Abstract
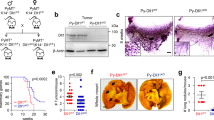
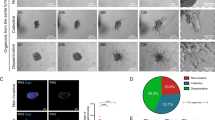
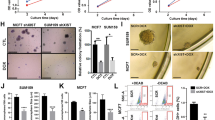
Inflammatory breast carcinoma (IBC) is characterized by exaggerated lymphovascular invasion (LVI), recapitulated in our human xenograft, MARY-X. This model exhibited lymphovascular emboli in vivo and corresponding spheroids in vitro. Owing to the morphological and gene profile resemblance of these spheroids to embryonal blastocysts, we wondered whether they might exhibit embryonic stem cell signaling. Specifically we investigated Notch and observed selective Notch 3 activation by expression profiling, reverse transcriptase– and real-time PCR, western blot and immunofluorescence in vitro, and immunohistochemistry in vivo. Notch 3 intracellular domain (N3icd) and six target genes, HES-5, HEY-1, c-Myc, Deltex-1, NRARP and PBX1, markedly increased in MARY-X. In addition, a significant percentage of MARY-X cells expressed aldehyde dehydrogenase (ALDH), a stem cell marker. Only the ALDH+ cells were capable of secondary spheroidgenesis, tumorigenicity and self-renewal. Inhibiting Notch 3 activation in vitro with γ-secretase inhibitors (GSIs) or small interfering RNA resulted in a downregulation of Notch target genes, including CD133, and an induction of caspase 3-mediated apoptosis. Transfection of N3icd but not Notch 1 intracellular domain into normal human mammary epithelial cells resulted in increased expression of Notch target genes and induction of spheroidgenesis. GSI in vivo resulted in inhibitory but diffusion-limited effects on Notch 3 signaling, resulting in xenograft growth reduction. The lymphovascular emboli of human IBC exhibited dual N3icd and ALDH1 immunoreactivities independently of molecular subtype. This Notch 3 addiction of lymphovascular emboli might be exploited in future therapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Alpaugh ML, Barsky SH . (2002). Reversible model of spheroid formation allows for high efficiency of gene delivery ex vivo and accurate gene assessment in vivo. Hum Gene Ther 13: 1245–1258.
Alpaugh ML, Tomlinson JS, Shao ZM, Barsky SH . (1999). A novel human xenograft model of inflammatory breast cancer. Cancer Res 59: 5079–5084.
Alpaugh ML, Tomlinson JS, Kasraeian S, Barsky SH . (2002a). Cooperative role of E-cadherin and sialyl-Lewis X/A-deficient MUC1 in the passive dissemination of tumor emboli in inflammatory breast carcinoma. Oncogene 21: 3631–3643.
Alpaugh ML, Tomlinson JS, Ye Y, Barsky SH . (2002b). Relationship of sialyl-Lewis x/a underexpression and E-cadherin overexpression in the lymphovascular embolus of inflammatory breast carcinoma. Am J Pathol 161: 619–628.
Androutsellis-Theotokis A, Leker RR, Soldner F, Hoeppner DJ, Ravin R, Poser SW et al. (2006). Notch signaling regulates stem cell numbers in vitro and in vivo. Nature 442: 823–826.
Ayyanan A, Civenni G, Ciarloni L, Morel C, Mueller N, Lefort K et al. (2006). Increased Wnt signaling triggers oncogenic conversion of human breast epithelial cells by a notch-dependent mechanism. Proc Natl Acad Sci USA 103: 3799–3804.
Barsky SH . (2003). Myoepithelial mRNA expression profiling reveals a common tumor suppressor phenotype. Exp Mol Pathol 74: 113–122.
Bellavia D, Checquolo S, Campese AF, Felli MP, Gulino A, Screpanti I . (2008). Notch3: from subtle structural differences to functional diversity. Oncogene 27: 5092–5098.
Bertucci F, Finetti P, Cervera N, Charafe-Jauffret E, Mamessier E, Adelaide J et al. (2006). Gene expression profiling shows medullary breast cancer is a subgroup of basal breast cancers. Cancer Res 66: 4636–4644.
Bertucci F, Finetti P, Rougemont J, Charafe-Jauffret, Cervera N, Tarpin C et al. (2005). Gene expression profiling identifies molecular subtypes of inflammatory breast cancer. Cancer Res 65: 2170–2178.
