Abstract
The DNA glycosylase MUTYH (mutY homolog (Escherichia coli)) counteracts the mutagenic effects of 8-oxo-7,8-dihydroguanine (8-oxodG) by removing adenine (A) misincorporated opposite the oxidized purine. Biallelic germline mutations in MUTYH cause the autosomal recessive MUTYH-associated adenomatous polyposis (MAP). Here we designed new tools to investigate the biochemical defects and biological consequences associated with different MUTYH mutations in human cells. To identify phenotype(s) associated with MUTYH mutations, lymphoblastoid cell lines (LCLs) were derived from seven MAP patients harboring missense as well as truncating mutations in MUTYH. These included homozygous p.Arg245His, p.Gly264TrpfsX7 or compound heterozygous variants (p.Gly396Asp/Arg245Cys, p.Gly396Asp/Tyr179Cys, p.Gly396Asp/Glu410GlyfsX43, p.Gly264TrpfsX7/Ala385ProfsX23 and p.Gly264TrpfsX7/Glu480del). DNA glycosylase assays of MAP LCL extracts confirmed that all these variants were defective in removing A from an 8-oxoG:A DNA substrate, but retained wild-type OGG1 activity. As a consequence of this defect, MAP LCLs accumulated DNA 8-oxodG in their genome and exhibited a fourfold increase in spontaneous mutagenesis at the PIG-A gene compared with LCLs from healthy donors. They were also hypermutable by KBrO3—a source of DNA 8-oxodG—indicating that the relatively modest spontaneous mutator phenotype associated with MUTYH loss can be significantly enhanced by conditions of oxidative stress. These observations identify accumulation of DNA 8-oxodG and a mutator phenotype as likely contributors to the pathogenesis of MUTYH variants.
Similar content being viewed by others
Introduction
The DNA glycosylase MUTYH is part of the base excision repair pathway, which counteracts the promutagenic properties of 8-oxo-7,8-dihydroguanine (8-oxodG), one of the most common base lesions produced by reactive oxygen species. It prevents G:C to T:A transversions by excising adenine (A) misincorporated opposite 8-oxodG in template DNA.1, 2, 3 The human MUTYH (mutY homolog (Escherichia coli)) gene, is located on chromosome 1p34.3–p32.1 with an open reading frame of 1.6 kb and 16 exons.4 Germline biallelic mutations in this gene are implicated in MUTYH-associated polyposis (MAP), a recessively heritable colorectal polyposis, linked to an increased risk of colorectal cancer (CRC).5, 6, 7, 8, 9 In contrast to familial adenomatous polyposis patients (FAP) who have germline mutations in the adenomatous polyposis coli gene, MAP patients show distinctive somatic G:C to T:A transversions in the adenomatous polyposis coli gene of their tumors. Following the initial discoveries, other groups confirmed these findings and concluded that biallelic mutations in the MUTYH gene drive genomic instability in colorectal epithelial cells resulting in an increased risk of neoplastic transformation in the colon of MAP patients.10, 11, 12, 13
To date, >300 MUTYH unique variants among MAP patients and/or controls have been described in the LOVD database,14 with the p.Tyr179Cys and p.Gly396Asp variants as the most common documented mutations in Caucasian populations.5, 15, 16, 17 Most MUTYH variants are missense mutations and their effects on protein function can be difficult to predict when present in homozygosity and even more in a genetic condition of compound heterozygosity. In addition, there is a need to set up functional tests that take into account the biological context in which the variants are present. These might represent the basis for establishing genotype–phenotype correlations, thus providing an important tool in clinical practice. Results of our previous study based on an assay in which single mutant MUTYH proteins were expressed in mouse embryonic fibroblasts (MEFs) derived from Mutyh-null mice, showed that accumulation of 8-oxodG is a common phenotype among the investigated MAP-associated variants.18 To extend these observations to human cells, we derived lymphoblastoid cell lines (LCLs) from MAP patients harboring missense and truncating MUTYH mutations, including compound heterozygotes—a frequent occurrence among Italian MAP patients. We investigated the biochemical defects and biological consequences associated with different MUTYH mutations. The results indicate that the mutator phenotype associated with loss of a functional MUTYH protein in human cells is relatively modest, but can be enhanced by exposure to an oxidative stress.
