Abstract
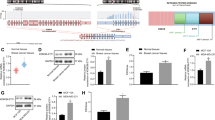
The transcription factor Ets-1 is implicated in various physiological processes and invasive pathologies. We identified a novel variant of ets-1, ets-1Δ(III–VI), resulting from the alternative splicing of exons III to VI. This variant encodes a 27 kDa isoform, named Ets-1 p27. Ets-1 p27 lacks the threonine-38 residue, the Pointed domain and the transactivation domain, all of which are required for the transactivation of Ets-1 target genes. Both inhibitory domains surrounding the DNA-binding domain are conserved, suggesting that Ets-1 p27, like the full-length Ets-1 p51 isoform, is autoinhibited for DNA binding. We showed that Ets-1 p27 binds DNA in the same way as Ets-1 p51 does and that it acts both at a transcriptional and a subcellular localization level, thereby constituting a dual-acting dominant negative of Ets-1 p51. Ets-1 p27 blocks Ets-1 p51-mediated transactivation of target genes and induces the translocation of Ets-1 p51 from the nucleus to the cytoplasm. Furthermore, Ets-1 p27 overexpression represses the tumor properties of MDA-MB-231 mammary carcinoma cells in correlation with the known implication of Ets-1 in various cellular mechanisms. Thus the dual-acting dominant-negative function of Ets-1 p27 gives to the Ets-1 p27/Ets-1 p51 ratio a determining effect on cell fate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Alipov G, Nakayama T, Ito M, Kawai K, Naito S, Nakashima M et al. (2005). Overexpression of Ets-1 proto-oncogene in latent and clinical prostatic carcinomas. Histopathology 46: 202–208.
Araud T, Genolet R, Jaquier-Gubler P, Curran J . (2007). Alternatively spliced isoforms of the human elk-1 mRNA within the 5′ UTR: implications for ELK-1 expression. Nucleic Acids Res 35: 4649–4663.
Baillat D, Begue A, Stehelin D, Aumercier M . (2002). ETS-1 transcription factor binds cooperatively to the palindromic head to head ETS-binding sites of the stromelysin-1 promoter by counteracting autoinhibition. J Biol Chem 277: 29386–29398.
Baillat D, Laitem C, Leprivier G, Margerin C, Aumercier M . (2009). Ets-1 binds cooperatively to the palindromic Ets-binding sites in the p53 promoter. Biochem Biophys Res Commun 378: 213–217.
Baillat D, Leprivier G, Regnier D, Vintonenko N, Begue A, Stehelin D et al. (2006). Stromelysin-1 expression is activated in vivo by Ets-1 through palindromic head-to-head Ets binding sites present in the promoter. Oncogene 25: 5764–5776.
Buttice G, Kurkinen M . (1993). A polyomavirus enhancer A-binding protein-3 site and Ets-2 protein have a major role in the 12-O-tetradecanoylphorbol-13-acetate response of the human stromelysin gene. J Biol Chem 268: 7196–7204.
Cho JY, Akbarali Y, Zerbini LF, Gu X, Boltax J, Wang Y et al. (2004). Isoforms of the Ets transcription factor NERF/ELF-2 physically interact with AML1 and mediate opposing effects on AML1-mediated transcription of the B cell-specific blk gene. J Biol Chem 279: 19512–19522.
Coutte L, Monte D, Imai K, Pouilly L, Dewitte F, Vidaud M et al. (1999). Characterization of the human and mouse ETV1/ER81 transcription factor genes: role of the two alternatively spliced isoforms in the human. Oncogene 18: 6278–6286.
Cowley DO, Graves BJ . (2000). Phosphorylation represses Ets-1 DNA binding by reinforcing autoinhibition. Genes Dev 14: 366–376.
De Haro L, Janknecht R . (2005). Cloning of the murine ER71 gene (Etsrp71) and initial characterization of its promoter. Genomics 85: 493–502.
de Nigris F, Mega T, Berger N, Barone MV, Santoro M, Viglietto G et al. (2001). Induction of ETS-1 and ETS-2 transcription factors is required for thyroid cell transformation. Cancer Res 61: 2267–2275.