Bonnet D, Dick J . (1997). Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat Med 3: 730–737.
Bray SJ . (2006). Notch signaling: a simple pathway becomes complex. Nat Rev Mol Cell Biol 7: 678–689.
Callahan R, Egan SE . (2004). Notch signaling in mammary development and oncogenesis. J Mammary Gland Biol Neoplasia 9: 145–163.
Cariati M, Bennett-Britton TM, Pinder SE, Purushotham AD . (2005). ‘Inflammatory’ breast cancer. Surg Oncol 14: 133–143.
Chiba S . (2006). Notch signaling in stem cell systems. Stem Cells 24: 2437–2447.
Clark AT, Rodriguez RT, Bodnar MS . (2004). Human STELLAR, NANOG, and GDF3 genes are expressed in pluripotent cells and map to chromosome 12p13, a hotspot for teratocarcinoma. Stem Cells 22: 169–179.
Cuevas IC, Slocum AL, Jun P, Costello JF, Bollen AW, Riggins GJ et al. (2005). Meningioma transcript profiles reveal deregulated notch signaling pathway. Cancer Res 65: 5070–5075.
Curry CL, Reed LL, Golde TE, Miele L, Nickoloff BJ, Foreman KE . (2005). Gamma-secretase inhibitor blocks notch activation and induces apoptosis in Kaposi's sarcoma tumor cells. Oncogene 24: 6333–6344.
Dievart A, Beaulieu N, Jolicoeur P . (1999). Involvement of notch1 in the development of mouse mammary tumors. Oncogene 18: 5973–5981.
Dontu G, Abdallah WM, Foley JM, Jackson KW, Michael FC, Kawamura MJ et al. (2003). In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev 17: 1253–1270.
Dontu G, Jackson KW, McNicholas E, Kawamura MJ, Abdallah WM, Wicha MS . (2004). Role of notch signaling in cell-fate determination of human mammary stem/progenitor cells. Breast Cancer Res 6: R605–R615.
Farnie G, Clarke RB, Spence K, Pinnock N, Brennan K, Anderson NG et al. (2007). Novel cell culture technique for primary ductal carcinoma in situ: role of notch and epidermal growth factor receptor signaling pathways. J Natl Cancer Inst 99: 616–627.
Gallahan D, Callahan R . (1997). The mouse mammary tumor associated gene INT3 is a unique member of the NOTCH gene family (NOTCH4). Oncogene 14: 1883–1890.
Ginestier C, Hur MH, Charafe-Jauffret E, Monville F, Dutcher J, Brown M et al. (2007). ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell 1: 555–567.
Giraldez AJ, Perez L, Cohen SM . (2002). A naturally occurring alternative product of the mastermind locus that represses notch signaling. Mech Dev 115: 101–105.
Girard L, Hanna Z, Beaulieu N, Hoemann CD, Simard C, Kozak CA et al. (1996). Frequent provirus insertional mutagenesis of notch1 in thymomas of MMTVD/myc transgenic mice suggests a collaboration of c-myc and notch1 for oncogenesis. Genes Dev 10: 1930–1944.
Gordon WR, Vardar-Ulu D, Histen G, Sanchez-Irizarry C, Aster JC, Blacklow SC . (2007). Structural basis for autoinhibition of notch. Nat Struct Mol Biol 14: 295–300.
Haruki N, Kawaguchi KS, Eichenberger S, Massion PP, Olson S, Gonzalez A et al. (2005). Dominant-negative notch3 receptor inhibits mitogen-activated protein kinase pathway and the growth of human lung cancers. Cancer Res 65: 3555–3561.
Hu C, Dievart A, Lupien M, Calvo E, Tremblay G, Jolicoeur P . (2006). Overexpression of activated murine notch1 and notch3 in transgenic mice blocks mammary gland development and induces mammary tumors. Am J Pathol 168: 973–990.
Iso T, Kedes L, Hamamori Y . (2003). HES and HERP families: multiple effectors of the notch signaling pathway. J Cell Physiol 194: 237–255.
Jain AN, Tokuyasu TA, Snijders AM, Segraves R, Albertson DG, Pinkel D . (2002). Fully automatic quantification of microarray image data. Genome Res 12: 325–332.