Results
Expression of MUTYH in LCLs derived from MAP patients
LCLs were established from seven MAP patients carrying biallelic, either compound heterozygous or homozygous, MUTYH variants. The phenotypic features of these patients are shown in Table 1. The family history in all biallelic mutation carriers corresponded to an autosomal recessive mode of inheritance, although four patients occurred as sporadic cases (FAP117, FAP278, FAP349 and FAP483). According to the average MAP features, all studied biallelic carriers were phenotypically similar to attenuated adenomatous polyposis coli-polyposis patients and characterized by a limited number of colorectal polyps (range 30–150), with the first diagnosis in adult ages (range 37–51 years). The pathology records of four patients (FAP236, FAP349, FAP483 and FAP527) reported, however, co-existing adenomatous and hyperplastic polyps. With the exception of one patient (FAP117), in which adenomas had been detected in small intestine, no apparent extracolonic disease manifestations were observed in the other biallelic mutation carriers.
The position of the variants in relation to the MUTYH coding region together with the estimated locations of the assumed functional domains of the MUTYH protein14 are shown in Figures 1a and b. The c.933+3A>C (IVS10+3A>C), a transversion causing an aberrant splicing process leading to skipping of exon 10 with the formation of a truncated protein (p.Gly264TrpfsX7)19 and the predicted truncated gene products of the p.Glu410GlyfsX43 and p.Ala385ProfsX23 mutations are shown in Figure 1b.
Localization along the MUTYH gene of the variants analyzed in this study. (a) Localization of missense (p.Tyr179Cys, p.Gly396Asp, p.Arg245Cys and p.Arg245His) and small in-frame deletion (p.Glu480del) mutations in the MUTYH gene with respect of the putative functional domains of the repair protein. (b) Graphic representation of the predicted truncated gene products of p.Gly264TrpfsX7, p.Ala385ProfsX23 and p.Glu410GlyfsX43 mutations.
Expression of MUTYH in these variants was analyzed by real-time PCR using specific primers and a probe located at the junction between exons 5 and 6 of the gene. This allows detecting both mitochondrial and nuclear isoforms of MUTYH. Data from three independent determinations are shown in Figure 2. The comparison of MUTYH expression levels, normalized using the mean of two housekeeping genes (β-actin and β-tubulin), between seven LCLs from MAP patients and three LCLs from healthy donors indicate that only in one instance MUTYH levels were similar to wild-type cells (p.Tyr179Cys/Gly396Asp; Figure 2a). Thus, a 20–35% decrease in MUTYH transcripts was observed in four LCLs harboring the p.Arg245Cys/Gly396Asp, p.Gly264TrpfsX7/Glu480del, p.Arg245His/Arg245His and p.Gly264TrpfsX7/Gly264TrpfsX7 variants, and these levels were further decreased to 50% of the wild-type protein in LCLs carrying the p.Gly264TrpfsX7/Ala385ProfsX23 and p.Gly396Asp/Glu410GlyfsX43 variants (Figures 2a and c).
Quantitative analysis of MUTYH in wild-type and MUTYH variant LCLs. (a) Total amount of MUTYH mRNA (nuclear and mitochondrial forms) as obtained from a MUTYH mRNA analysis performed by real-time PCR on three different wild-type and variant LCLs. MUTYH signal was normalized by quantization with two different housekeeping genes (β-actin and β-tubulin) as described in Materials and methods section. Data are the mean±s.d. of three independent determinations. ***P<0.001 (Student’s t-test). (b) A representative western blot of the MUTYH protein in whole-cell extracts is also shown. Blots were probed for MUTYH using an antibody specific for a C-terminal epitope (aa 435–535) and for β-tubulin protein. Relative expression of MUTYH protein in whole-cell extracts from MUTYH variants vs the wild-type BR806 cell line. A second wild-type cell line (BR805) is also shown for comparison. Data are the mean±s.d. from western blotting analyses performed in three independent experiments. Lanes 1 to 9 correspond to the LCLs indicated in panel c. (c) Comparison of relative levels of MUTYH transcript and protein in the MUTYH variant LCLs. Reference values for transcripts are the mean of MUTYH expression in the wild-type BR806, BR805 and BR77 LCLs. Lanes 1 to 9 correspond to data presented in panel b.