Dittmer J . (2003). The biology of the Ets1 proto-oncogene. Mol Cancer 2: 29.
Fisher RJ, Fivash M, Casas-Finet J, Erickson JW, Kondoh A, Bladen SV et al. (1994). Real-time DNA binding measurements of the ETS1 recombinant oncoproteins reveal significant kinetic differences between the p42 and p51 isoforms. Protein Sci 3: 257–266.
Foulds CE, Nelson ML, Blaszczak AG, Graves BJ . (2004). Ras/mitogen-activated protein kinase signaling activates Ets-1 and Ets-2 by CBP/p300 recruitment. Mol Cell Biol 24: 10954–10964.
Hewett PW, Nishi K, Daft EL, Clifford Murray J . (2001). Selective expression of erg isoforms in human endothelial cells. Int J Biochem Cell Biol 33: 347–355.
Higashino F, Yoshida K, Noumi T, Seiki M, Fujinaga K . (1995). Ets-related protein E1A-F can activate three different matrix metalloproteinase gene promoters. Oncogene 10: 1461–1463.
Higuchi T, Bartel FO, Masuya M, Deguchi T, Henderson KW, Li R et al. (2007). Thymomegaly, microsplenia, and defective homeostatic proliferation of peripheral lymphocytes in p51-Ets1 isoform-specific null mice. Mol Cell Biol 27: 3353–3366.
Jayaraman G, Srinivas R, Duggan C, Ferreira E, Swaminathan S, Somasundaram K et al. (1999). p300/cAMP-responsive element-binding protein interactions with ets-1 and ets-2 in the transcriptional activation of the human stromelysin promoter. J Biol Chem 274: 17342–17352.
John SA, Clements JL, Russell LM, Garrett-Sinha LA . (2007). Ets-1 regulates plasma cell differentiation by interfering with the activity of the transcription factor Blimp-1. J Biol Chem 283: 951–962.
Jorcyk CL, Watson DK, Mavrothalassitis GJ, Papas TS . (1991). The human ETS1 gene: genomic structure, promoter characterization and alternative splicing. Oncogene 6: 523–532.
Katayama S, Nakayama T, Ito M, Naito S, Sekine I . (2005). Expression of the ets-1 proto-oncogene in human breast carcinoma: differential expression with histological grading and growth pattern. Histol Histopathol 20: 119–126.
Kherrouche Z, Blais A, Ferreira E, De Launoit Y, Monte D . (2006). ASK-1 (apoptosis signal-regulating kinase 1) is a direct E2F target gene. Biochem J 396: 547–556.
Kita D, Takino T, Nakada M, Takahashi T, Yamashita J, Sato H . (2001). Expression of dominant-negative form of Ets-1 suppresses fibronectin-stimulated cell adhesion and migration through down-regulation of integrin alpha5 expression in U251 glioma cell line. Cancer Res 61: 7985–7991.
Kitange G, Kishikawa M, Nakayama T, Naito S, Iseki M, Shibata S . (1999). Expression of the Ets-1 proto-oncogene correlates with malignant potential in human astrocytic tumors. Mod Pathol 12: 618–626.
Koizumi S, Fisher RJ, Fujiwara S, Jorcyk C, Bhat NK, Seth A et al. (1990). Isoforms of the human ets-1 protein: generation by alternative splicing and differential phosphorylation. Oncogene 5: 675–681.
Laitem C, Choul-Li S, Baillat D, Begue A, Aumercier M . (2008). Efficient system for biotinylated recombinant Ets-1 production in Escherichia coli: a useful tool for studying interactions between Ets-1 and its partners. Protein Expr Purif 62: 53–63.
Lee GM, Donaldson LW, Pufall MA, Kang HS, Pot I, Graves BJ et al. (2005). The structural and dynamic basis of Ets-1 DNA binding autoinhibition. J Biol Chem 280: 7088–7099.