Jhappan C, Gallahan D, Stahle C, Chu E, Smith GH, Merlino G et al. (1992). Expression of an activated Notch-related int-3 transgene interferes with cell differentiation and induces neoplastic transformation in mammary and salivary glands. Genes Dev 6: 345–355.
Klinakis A, Szabolcs M, Politi K, Kiaris H, Atavanis-Tsakonas S, Efstratiadis A . (2006). Myc is a Notch1 transcriptional target and a requisite for Notch1-induced mammary tumorigenesis in mice. Proc Natl Acad Sci USA 103: 9262–9267.
Krzywinski M, Bosdet I, Smailus D, Chiu R, Mathewson C, Wye N et al. (2004). A set of BAC clones spanning the human genome. Nucelic Acids Res 32: 3651–3660.
Kopan R, Llagan MXG . (2009). The canonical notch signaling pathway: unfolding the activation mechanism. Cell 137: 216–233.
Kubota H, Avarbock MR, Brinster R . (2003). Spermatogonial stem cells share some, but not all, phenotypic and functional characteristics with other stem cells. Proc Natl Acad Sci USA 100: 6487–6492.
Laurie AB, Tong IL, Megan FC, Sarah EJ, Stuart SL, Jacob PZ et al. (2005). Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122: 947–956.
Leong KG, Karsan A . (2006). Recent insights into the role of notch signaling in tumorigenesis. Blood 107: 2223–2233.
Leong KG, Niessen K, Kulic I, Raouf A, Eaves C, Pollet I et al. (2007). Jagged1-mediated notch activation induces epithelial-to-mesenchymal transition through slug-induced repression of E-cadherin. J Exp Med 204: 2935–2948.
Miyamoto Y, Maitra A, Ghosh B, Zechner U, Argani P, Iacobuzio-Donahue CA et al. (2003). Notch mediates TGF alpha-induced changes in epithelial differentiation during pancreatic tumorigenesis. Cancer Cell 3: 565–576.
Palangie T, Mosseri V, Mihura J, Campana F, Beuzeboc P, Dorval T et al. (1994). Prognostic factors in inflammatory breast cancer and therapeutic implications. Eur J Cancer 30A: 921–927.
Palomero T, Lim WK, Odom DT, Sulis ML, Real PJ, Margolin A et al. (2006). Notch1 directly regulates c-MYC and activates a feed-forward-loop transcriptional network promoting leukemic cell growth. Proc Natl Acad Sci USA 103: 18261–18266.
Park JT, Li M, Nakayama K, Mao TL, Davidson B, Zhang Z et al. (2006). Notch 3 gene amplification in ovarian cancer. Cancer Res 66: 6312–6318.
Park JT, Shih IM, Wang TL . (2008). Identification of Pbx1, a potential oncogene, as a notch 3 target gene in ovarian cancer. Cancer Res 68: 8852–8860.
Politi K, Feirt N, Kitajewski J . (2006). Notch in mammary gland development and breast cancer. Semin Cancer Biol 14: 341–347.
Purow BW, Haque RM, Noel MW, Su Q, Burdick MJ, Lee J et al. (2005). Expression of Notch-1 and its ligands, delta-like-1 and jagged-1, is critical for glioma cell survival and proliferation. Cancer Res 65: 2353–2363.
Rachel E, Maya S, Micha D, Ofra Y, Joseph IE, Nissim B . (2001). Establishment of human embryonic stem cell-transfected clones carrying a marker for undifferentiated cells. Curr Biol 11: 514–518.
Reya T, Morrison SJ, Clarke MF, Weissman IL . (2001). Stem cells, cancer, and cancer stem cells. Nature 414: 105–111.
Rizzo P, Osipo C, Foreman K, Golde T, Osborne B, Miele L . (2008). Rationale targeting of notch signaling in cancer. Oncogene 27: 5124–5131.
Sanchez-Irizarry C, Carpenter AC, Weng AP, Pear WS, Aster JC, Blacklow SC . (2004). Notch subunit heterodimerization and prevention of ligand-independent proteolytic activation depend, respectively, on a novel domain and the LNR repeats. Mol Cell Biol 24: 9265–9273.
Sansone P, Storci G, Giovannini C, Pandolfi S, Pianetti S, Taffurelli M et al. (2007a). p66Shc/Notch-3 interplay controls self-renewal and hypoxia survival in human stem/progenitor cells of the mammary gland expanded in vitro as mammospheres. Stem Cells 25: 807–815.