Protein levels were then measured by western blotting (Figure 2b). Reproducible levels of the MUTYH protein were observed in three independent experiments and a comparison with two wild-type LCLs (lanes 5 and 6) from healthy donors confirms that in some cell lines (FAP236, FAP182 and FAP527) there is a good correlation between transcripts and protein levels (lanes 1, 2 and 8; Figure 2c). In other instances, often in LCLs harboring frameshift mutations, no MUTYH or traces of the full-length protein are detectable (lanes 4 and 9, respectively). It is possible that these truncated proteins might not be identifiable by our antibody that recognizes an epitope at the C-terminus of the protein (aa 435–535). Finally, a low level of the MUTYH protein is also observed in FAP527 LCLs with the homozygous Arg245His mutation (lane 7) suggesting that this specific mutation confers instability to the MUTYH protein.
DNA glycosylase activity in LCLs from MAP patients
MUTYH DNA glycosylase activity was assayed by measuring cleavage of 30-mer oligonucleotides containing a single 8-oxodG:A mispair. Extracts of two independent wild-type LCLs, BR806 and BR77 were proficient in A excision (lanes 3–5; Figures 3a and b). Although MUTYH activity activity is low, the assay reproducibly measured reductions of ⩾50% (examples are shown in Supplementary Figure S1). MUTYH DNA glycosylase activity was undetectable in extracts of MAP LCLs (Figures 4a and b). In the same extract, a dose response for the OGG1-dependent removal of 8-oxodG from a duplex with an 8-oxoG:C mispair was also identified (Supplementary Figures S2a and b). The activity of OGG1 was similar in MAP and wild-type extracts (examples are shown in Supplementary Figure S2c). These data confirm the substantially reduced MUTYH activity in MAP LCLs. They indicate further that these cells retain normal levels of OGG1 DNA glycosylase.
Dose response of MUTYH DNA glycosylase activity in cell extracts from wild-type LCLs. (a) A representative gel of the reaction products because of MUTYH activity on an 8-oxoG:A substrate is shown. A 3′-end fluorescent labeled DNA substrate (150 fmol) was incubated with increasing amount of cell extracts at 37 °C for 1 h, treated with NaOH and analyzed by 20% denaturing polyacrylamide gel electrophoresis (PAGE) in 1 × TBE at 500 V for 2 h 30 min. The product of the DNA glycosylase activity is a 16-mer fragment (indicated by an arrow) that co-migrates with the reaction product of the purified MUTYH protein. Control DNA (lane 1); reaction product of a purified MUTYH protein (lane 2) and following incubation with 30, 60 and 90 μg of cell extracts from BR77 LCL (lanes 3–5). (b) Fluorescent band intensities from triplicate gels (as in a) were quantified using the public domain NIH ImageJ software and data analysis was performed with Kaleidagraph software (Synergy Software, Reading, PA, USA). The percentage of reaction product following incubation with increasing amount of cell extracts from the wild-type BR77 cells (empty circle) are shown together with values obtained in another wild-type cell line (BR806, full triangle).
MUTYH DNA glycosylase activity in cell extracts from LCLs derived from different MAP patients. (a) Representative gels of MUTYH activity in different LCLs. 6-Carboxyfluorescein (6-FAM)-labeled 8-oxoG:A substrate was reacted with 90 μg of cell extracts at 37 °C for 1 h and treated with NaOH. Positive control with MUTYH purified protein (6 ng, lanes 1, 12; 13 ng, lane 8); control DNA (lanes 2, 7); reaction products following incubation with wild-type BR77 (lane 13), p.Tyr179Cys/Gly396Asp (lane 3), p.ArgHis/Arg245His (lane 4), p.Gly264TrpfsX7/Ala385ProfsX23 (lane 5), p.Gly264TrpfsX7/Gly264TrpfsX7 (lane 6), p.Gly264TrpfsX7/Glu480del (lane 9), pArg245Cys/Gly396Asp (lane 10) and pGly396Asp/Glu410GlyfsX43 (lane 11). (b) The percentage of product obtained after reaction of the 8-oxoG:A substrate with LCL cell extracts as described in a is shown.
