Abstract
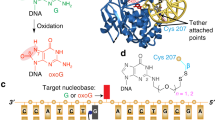
Uracil DNA glycosylase (UDG) removes uracil from U·A or U·G base pairs in genomic DNA by extruding the aberrant uracil from the DNA base stack. A question in enzymatic DNA repair is whether UDG and related glycosylases also use an extrahelical recognition mechanism to inspect the integrity of undamaged base pairs. Using NMR imino proton exchange measurements we find that UDG substantially increases the equilibrium constant for opening of T-A base pairs by almost two orders of magnitude relative to free B-DNA. This increase is brought about by enzymatic stabilization of an open state of the base pair without increasing the rate constant for spontaneous base pair opening. These findings indicate a passive search mechanism in which UDG uses the spontaneous opening dynamics of DNA to inspect normal base pairs in a rapid genome-wide search for uracil in DNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Stivers, J.T. & Jiang, Y.L. A mechanistic perspective on the chemistry of DNA repair glycosylases. Chem. Rev. 103, 2729–2759 (2003).
Stivers, J.T. Site-specific DNA damage recognition by enzyme-induced base flipping. Prog. Nucleic Acid Res. Mol. Biol. 77, 37–65 (2004).
Parikh, S.S. et al. Base excision repair initiation revealed by crystal structures and binding kinetics of human uracil-DNA glycosylase with DNA. EMBO J. 17, 5214–5226 (1998).
Parikh, S.S. et al. Uracil-DNA glycosylase-DNA substrate and product structures: conformational strain promotes catalytic efficiency by coupled stereoelectronic effects. Proc. Natl. Acad. Sci. USA 97, 5083–5088 (2000).
Werner, R.M. et al. Stressing-out DNA? The contribution of serine-phosphodiester interactions in catalysis by uracil DNA glycosylase. Biochemistry 39, 12585–12594 (2000).
Drohat, A.C. & Stivers, J.T. NMR evidence for an unusually low N1 pKa for uracil bound to uracil DNA glycosylase: Implications for catalysis. J. Am. Chem. Soc. 122, 1840–1841 (2000).
Kwon, K., Jiang, Y. & Stivers, J. Rational engineering of a DNA glycosylase specific for an unnatural cytosine:pyrene base pair. Chem. Biol. 10, 1–20 (2003).
Cao, C., Kwon, K., Jiang, Y.L., Drohat, A.C. & Stivers, J.T. Solution structure and base perturbation studies reveal a novel mode of alkylated base recognition by 3-methyladenine DNA glycosylase I. J. Biol. Chem. 48012–48021 (2003).
Slupphaug, G. et al. A nucleotide-flipping mechanism from the structure of human uracil-DNA glycosylase bound to DNA [see comments]. Nature 384, 87–92 (1996).
Lau, A.Y., Scharer, O.D., Samson, L., Verdine, G.L. & Ellenberger, T. Crystal structure of a human alkylbase-DNA repair enzyme complexed to DNA: mechanisms for nucleotide flipping and base excision. Cell 95, 249–258 (1998).
Lau, A.Y., Wyatt, M.D., Glassner, B.J., Samson, L.D. & Ellenberger, T. Molecular basis for discriminating between normal and damaged bases by the human alkyladenine glycosylase, AAG. Proc. Natl. Acad. Sci. USA 97, 13573–13578 (2000).
Jiang, Y.L. & Stivers, J.T. Mutational analysis of the base flipping mechanism of uracil DNA glycosylase. Biochemistry 41, 11236–11247 (2002).
Stivers, J.T., Pankiewicz, K.W. & Watanabe, K.A. Kinetic mechanism of damage site recognition and uracil flipping by Escherichia coli uracil DNA glycosylase. Biochemistry 38, 952–963 (1999).
Roberts, R.J. On base flipping. Cell 82, 9–12 (1995).
Krosky, D.J., Schwarz, F.P. & Stivers, J.T. Linear free energy correlations for enzymatic base flipping: how do damaged base pairs facilitate specific recognition? Biochemistry 43, 4188–4195 (2004).
Jencks, W.P. When is an intermediate not an intermediate—enforced mechanisms of general acid-base catalyzed, carbocation, carbanion, and ligand-exchange reactions. Acc. Chem. Res. 13, 161–169 (1980).
Chen, L., Haushalter, K.A., Lieber, C.M. & Verdine, G.L. Direct visualization of a DNA glycosylase searching for damage. Chem. Biol. 9, 345–350 (2002).
Verdine, G.L. & Bruner, S.D. How do DNA repair proteins locate damaged bases in the genome? Chem. Biol. 4, 329–334 (1997).
Seibert, E., Ross, J.B. & Osman, R. Role of DNA flexibility in sequence-dependent activity of uracil DNA glycosylase. Biochemistry 41, 10976–10984 (2002).
