–12898 (2004).
Plumbridge, J.A., Baumert, H.G., Ehrenberg, M. & Rigler, R. Characterisation of a new, fully active fluorescent derivative of E. coli tRNA Phe. Nucleic Acids Res. 8, 827–843 (1980).
Cundliffe, E. Antibiotics and prokaryotic ribosomes: action interaction and resistance. In Ribosomes, Structure, Function, and Genetics. Proceedings of the 9th Steenbock Symposium (eds. Chambliss, G. et al.) 555–581 (University Park Press, Baltimore, 1980).
Rodnina, M.V., Fricke, R. & Wintermeyer, W. Transient conformational states of aminoacyl-tRNA during ribosome binding catalyzed by elongation factor Tu. Biochemistry 33, 12267–12275 (1994).
Vogeley, L., Palm, G.J., Mesters, J.R. & Hilgenfeld, R. Conformational change of elongation factor Tu (EF-Tu) induced by antibiotic binding. Crystal structure of the complex between EF-Tu.GDP and aurodox. J. Biol. Chem. 276, 17149–17155 (2001).
Wolf, H., Chinali, G. & Parmeggiani, A. Mechanism of the inhibition of protein synthesis by kirromycin. Eur. J. Biochem. 75, 67–75 (1977).
Moazed, D., Robertson, J.M. & Noller, H.F. Interaction of elongation factors EF-G and EF-Tu with a conserved loop in 23S RNA. Nature 334, 362–364 (1988).
Hausner, T.P., Atmadja, J. & Nierhaus, K.H. Evidence that the G2661 region of 23S rRNA is located at the ribosomal binding sites of both elongation factors. Biochimie 69, 911–923 (1987).
Kurland, C.G., Hughes, D. & Ehrenberg, M. Limitations of Translational Accuracy 979–1004 (American Society for Microbiology Press, Washington, DC, 1996).
Bilgin, N. & Ehrenberg, M. Mutations in 23S ribosomal RNA perturb transfer RNA selection and can lead to streptomycin dependence. J. Mol. Biol. 235, 813–824 (1994).
Valle, M. et al. Cryo-EM reveals an active role for aminoacyl-tRNA in the accommodation process. EMBO J. 21, 3557–3567 (2002).
Stark, H. et al. Ribosome interactions of aminoacyl-tRNA and elongation factor Tu in the codon-recognition complex. Nat. Struct. Biol. 9, 849–854 (2002).
Pape, T., Wintermeyer, W. & Rodnina, M.V. Complete kinetic mechanism of elongation factor Tu-dependent binding of aminoacyl-tRNA to the A site of the E. coli ribosome. EMBO J. 17, 7490–7497 (1998).
Brodersen, D.E. et al. The structural basis for the action of the antibiotics tetracycline, pactamycin, and hygromycin B on the 30S ribosomal subunit. Cell 103, 1143–1154 (2000).
Gordon, J. Hydrolysis of guanosine 5′-triphosphate associated with binding of aminoacyl transfer ribonucleic acid to ribosomes. J. Biol. Chem. 244, 5680–5686 (1969).
Ogle, J.M., Murphy, F.V., Tarry, M.J. & Ramakrishnan, V. Selection of tRNA by the ribosome requires a transition from an open to a closed form. Cell 111, 721–732 (2002).
Thompson, R.C., Dix, D.B., Gerson, R.B. & Karim, A.M. Effect of Mg2+ concentration, polyamines, streptomycin, and mutations in ribosomal proteins on the accuracy of the two-step selection of aminoacyl-tRNAs in protein biosynthesis. J. Biol. Chem. 256, 6676–6681 (1981).
Janiak, F. et al. Fluorescence characterization of the interaction of various transfer RNA species with elongation factor Tu.GTP: evidence for a new functional role for elongation factor Tu in protein biosynthesis. Biochemistry 29, 4268–4277 (1990).
O'Connor, M.O. & Dahlberg, A.E. The involvement of two distinct regions of 23S ribosomal RNA in tRNA selection. J. Mol. Biol. 254, 838–847 (1995).
Muth, G.W., Chen, L., Kosek, A.B. & Strobel, S.A. pH-dependent conformational flexibility within the ribosomal peptidyl transferase center. RNA 7, 1403–1415 (2001).