Steady-state levels and kinetics of repair of DNA 8-oxodG
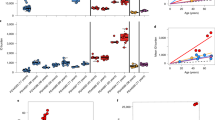
Inactivation of the mouse MUTYH gene is associated with an accumulation of DNA 8-oxodG observed in MEFs as well as in vivo.18, 20 To examine the effects of the MUTYH mutations in human cells, we compared steady-state levels of DNA 8-oxodG in MAP LCLs and the two control LCLs. Basal levels of DNA 8-oxodG in the BR77 and BR806 wild-type cell lines were 0.44 and 0.49 residues × 10−6 dG, respectively (Figure 5a). Expression of variant MUTYH was associated with increases of 1.3- up to 1.7-fold in DNA 8-oxodG levels (P⩽0.02–0.001; Figure 5a). The two exceptions were cells harboring the p.Gly264TrpfsX7 variant either in homozygosity or in combination with the frameshift mutation p.Ala385ProfsX23. These cell lines, in which no detectable expression or traces of the MUTYH protein could be identified, showed levels of DNA 8-oxodG similar to wild-type cells.
DNA 8-oxodG levels in wild-type and MUTYH variant LCLs. (a) Steady-state DNA 8-oxodG levels were measured in wild-type BR77 and BR806 LCLs by high-performance liquid chromatography with electrochemical detection (HPLC/EC). Data are mean±s.e. from 6 to 11 independent measurements. **P<0.01, ***P<0.001 (Student’s t-test against wild-type cultures). (b) Repair kinetics of DNA 8-oxodG following exposure to a 30-min treatment with 20 mM KBrO3 in wild-type and MUTYH variant LCLs. Rate of DNA 8-oxodG removal was measured at the indicated time points. Data are mean±s.e. from five experiments.
To examine whether exposure to an oxidative stress might exacerbate the biological consequences of a MUTYH defect in p.Gly264TrpfsX7/Ala385ProfsX23 LCL, this cell line was compared with wild-type BR77 cells following treatment with the KBrO3 oxidant (Figure 5b). The defective processing of 8-oxodG:A mispairs resulted in 8-oxodG repair kinetics much slower than those observed in the wild-type cells, with almost a doubling in the half-life of the lesion (360 vs 180 min). This phenotype is similar to that observed in two other MUTYH defective LCLs expressing 20% or 50% of the MUTYH protein (p.Gly264TrpfsX7/Glu480del and p.Gly396Asp/Glu410GlyfsX43, respectively; Figure 5b). Thus, independently from the level of expression of the mutant MUTYH protein, removal of 8-oxodG from the genome was clearly impaired in LCLs derived from MAP patients.
Our findings indicate that MUTYH defects cause an increased steady-state level of DNA 8-oxodG. The differential 8-oxoG accumulation is more marked under conditions of oxidative stress. Expression of significant levels of mutated protein can have a more profound effect that simple loss of MUTYH expression suggesting possible dominant-negative effects.
Measurements of mutation frequencies at the PIG-A gene
To investigate whether the defective MUTYH activity in LCLs from MAP patients is associated with increased mutagenesis, we compared the mutation frequency at the X-linked PIG-A locus. Mutations in this gene result in either complete or partial deficiency of membrane glycosylphosphatidylinositol (GPI)-linked proteins and CD48, CD59 and CD55-negative mutant cells can be identified by fluorescence-activated cell sorting analysis21, 22 (Figure 6a). Measurements of mutation frequency in three wild-type cell lines (BR77, BR805 and BR806) yielded a mean value of 21.7 × 10−6 GPI-negative events (10.9, 31.7 and 22.5 × 10−6, respectively) with modest variations among different determinations (Figure 6b). This value is in the range of values previously reported for wild-type human cell lines.21, 22 The mutation frequencies in MAP LCLs were higher with a mean value of 87.6 × 1016 GPI-negative events, ranging from 42.2 (the homozygous p.Gly264TrpfsX7) to 157 × 10−6 GPI-negative events (p.Arg245His/Arg245His) (Student’s t-test, P=0.04; Figure 6b).
