Abstract
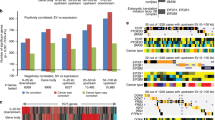
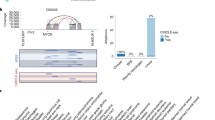
Deletion of 11q23–q24 is frequent in a diverse variety of malignancies, including breast and colorectal carcinoma, implicating the presence of a tumor suppressor gene at that chromosomal region. We examined a 6-Mb region on 11q23 by high-resolution deletion mapping, using both loss of heterozygosity analysis and customized microarray comparative genomic hybridization. LARG (leukemia-associated Rho guanine-nucleotide exchange factor) (also called ARHGEF12), identified from the analysed region, is frequently underexpressed in breast and colorectal carcinomas with a reduced expression observed in all breast cancer cell lines (n=11), in 12 of 38 (32%) primary breast cancers, 5 of 10 (50%) colorectal cell lines and in 20 of 37 (54%) primary colorectal cancers. Underexpression of the LARG transcript was significantly associated with genomic loss (P=0.00334). Hypermethylation of the LARG promoter was not detected in either breast or colorectal cancer, and treatment of four breast and four colorectal cancer cell lines with 5-aza-2′-deoxycytidine and/or trichostatin A did not result in a reactivation of LARG. Enforced expression of LARG in breast and colorectal cancer cells by stable transfection resulted in reduced cell proliferation and colony formation, as well as in a markedly slower cell migration rate in colorectal cancer cells, providing functional evidence for LARG as a candidate tumor suppressor gene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Baffa R, Negrini M, Mandes B, Rugge M, Ranzani GN, Hirohashi S et al. (1996). Loss of heterozygosity for chromosome 11 in adenocarcinoma of the stomach. Cancer Res 56: 268–272.
Bourguignon LY, Gilad E, Brightman A, Diedrich F, Singleton P . (2006). Hyaluronan–CD44 interaction with leukemia-associated RhoGEF and epidermal growth factor receptor promotes Rho/Ras co-activation, phospholipase C epsilon-Ca2+ signaling, and cytoskeleton modification in head and neck squamous cell carcinoma cells. J Biol Chem 281: 14026–14040.
Brooks CL, Gu W . (2009). How does SIRT1 affect metabolism, senescence and cancer? Nat Rev Cancer 9: 123–128.
Dotto GP . (2008). Notch tumor suppressor function. Oncogene 27: 5115–5123.
Ehrich M, Nelson MR, Stanssens P, Zabeau M, Liloglou T, Xinarianos G et al. (2005). Quantitative high-throughput analysis of DNA methylation patterns by base-specific cleavage and mass spectrometry. Proc Natl Acad Sci USA 102: 15785–15790.
Esteller M . (2007). Cancer epigenomics: DNA methylomes and histone-modification maps. Nat Rev Genet 8: 286–298.
Fukuhara S, Chikumi H, Gutkind JS . (2000). Leukemia-associated Rho guanine nucleotide exchange factor (LARG) links heterotrimeric G proteins of the G(12) family to Rho. FEBS Lett 485: 183–188.
Gabra H, Watson JE, Taylor KJ, Mackay J, Leonard RC, Steel CM et al. (1996). Definition and refinement of a region of loss of heterozygosity at 11q23.3–q24.3 in epithelial ovarian cancer associated with poor prognosis. Cancer Res 56: 950–954.
Harris BZ, Lim WA . (2001). Mechanism and role of PDZ domains in signaling complex assembly. J Cell Sci 114: 3219–3231.
Hui AB, Lo KW, Leung SF, Choi PH, Fong Y, Lee JC et al. (1996). Loss of heterozygosity on the long arm of chromosome 11 in nasopharyngeal carcinoma. Cancer Res 56: 3225–3229.
Jain AN, Tokuyasu TA, Snijders AM, Segraves R, Albertson DG, Pinkel D . (2002). Fully automatic quantification of microarray image data. Genome Res 12: 325–332.
Koreth J, Bakkenist CJ, Larin Z, Hunt NC, James MR, McGee JO . (1999). 11q23.1 and 11q25-qter YACs suppress tumour growth in vivo. Oncogene 18: 1157–1164.
Koreth J, Bakkenist CJ, McGee JO . (1997). Allelic deletions at chromosome 11q22–q23.1 and 11q25-qterm are frequent in sporadic breast but not colorectal cancers. Oncogene 14: 431–437.