Sansone P, Storci G, Tavolari S, Guarnieri T, Giovannini C, Taffurelli M et al. (2007b). Il-6 triggers malignant features in mammospheres from human ductal breast carcinoma and normal mammary gland. J Clin Invest 117: 3988–4002.
Santagata S, Demichelis F, Riva A, Varambally S, Hofer MD, Kutok JL et al. (2004). JAGGED1 expression is associated with prostate cancer metastasis and recurrence. Cancer Res 64: 6854–6857.
Sheila K, Cynthia H, Clarke ID, Jeremy A, Jane B, Takuichiro H et al. (2004). Identification of human brain tumour initiating cells. Nature 432: 396–401.
Shih IeM, Wang TL . (2007). Notch signaling, gamma-secretase inhibitors, and cancer therapy. Cancer Res 67: 1879–1882.
Shipitsin M, Campbell LL, Argani P, Weremowicz S, Bloushtain-Qimron N, Yao J et al. (2007). Molecular definition of breast tumor heterogeneity. Cancer Cell 11: 259–273.
Sternlicht M, Safarians S, Calcaterra TC, Barsky SH . (1996). Establishment and characterization of a novel human myoepithelial cell line and matrix-producing xenograft from a parotid basal cell adenocarcinoma. In vitro Cell Dev Biol 32: 550–563.
Stylianou S, Clarke RB, Brennan K . (2006). Aberrant activation of notch signaling in human breast cancer. Cancer Res 66: 1517–1525.
Sun Y, Lowther W, Kato K, Bianco C, Kenney N, Strizzi L et al. (2005). Notch4 intracellular domain binding to Smad3 and inhibition of the TGF-β signaling. Oncogene 24: 5365–5374.
Tomlinson JS, Alpaugh ML, Barsky SH . (2001). An intact overexpressed E-cadherin/alpha, beta-catenin axis characterizes the lymphovascular emboli of inflammatory breast carcinoma. Cancer Res 61: 5231–5241.
Weng AP, Ferrando AA, Lee W, Morris JP, Silverman LB, Sanchez-Irizarry C et al. (2004). Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 306: 269–271.
Wicha MS, Liu SL, Dontu G . (2006). Cancer stem cells: an old idea-a paradigm shift. Cancer Res 66: 1883–1890.
Xiao Y, Yearsley K, Jones S, Barsky SH . (2008). The lymphovascular embolus of inflammatory breast cancer expresses a stem cell-like phenotype. Am J Pathol 173: 561–574.
Yamaguchi N, Oyama T, Ito E, Satoh H, Azuma S, Hayashi M et al. (2008). Notch3 signaling pathway plays crucial roles in the proliferation of erbB2-negative human breast cancer cells. Cancer Res 68: 1881–1888.
Zagouras P, Stifani S, Blaumueller CM, Carcangiu ML, Srtavanis-Tsakonas A . (1995). Alterations in notch signaling in neoplastic lesions of the human cervix. Proc Natl Acad Sci USA 92: 6414–6418.
Acknowledgements
This study was supported by the American Airlines-Susan G Komen for the Cure Promise Grant KGO81287, Department of Defense Breast Cancer Research Program Grants BC990959, BC024258, BC053405, the Strategic Initiative Grant Program at Ohio State and The Donald A Senhauser Endowment. Figures 1a, b, c and 2e were, in part, reprinted from Am J Pathol 2008, 173: 561–574 with permission from the American Society For Investigative Pathology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Xiao, Y., Ye, Y., Zou, X. et al. The lymphovascular embolus of inflammatory breast cancer exhibits a Notch 3 addiction. Oncogene 30, 287–300 (2011). https://doi.org/10.1038/onc.2010.405
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.405
Keywords
This article is cited by
-
KK-LC-1 as a therapeutic target to eliminate ALDH+ stem cells in triple negative breast cancer
Nature Communications (2023)
-
Geometric tumor embolic budding characterizes inflammatory breast cancer
Breast Cancer Research and Treatment (2023)
-
Comparative transcriptional analyses of preclinical models and patient samples reveal MYC and RELA driven expression patterns that define the molecular landscape of IBC
npj Breast Cancer (2022)
-
Inflammatory breast cancer biology: the tumour microenvironment is key
Nature Reviews Cancer (2018)
-
Inflammatory breast cancer: a model for investigating cluster-based dissemination
npj Breast Cancer (2017)