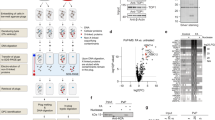
Mutation frequency at the PIG-A gene. (a) Flow cytometry methodology used to establish the population to be analyzed for the measurement of mutation frequency. Dead cells were excluded by propidium iodide staining (PI/FSC-A), whereas cell doublets were discriminated and excluded by comparing Area and Height signals of Forward scatter parameters (FSC-H/FSC-A; first two panels). The fluorescein isothiocyanate (FITC)-human leukocyte antigen (HLA)-DR positive population was gated (third panel) and evaluated in the phycoerythrin (PE) channel (last panel), in order to obtain a correct compensation of FITC emission and to establish at the same time the gating necessary to determine the distribution of mutant cells in the PE channel. (b) Representative flow cytometry dot plot analysis of WTBR77 (first panel) and p.Gly396Asp/Glu410GlyfsX43 (second panel) LCLs stained with antibodies against CD48, CD55, CD59 (all GPI-anchored proteins) and HLA-DR (non GPI-anchored protein) antigens. Mutant cells, represented by GPI-negative events (lower quadrant of panels) do not express CD48, CD55, CD59 but do express HLA-DR. Spontaneous mutation frequencies, calculated as the fraction of GPI-negative events in HLA-DR positive cells (third panel) measured at 1-week intervals in wild-type (BR77, BR805 and BR806) and mutant MUTYH LCLs. Each bar represents the mean of 4–11 independent values. (c) Cytometry dot plots of p.Gly396Asp/Glu410GlyfsX43 cells before (left) and after (center) sorting for GPI-positive population. Re-analysis of the same sorted population after several days in culture (right) shows the reappearance of GPI-negative cells.
In order to exclude that this high mutational load was due to a genetic drift of the cell population, GPI-negative cells were eliminated by flow sorting in the p.Gly396Asp/Glu410GlyfsX43 cell line. This sorted cell population was then expanded in vitro for several days and re-analyzed for mutation frequency at the PIG-A gene (Figure 6c). Once the initial GPI-negative events had been cleared, mutation frequency measured after several days in culture raised up to 149 mutation × 10−6 cells. Thus, fluorescence-activated cell sorting analysis of this cell population revealed the reappearance of the GPI-negative cells, demonstrating the tendency of this MUTYH-defective cell line to accumulate mutations in the absence of any mutagenic stress. We can conclude that a spontaneous mutator phenotype is a characteristic feature of cells with a defective MUTYH activity.
We have previously shown that MUTYH-defective variants are hypersensitive to killing by KBrO3 exposure.18 To investigate whether this sensitivity was associated with enhanced mutability, we compared mutation frequencies in KBrO3-treated wild-type BR77 and p.Gly396Asp/Glu410GlyfsX43 LCLs (Figure 7). These MUTYH-defective cells were only slightly more sensitive than BR77 cells to killing by KBrO3 (Figure 7a). The number of mutations introduced into the genome by KBrO3 exposure was, however, much larger in p.Gly396Asp/Glu410GlyfsX43 than in BR77 cells. As a linear increase in mutation frequency was observed as a function of dose in both cell lines (Figures 7b and c) a comparison of the respective slopes (y=58.5x+382 vs y=14x−5, respectively) showed an increment of 4.2-fold in the mutagenic response of the MUTYH-defective cell line (Figure 7d). These data indicate that defective MUTYH confers a hypermutable phenotype under conditions of oxidative stress.
Mutations induced at the PIG-A gene in the wild-type BR77 cells and the MUTYH-defective p.Gly396Asp/Glu410GlyfsX43 cells following exposure to KBrO3. (a) Survival measured in wild-type BR77 cells (black bar) and p.Gly396Asp/Glu410GlyfsX43 cells (white bar) 7 days after a 30-min exposure to increasing concentrations of KBrO3. (b, c) Mutation frequency measured 7 days after a 30-min treatment with increasing concentrations of KBrO3 in wild-type BR77 (b) and p.Gly396Asp/Glu410GlyfsX43 (c) cells. Values are the mean and s.d. of two independent experiments. (d) Comparison of the dose response curves for mutation induction by KBrO3 exposure in BR77 and p.Gly396Asp/Glu410GlyfsX43 LCLs.