Kourlas PJ, Strout MP, Becknell B, Veronese ML, Croce CM, Theil KS et al. (2000). Identification of a gene at 11q23 encoding a guanine nucleotide exchange factor: evidence for its fusion with MLL in acute myeloid leukemia. Proc Natl Acad Sci USA 97: 2145–2150.
Kuramochi M, Fukuhara H, Nobukuni T, Kanbe T, Maruyama T, Ghosh HP et al. (2001). TSLC1 is a tumor-suppressor gene in human non-small-cell lung cancer. Nat Genet 27: 427–430.
Lee AS, Ho GH, Oh PC, Balram C, Ooi LL, Lim DT et al. (2003). Founder mutation in the BRCA1 gene in Malay breast cancer patients from Singapore. Hum Mutat 22: 178.
Lee AS, Rudduck-Sivaswaren C, Khun-Hong Lie D, Li-Ming Chua C, Tien SL, Morsberger L et al. (2004). Overlapping deletion regions at 11q23 in myelodysplastic syndrome and chronic lymphocytic leukemia, characterized by a novel BAC probe set. Cancer Genet Cytogenet 153: 151–157.
Lee AS, Seo YC, Chang A, Tohari S, Eu KW, Seow-Choen F et al. (2000). Detailed deletion mapping at chromosome 11q23 in colorectal carcinoma. Br J Cancer 83: 750–755.
Liu X, Wang H, Eberstadt M, Schnuchel A, Olejniczak ET, Meadows RP et al. (1998). NMR structure and mutagenesis of the N-terminal Dbl homology domain of the nucleotide exchange factor Trio. Cell 95: 269–277.
Martin ES, Cesari R, Pentimalli F, Yoder K, Fishel R, Himelstein AL et al. (2003). The BCSC-1 locus at chromosome 11q23–q24 is a candidate tumor suppressor gene. Proc Natl Acad Sci USA 100: 11517–11522.
Massion PP, Kuo WL, Stokoe D, Olshen AB, Treseler PA, Chin K et al. (2002). Genomic copy number analysis of non-small cell lung cancer using array comparative genomic hybridization: implications of the phosphatidylinositol 3-kinase pathway. Cancer Res 62: 3636–3640.
Murakami Y, Nobukuni T, Tamura K, Maruyama T, Sekiya T, Arai Y et al. (1998). Localization of tumor suppressor activity important in non small cell lung carcinoma on chromosome 11q. Proc Natl Acad Sci USA 95: 8153–8158.
Negrini M, Rasio D, Hampton GM, Sabbioni S, Rattan S, Carter SL et al. (1995). Definition and refinement of chromosome 11 regions of loss of heterozygosity in breast cancer: identification of a new region at 11q23.3. Cancer Res 55: 3003–3007.
Negrini M, Sabbioni S, Possati L, Rattan S, Corallini A, Barbanti-Brodano G et al. (1994). Suppression of tumorigenicity of breast cancer cells by microcell-mediated chromosome transfer: studies on chromosomes 6 and 11. Cancer Res 54: 1331–1336.
Plass C . (2002). Cancer epigenomics. Hum Mol Genet 11: 2479–2488.
Pulido HA, Fakruddin MJ, Chatterjee A, Esplin ED, Beleno N, Martinez G et al. (2000). Identification of a 6-cM minimal deletion at 11q23.1–23.2 and exclusion of PPP2R1B gene as a deletion target in cervical cancer. Cancer Res 60: 6677–6682.
Radtke F, Raj K . (2003). The role of Notch in tumorigenesis: oncogene or tumour suppressor? Nat Rev Cancer 3: 756–767.
Rasio D, Negrini M, Manenti G, Dragani TA, Croce CM . (1995). Loss of heterozygosity at chromosome 11q in lung adenocarcinoma: identification of three independent regions. Cancer Res 55: 3988–3991.
Rice KL, Hormaeche I, Licht JD . (2007). Epigenetic regulation of normal and malignant hematopoiesis. Oncogene 26: 6697–6714.
Robertson G, Coleman A, Lugo TG . (1996). A malignant melanoma tumor suppressor on human chromosome 11. Cancer Res 56: 4487–4492.
Rossman KL, Der CJ, Sondek J . (2005). GEF means go: turning on RHO GTPases with guanine nucleotide-exchange factors. Nat Rev Mol Cell Biol 6: 167–180.