Banavali, N.K. & MacKerell, A.D. Jr. Free energy and structural pathways of base flipping in a DNA GCGC containing sequence. J. Mol. Biol. 319, 141–160 (2002).
Gueron, M. & Leroy, J.-L. Studies of Base Pair Kinetics by NMR Measurement of Proton Exchange. Methods Enzymol. 261, 383–413 (1995).
Guéron, M. et al. Applications to imino proton exchange to nucleic acid kinetics and structure. In Structure and Methods (eds. Sarma, R.H. & Sarma, M.H.) 113–137 (Adenine Press, Guilderland, New York, 1990).
Kavli, B. et al. Excision of cytosine and thymine from DNA by mutants of human uracil-DNA glycosylase. EMBO J. 15, 3442–3447 (1996).
Gueron, M., Kochoyan, M. & Leroy, J.L. A single mode of DNA base-pair opening drives imino proton exchange. Nature 328, 89–92 (1987).
Warmlander, S., Sen, A. & Leijon, M. Imino proton exchange in DNA catalyzed by ammonia and trimethylamine: evidence for a secondary long-lived open state of the base pair. Biochemistry 39, 607–615 (2000).
Handa, P., Acharya, N. & Varshney, U. Effects of mutations at tyrosine 66 and asparagine 123 in the active site pocket of Escherichia coli uracil DNA glycosylase on uracil excision from synthetic DNA oligomers: evidence for the occurrence of long-range interactions between the enzyme and substrate. Nucleic Acids Res. 30, 3086–3095 (2002).
Jiang, Y.L., Song, F. & Stivers, J.T. Base flipping mutations of uracil DNA glycosylase: substrate rescue using a pyrene nucleotide wedge. Biochemistry 41, 11248–11254 (2002).
Bommarito, S., Peyret, N. & SantaLucia, J. Jr. Thermodynamic parameters for DNA sequences with dangling ends. Nucleic Acids Res. 28, 1929–1934 (2000).
Stivers, J.T., Jagadeesh, G.J., Nawrot, B., Stec, W.J. & Shuman, S. Stereochemical outcome and kinetic effects of Rp- and Sp- phosphorothioate substitutions at the cleavage site of vaccinia type I DNA topoisomerase. Biochemistry 39, 5561–5572 (2000).
Drohat, A.C., Jagadeesh, J., Ferguson, E. & Stivers, J.T. Role of electrophilic and general base catalysis in the mechanism of Escherichia coli uracil DNA glycosylase. Biochemistry 38, 11866–11875 (1999).
Jiang, Y.L., Kwon, K. & Stivers, J.T. Turning on uracil-DNA glycosylase using a pyrene nucleotide switch. J. Biol. Chem. 276, 42347–42354 (2001).
Jiang, Y.L., Ichikawa, Y. & Stivers, J.T. Inhibition of uracil DNA glycosylase by an oxacarbenium ion mimic. Biochemistry 41, 7116–7124 (2002).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Snoussi, K. & Leroy, J.L. Imino proton exchange and base-pair kinetics in RNA duplexes. Biochemistry 40, 8898–9904 (2001).
Benight, A.S., Schurr, J.M., Flynn, P.F., Reid, B.R. & Wemmer, D.E. Melting of a self-complementary DNA minicircle. Comparison of optical melting theory with exchange broadening of the nuclear magnetic resonance spectrum. J. Mol. Biol. 200, 377–399 (1988).
Acknowledgements
This work was supported by US National Institutes of Health grant GM56834-09 to J.T.S.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Determination of the equilibrium dissociation constants for the T-A and G-C duplexes. (PDF 52 kb)
Supplementary Table 1
Rate and equilibrium constants for base pair opening in free and bound T/A duplex DNA. (PDF 43 kb)
Supplementary Table 2
Rate and equilibrium constants for base pair opening in 2′-FU·A duplex DNA. (PDF 34 kb)
Supplementary Table 3
Rate and equilibrium constants for base pair opening in free and bound G-C duplex DNA. (PDF 36 kb)
Rights and permissions
About this article
Cite this article
Cao, C., Jiang, Y., Stivers, J. et al. Dynamic opening of DNA during the enzymatic search for a damaged base. Nat Struct Mol Biol 11, 1230–1236 (2004). https://doi.org/10.1038/nsmb864
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb864
This article is cited by
-
Uracil-DNA glycosylase efficiency is modulated by substrate rigidity
Scientific Reports (2023)
-
Binding of undamaged double stranded DNA to vaccinia virus uracil-DNA Glycosylase
BMC Structural Biology (2015)
-
Identification of DNA lesions using a third base pair for amplification and nanopore sequencing
Nature Communications (2015)
-
Timing facilitated site transfer of an enzyme on DNA
Nature Chemical Biology (2012)
-
Duplex interrogation by a direct DNA repair protein in search of base damage
Nature Structural & Molecular Biology (2012)