Discussion
MAP is an increasingly studied autosomal recessive disorder. Analysis of the effects of MUTYH mutations is more limited and mainly confined to the most common missense variants. In this study, we characterized several LCLs from MAP patients carrying both compound heterozygous and homozygous mutations. Analysis of protein expression in these MAP LCLs indicates that MUTYH transcript levels are not always informative. Indeed the substitution of Arg 245 with His, while modestly affecting the level of transcript, is associated with a significantly reduced protein level. It seems likely that this mutation, which is localized near the 4Fe–4S cluster domain affects protein stability.
When LCLs from MAP patients were analyzed for their glycosylase activity, an impairment in cleavage of an 8-oxodG:A-containing DNA substrate was identified in all cases. In published reports, with a few exceptions,23, 24 purified or partially purified MUTYH variants were used to investigate DNA glycosylase activity.25, 26, 27, 28, 29, 30, 31 The p.Tyr179Cys and p.Arg245His substitutions result in the complete inactivation of MUTYH glycosylase activity,28, 29, 30, 31 whereas the enzymatic defect conferred by the p.Gly396Asp substitution causes a less severe activity reduction.27, 28, 30 Our data in LCLs expressing the p.Tyr179Cys/Gly396Asp (FAP527) and p.Arg245His/Arg245His (FAP483) variants indicate that these modifications are associated with a complete loss of MUTYH enzymatic activity in vivo. In contrast, although the p.Arg245Cys is present in the LOVD variant listing14 no biochemical data using the purified variant protein are available. By analogy to the His and Leu modifications at this position, which have been demonstrated to inactivate the enzymatic function,27, 31 structural analysis suggests the pathogenicity of this substitution. The Arginine 245 residue is strictly conserved and is placed in an α-helix perpendicular to the HhH motif. Besides that, because of the close proximity to the iron–sulfur cluster, the presence of an additional cysteine might cause a local rearrangement. The absence of DNA glycosylase activity in FAP182 LCLs where this variant is present in combination with p.Gly396Asp, supports the pathogenetic role of this mutation in MAP. Finally, the frameshift mutations present in FAP349, FAP278 and FAP236 (p.Gly264TrpfsX7, p.Ala385ProfsX23 and p.Glu410GlyfsX43) resulted in severely truncated proteins with loss of 278, 141 and 97 amino acids, respectively. As the deleted regions contain the nucleoside diphosphate linked to moiety X (NUDIX) domain as well as the proliferating cell nuclear antigen-binding region, the absence of MUTYH glycosylase activity in these cells, possibly because of defective binding, is hardly surprising.
We previously demonstrated in rodent cell lines that Mutyh loss is associated with increased steady-state levels of DNA 8-oxodG and expression of some human MUTYH variants leads to a more profound effect in the removal of the oxidized purine than the simple protein loss.18 This phenotype can also be observed in human cells with defective MUTYH DNA glycosylase activity and an accumulation of DNA 8-oxodG was observed following an oxidative stress induced by KBrO3. Even in the absence of exogenous oxidation, the majority of LCLs harboring MUTYH variants showed increased steady-state levels of DNA 8-oxodG when compared with wild-type cells. Interestingly, LCLs harboring frameshift mutations with no detectable expression of the MUTYH protein (the p.Gly264TrpfsX7 and p.Ala385ProfsX23) show a more moderate phenotype. We suggest that the presence of a mutant MUTYH protein might have a dominant-negative effect, possibly interfering with others DNA repair pathways involved in the removal of 8-oxodG. These factors might include mismatch repair or other base excision repair proteins, nucleotide excision repair, as well as components of the DNA damage response (for example, the RAD9-RAD1-HUS1 complex).32
At variance with the mouse model where mutant MUTYH proteins are expressed in the same genetic background (Mutyh−/− MEFs), LCLs from different patients might have an ample range of cellular responses to oxidative stress. This is one of the reasons why both MUTYH and OGG1 activity were measured in parallel in cell-free extracts. Indeed the role of MUTYH in removal of 8-oxodG from the genome is indirect and several lines of evidence indicate that the OGG1 DNA glycosylase is one of the major contributors to the repair of this oxidized purine. The observation that all LCLs show wild-type OGG1 activity allows concluding that the accumulation of DNA 8-oxodG is a direct consequence of the MUTYH defect.