Bayfield, M.A., Dahlberg, A.E., Schulmeister, U., Dorner, S. & Barta, A. A conformational change in the ribosomal peptidyl transferase center upon active/inactive transition. Proc. Natl. Acad. Sci. USA 98, 10096–10101 (2001).
Harms, J. et al. High resolution structure of the large ribosomal subunit from a mesophilic eubacterium. Cell 107, 679–688 (2001).
Ban, N., Nissen, P., Hansen, J., Moore, P.B. & Steitz, T.A. The complete atomic structure of the large ribosomal subunit at 2.4 Å resolution. Science 289, 905–920 (20).
Acknowledgements
S.C.B. is supported by the Giannini Family Foundation, and R.L.G. is supported by the American Cancer Society. Restrictocin was a gift of C. Correll (University of Chicago). This work was supported by grants to J.D.P. from the US National Institutes of Health (GM51266) and the David and Lucille Packard Foundation, to S.C. from the US National Science Foundation, the US National Aeronautics and Space Administration and the US Air Force Office of Scientific Research, and to S.C. and J.D.P. from the David and Lucille Packard Foundation Interdisciplinary Science Program (grant 2000-01671). The authors thank E. Lau for technical support, J. Hoch for suggestions relating to fluorescence quenching, E.V. Puglisi for critical discussions and scientific input, and M. Dorywalska and J. Choy for comments on the written manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Histogram analysis of FRET states achieved after EF-Tu(GTP)Phe-tRNAPhe delivery to surface immobilized ribosome complexes. (PDF 58 kb)
Supplementary Fig. 2
FRET arrival time plot. (PDF 58 kb)
Supplementary Fig. 3
Time evolution of FRET states in the delivery of EF-Tu(GTP)Phe-tRNAPhe (Cy5-acp3U) to surface immobilized 70S complexes initiated with fMet-tRNAfMet (Cy3-s4U) in the P site and a phenylalanine codon (UUU) in the A site. (PDF 58 kb)
Supplementary Fig. 4
Lifetime analysis of the 0.35 FRET state in tetracycline-stalled EF-Tu(GTP)Phe-tRNAPhe delivery to the ribosome. (PDF 36 kb)
Supplementary Fig. 5
The lifetime of the 0.5 FRET state (pre-GTP hydrolysis). GDPNP is a non-hydrolyzable GTP analogue. (PDF 370 kb)
Supplementary Fig. 6
Cleavage of the SRL within tight coupled-70S particles was achieved using the enzyme restrictocin. (PDF 114 kb)
Supplementary Fig. 7
EF-Tu(GTP)Phe-tRNAPhe (Cy5-acp3U) is stalled in the 0.5 FRET state when delivered to immobilized ribosome complexes cleaved at the SRL. (PDF 29 kb)
Supplementary Fig. 8
Lifetime analysis of the 0.35 FRET state in delivery of EF-Tu(GDPNP)Phe-tRNAPhe to ribosome complexes programmed with a cognate (UUU). (PDF 29 kb)
Supplementary Fig. 9
Lifetime analysis of the 0.35 FRET state prior to transition to the zero FRET state in delivery of EF-Tu(GTP)Phe-tRNAPhe to ribosome complexes programmed with a cognate (UUU) and near-cognate (CUU) codon. (PDF 32 kb)
Supplementary Table 1
Fidelity calculations. (PDF 25 kb)
Rights and permissions
About this article
Cite this article
Blanchard, S., Gonzalez, R., Kim, H. et al. tRNA selection and kinetic proofreading in translation. Nat Struct Mol Biol 11, 1008–1014 (2004). https://doi.org/10.1038/nsmb831
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb831
This article is cited by
-
Geometric alignment of aminoacyl-tRNA relative to catalytic centers of the ribosome underpins accurate mRNA decoding
Nature Communications (2023)
-
Dynamics of the context-specific translation arrest by chloramphenicol and linezolid
Nature Chemical Biology (2020)
-
Multiplexed genomic encoding of non-canonical amino acids for labeling large complexes
Nature Chemical Biology (2020)
-
2′-O-methylation in mRNA disrupts tRNA decoding during translation elongation
Nature Structural & Molecular Biology (2018)
-
Hierarchical mechanism of amino acid sensing by the T-box riboswitch
Nature Communications (2018)