Shilatifard A . (2006). Chromatin modifications by methylation and ubiquitination: implications in the regulation of gene expression. Annu Rev Biochem 75: 243–269.
Siderovski DP, Strockbine B, Behe CI . (1999). Whither goest the RGS proteins? Crit Rev Biochem Mol Biol 34: 215–251.
Smietana K, Kasztura M, Paduch M, Derewenda U, Derewenda ZS, Otlewski J . (2008). Degenerate specificity of PDZ domains from RhoA-specific nucleotide exchange factors PDZRhoGEF and LARG. Acta Biochim Pol 55: 269–280.
Swiercz JM, Kuner R, Behrens J, Offermanns S . (2002). Plexin-B1 directly interacts with PDZ-RhoGEF/LARG to regulate RhoA and growth cone morphology. Neuron 35: 51–63.
Taya S, Inagaki N, Sengiku H, Makino H, Iwamatsu A, Urakawa I et al. (2001). Direct interaction of insulin-like growth factor-1 receptor with leukemia-associated RhoGEF. J Cell Biol 155: 809–820.
Ting AH, McGarvey KM, Baylin SB . (2006). The cancer epigenome—components and functional correlates. Genes Dev 20: 3215–3231.
Tomlinson IP, Nicolai H, Solomon E, Bodmer WF . (1996). The frequency and mechanism of loss of heterozygosity on chromosome 11q in breast cancer. J Pathol 180: 38–43.
Tomlinson IP, Strickland JE, Lee AS, Bromley L, Evans MF, Morton J et al. (1995). Loss of heterozygosity on chromosome 11 q in breast cancer. J Clin Pathol 48: 424–428.
Van Aelst L, D’Souza-Schorey C . (1997). Rho GTPases and signaling networks. Genes Dev 11: 2295–2322.
Whitehead IP, Campbell S, Rossman KL, Der CJ . (1997). Dbl family proteins. Biochim Biophys Acta 1332: F1–23.
Yamada T, Ohoka Y, Kogo M, Inagaki S . (2005). Physical and functional interactions of the lysophosphatidic acid receptors with PDZ domain-containing Rho guanine nucleotide exchange factors (RhoGEFs). J Biol Chem 280: 19358–19363.
Acknowledgements
We thank Dr Glenn Koh for assistance with review of case notes; YC Seo, Angela Chang, S Tohari, Irene HK Lim and Gan Yar Chze for excellent technical assistance; and Dr Eric Yap for helpful discussions. This study was supported by Grants from the National Medical Research Council (NMRC) of Singapore (NMRC/0076/1995, NMRC/0440/2000, NMRC/0570/2001, NMRC/0843/2004); SingHealth Foundation (SHF/FG235P/2005); the Singapore Cancer Society, SGH Research Fund, Cancer Research Education Fund, NCC and Department of Clinical Research, SGH, to AL. We gratefully acknowledge the grant support from the US Department of Energy under Contract No. DE-AC02-05CH11231, USAMRMC BC 061995; National Institutes of Health, National Cancer Institute (P50 CA 58207, P50 CA 83639, P30 CA 82103, U54 CA 112970, U24 CA 126477 P01 CA 64602); National Human Genome Research Institute (U24 CA 126551) and SmithKline Beecham Corporation, to JWG.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Supplementary information
Rights and permissions
About this article
Cite this article
Ong, D., Ho, Y., Rudduck, C. et al. LARG at chromosome 11q23 has functional characteristics of a tumor suppressor in human breast and colorectal cancer. Oncogene 28, 4189–4200 (2009). https://doi.org/10.1038/onc.2009.266
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2009.266
Keywords
This article is cited by
-
Targeting ARHGEF12 promotes neuroblastoma differentiation, MYCN degradation, and reduces tumorigenicity
Cellular Oncology (2023)
-
Investigation of the effects of the royal jelly on genomic demethylation and tumor suppressor genes in human cancer cells
Medical Oncology (2022)
-
Dysregulation of mitotic machinery genes precedes genome instability during spontaneous pre-malignant transformation of mouse ovarian surface epithelial cells
BMC Genomics (2016)
-
Expanding the genetic basis of copy number variation in familial breast cancer
Hereditary Cancer in Clinical Practice (2014)
-
Genome-wide investigation of gene–environment interactions in colorectal cancer
Human Genetics (2013)