Expression of MUTYH variants in Mutyh−/− MEFs conferred a clear hypersensitivity to killing by KBrO3 (but not by γ-irradiation).18 This phenotype was much more moderate in the case of human LCLs, as shown by data obtained in the p.Gly396Asp/Glu410GlyfsX43 variant and confirmed in the p.Gly264TrpfsX7/Ala385ProfsX23 (data not shown). It is possible that establishment of LCLs by Epstein–Barr virus infection might alter the phenotype associated with MUTYH deficiency. Indeed it has been shown that the EBNA1 protein destabilizes the p53 protein, and decreased level of p53 may increase resistance to apoptosis under oxidative stress.33
The information on the mutator phenotype associated with MUTYH inactivation is limited to MEFs and embryonic stem cells from Mutyh−/− mice, in which increases in mutation rates varied from 2- to 10-fold.20, 34 The presence of specific G>T transversions in adenomatous polyposis coli or KRAS genes in tumors occurring in MAP patients can also be considered an indirect evidence of a mutator phenotype associated with the presence of a non-functional MUTYH.5, 6, 35, 36 Here we investigated for the first time to which extent MUTYH impairment results in an increased mutation frequency in human cells. The gene we chose for this analysis is the X-linked PIG-A gene that has been successfully used to characterize the mutator phenotype associated with DNA repair-defective human syndromes (Ataxia telangiectasia, Fanconi anemia).21 When compared with the 21.7 × 10−6 mutational events at the PIG-A gene observed in wild-type cells, the group of LCLs from MAP patients showed a fourfold increased mutation frequency. The reappearance of PIG-A mutants in a cell line with one of the highest mutant frequency (p.Gly396Asp/Glu410GlyfsX43), following flow sorting to eliminate the pre-existing mutant cells, demonstrates that this phenotype is not due to a genetic drift of the cell population but it is an intrinsic feature of these cells. The reported spontaneous mutation rate (m) at this gene for normal cells ranges from 8.8 to 16.5 × 10−7 mutation per cell division.22 A tentative m-value of 299 × 10−7 mutation per cell division was calculated on the p.Gly396Asp/Glu410GlyfsX43 LCLs following measurement of mutation frequency after six doublings. This value is at least 10-times higher that m-values for normal human cells.22 These data support the presence of a spontaneous mutator phenotype associated with defective MUTYH proteins and fit quite well with the proposed role of MUTYH variants in favoring the appearance of mutations in oncogenes relevant in the process of colorectal carcinogenesis.5, 6
As oxidative damage in DNA can contribute to genome instability, we also investigated whether MUTYH inactivation exposes cells to a further increase in their mutational burden in a condition of oxidative stress. Thus, treatment of the biallelic p.Gly396Asp/Glu410GlyfsX43 variant LCLs with an oxidizing agent results indeed in a hypermutable phenotype. Oxidative stress might then have an independent and major role in causing genome instability in the presence of an inactivating MUTYH mutation. Although the mutator phenotype associated with MUTYH defects is considerably milder than that observed in mismatch repair-defective cells, the hypermutability induced by specific genotoxic agents is a shared characteristic of cells with inactivation in either pathway.37, 38, 39, 40 Thus, we suggest that the level of oxidative damage to DNA bases undergoing in specific districts of the body (and specifically in the gastrointestinal tract) might be a relevant issue to explain the tissue specificity of the increased cancer susceptibility of MUTYH- and mismatch repair-defective patients.
Taken together, these findings support the pathogenic role of the MUTYH mutations in the clinical histories of the MAP patients we analyzed. In addition, the results obtained by these novel functional assays identify the mutator phenotype and oxidative DNA damage as relevant factors, which might contribute to establish informative genotype–phenotype relationships.