Lei W, Jaramillo RJ, Harrod KS . (2007). Transactivation of lung lysozyme expression by Ets family member ESE-1. Am J Physiol Lung Cell Mol Physiol 293: L1359–L1368.
Li B, Lager J, Wang D, Li D . (2007). Ets-1 participates in and facilitates developmental expression of hypoxia-induced mitogenic factor in mouse lung. Front Biosci 12: 2269–2278.
Li R, Pei H, Papas T . (1999). The p42 variant of ETS1 protein rescues defective Fas-induced apoptosis in colon carcinoma cells. Proc Natl Acad Sci USA 96: 3876–3881.
Lulli V, Romania P, Morsilli O, Gabbianelli M, Pagliuca A, Mazzeo S et al. (2006). Overexpression of Ets-1 in human hematopoietic progenitor cells blocks erythroid and promotes megakaryocytic differentiation. Cell Death Differ 13: 1064–1074.
Maroulakou IG, Bowe DB . (2000). Expression and function of Ets transcription factors in mammalian development: a regulatory network. Oncogene 19: 6432–6442.
Mylona EE, Alexandrou PT, Giannopoulou IA, Rafailidis PI, Markaki S, Keramopoulos A et al. (2006). Study of the topographic distribution of ets-1 protein expression in invasive breast carcinomas in relation to tumor phenotype. Cancer Detect Prev 30: 111–117.
Nakayama T, Ito M, Ohtsuru A, Naito S, Sekine I . (2001). Expression of the ets-1 proto-oncogene in human colorectal carcinoma. Mod Pathol 14: 415–422.
O’Leary DA, Koleski D, Kola I, Hertzog PJ, Ristevski S . (2005). Identification and expression analysis of alternative transcripts of the mouse GA-binding protein (Gabp) subunits alpha and beta1. Gene 344: 79–92.
Pognonec P, Boulukos KE, Bosselut R, Boyer C, Schmitt-Verhulst AM, Ghysdael J . (1990). Identification of a Ets1 variant protein unaffected in its chromatin and in vitro DNA binding capacities by T cell antigen receptor triggering and intracellular calcium rises. Oncogene 5: 603–610.
Pufall MA, Graves BJ . (2002). Autoinhibitory domains: modular effectors of cellular regulation. Annu Rev Cell Dev Biol 18: 421–462.
Raffetseder U, Wernert N, Ostendorf T, van Roeyen C, Rauen T, Behrens P et al. (2004). Mesangial cell expression of proto-oncogene Ets-1 during progression of mesangioproliferative glomerulonephritis. Kidney Int 66: 622–632.
Ray-Gallet D, Tavitian A, Moreau-Gachelin F . (1996). An alternatively spliced isoform of the Spi-B transcription factor. Biochem Biophys Res Commun 223: 257–263.
Redlich K, Kiener HP, Schett G, Tohidast-Akrad M, Selzer E, Radda I et al. (2001). Overexpression of transcription factor Ets-1 in rheumatoid arthritis synovial membrane: regulation of expression and activation by interleukin-1 and tumor necrosis factor alpha. Arthritis Rheum 44: 266–274.
Rowe A, Propst F . (1992). Ets-1 and Ets-2 protooncogene expression in theca cells of the adult mouse ovary. Exp Cell Res 202: 199–202.
Sarrazin S, Starck J, Gonnet C, Doubeikovski A, Melet F, Morle F . (2000). Negative and translation termination-dependent positive control of FLI-1 protein synthesis by conserved overlapping 5' upstream open reading frames in Fli-1 mRNA. Mol Cell Biol 20: 2959–2969.
Sasaki K, Nakamura Y, Maki K, Waga K, Nakamura F, Arai H et al. (2004). Functional analysis of a dominant-negative DeltaETS TEL/ETV6 isoform. Biochem Biophys Res Commun 317: 1128–1137.
Seidel JJ, Graves BJ . (2002). An ERK2 docking site in the Pointed domain distinguishes a subset of ETS transcription factors. Genes Dev 16: 127–137.
Shin C, Manley JL . (2004). Cell signalling and the control of pre-mRNA splicing. Nat Rev Mol Cell Biol 5: 727–738.
Smith CW, Valcarcel J . (2000). Alternative pre-mRNA splicing: the logic of combinatorial control. Trends Biochem Sci 25: 381–388.
Stamm S, Ben-Ari S, Rafalska I, Tang Y, Zhang Z, Toiber D et al. (2005). Function of alternative splicing. Gene 344: 1–20.
Takai N, Miyazaki T, Fujisawa K, Nasu K, Miyakawa I . (2000). Expression of c-Ets1 is associated with malignant potential in endometrial carcinoma. Cancer 89: 2059–2067.
Takai N, Miyazaki T, Nishida M, Nasu K, Miyakawa I . (2002). c-Ets1 is a promising marker in epithelial ovarian cancer. Int J Mol Med 9: 287–292.
Takai N, Ueda T, Narahara H, Miyakawa I . (2006). Expression of c-Ets1 protein in normal human placenta. Gynecol Obstet Invest 61: 15–20.
Taniguchi H, Fujiwara Y, Doki Y, Sugita Y, Sohma I, Miyata H et al. (2007). Gene therapy using ets-1 transcription factor decoy for peritoneal dissemination of gastric cancer. Int J Cancer 121: 1609–1617.
Teruyama K, Abe M, Nakano T, Iwasaka-Yagi C, Takahashi S, Yamada S et al. (2001). Role of transcription factor Ets-1 in the apoptosis of human vascular endothelial cells. J Cell Physiol 188: 243–252.
Valter MM, Hugel A, Huang HJ, Cavenee WK, Wiestler OD, Pietsch T et al. (1999). Expression of the Ets-1 transcription factor in human astrocytomas is associated with Fms-like tyrosine kinase-1 (Flt-1)/vascular endothelial growth factor receptor-1 synthesis and neoangiogenesis. Cancer Res 59: 5608–5614.
Wasylyk C, Gutman A, Nicholson R, Wasylyk B . (1991). The c-Ets oncoprotein activates the stromelysin promoter through the same elements as several non-nuclear oncoproteins. EMBO J 10: 1127–1134.
Acknowledgements
We thank Isabelle Roland, Anne-Claire Flourens, Nathalie Tomavo and Arnaud Legrand for technical assistance, and Dr Yvan de Launoit for stimulating discussions. We thank the Microscopy Facility of the Institut Pasteur de Lille Campus. This work was supported by the Centre National de la Recherche Scientifique (CNRS) and by grants from la Ligue contre le Cancer-Comité du Pas-de-Calais and from the Fondation pour la Recherche Médicale-Comité Nord-Pas-de-Calais (FRM). The Ministère de la Recherche et de l’Enseignement Supérieur provided a student fellowship to Clélia Laitem. La Ligue contre le Cancer provided a student fellowship to Gabriel Leprivier. The CNRS and the Conseil Régional du Nord-Pas-de-Calais provided a PhD fellowship (BDI) to Souhaila Choul-li.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Supplementary information
Rights and permissions
About this article
Cite this article
Laitem, C., Leprivier, G., Choul-Li, S. et al. Ets-1 p27: a novel Ets-1 isoform with dominant-negative effects on the transcriptional properties and the subcellular localization of Ets-1 p51. Oncogene 28, 2087–2099 (2009). https://doi.org/10.1038/onc.2009.72
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2009.72
Keywords
This article is cited by
-
Genome-wide identification and characteristic analysis of ETS gene family in blood clam Tegillarca granosa
BMC Genomics (2023)
-
Identification and characterization of novel ETV4 splice variants in prostate cancer
Scientific Reports (2023)
-
Splice variants as novel targets in pancreatic ductal adenocarcinoma
Scientific Reports (2017)
-
Review of Ets1 structure, function, and roles in immunity
Cellular and Molecular Life Sciences (2013)
-
Regulation of protein function by ‘microProteins’
EMBO reports (2011)