Materials and methods
Sample collection and LCL establishment
Lymphocytes of patients and controls were collected from 20 ml of peripheral blood using Lymphoprep solution (Axis-Shield, Dundee, Scotland) according to the manufacturer’s instruction and resuspended in 90% fetal bovine serum, 10% dimethylsulphoxide, frozen at −80 °C and then stored in liquid nitrogen. MUTYH genetic testing was carried out on genomic DNA extracted from lymphocytes. Direct DNA sequencing was carried out on PCR products of exons 1–16 and adjacent intronic regions5 using the BigDye Terminator v3.1 kit and an ABI3100 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA). To establish LCLs, lymphocytes were thawed rapidly at 37 °C and immortalized via Epstein–Barr virus infection in the presence of cyclosporin A. Immortalization was recognized by the appearance of floating clumps of cells. LCL were grown in RPMI supplemented with 15% fetal bovine serum and 1% penicillin–streptomycin at 37 °C and 5% CO2.
MUTYH mRNA analysis
Lymphoblastoid cells (2–3 × 106) from each MAP patient carrying biallelic MUTYH mutations (FAP117, FAP182, FAP236, FAP278, FAP349, FAP483 and FAP527; Table 1) and three controls without MUTYH mutation were pelleted and RNA extracted by EZ1 QIAgen RNA extraction Mini-kit (Qiagen, Germantown, MD, USA). Complementary DNAs were prepared using a Reverse Transcription System kit (Promega, Madison, WI, USA) starting from 1 μg of total RNA. For the quantitative analysis of the MUTYH transcripts, real-time PCR was performed on complementary DNA using the TaqMan Gene Expression Master Mix (Applied Biosystems) and a CFX96 thermal cycler (Bio-Rad, Hercules, CA, USA). Total amount of MUTYH mRNA (nuclear and mitochondrial forms, wild-type and MUT) was detected with a predeveloped TaqMan Gene Expression Assay (Applied Biosystems) containing a FAM-MGB probe directed at the junction between exons 5 and 6. MUTYH signal was normalized by quantization of two different housekeeping genes (β-actin and β-tubulin) with predeveloped assays containing a VIC-MGB probe (Applied Biosystems), allowing to work in duplex condition. To avoid possible contamination with genomic DNA, RNA samples were treated with DNAse during RNA extraction and TaqMan probes sited in exons’ junctions were used. Quantitative evaluation of MUTYH expression was carried out by comparing mutant and wild-type samples by the ΔΔCt method.
Western blotting
Following centrifugation, lymphoblastoid cells (5 × 106) were washed once with phosphate-buffered saline, resuspended in lysis buffer (20 mM Tris-HCl, 100 mM NaCl, 5 mM MgCl2, 0.2 mM EDTA, 0.1% NP40) containing protease and phosphatase inhibitors (Sigma, St Louis, MO, USA) and incubated on ice for 30 min. Lysed cells were then centrifuged in a microfuge and protein concentration was determined by Bradford assay (Bio-Rad). Proteins (100–150 μg) were loaded on 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis precast gels, transferred to nitrocellulose membrane (Protran, Schleicher & Schuell, Dassel, Germany) and incubated overnight with a mouse monoclonal antibody (dilution 1:150; Ab 55551; Abcam, Cambridge, UK) specific for a C-terminal protein epitope. The signal was amplified by an horseradish peroxidase-conjugated secondary antibody (dilution 1:2000, NA931, GE Healthcare, formerly Amersham Biosciences, Piscataway, NJ, USA) and revealed by Enhanced ChemiLuminescence (GE Healthcare) following the suggested protocol. MUTYH signals were normalized to β-tubulin protein.
Determination of 8-oxodG
8-OxodG was measured by high-performance liquid chromatography with electrochemical detection as described in ref. 41. To determine 8-oxodG repair kinetics, cells were treated for 30 min with 20 mM KBrO3 and at each time point, 1 × 106 cells were removed from dishes, DNA was extracted, and 8-oxodG levels measured as described above.
Preparation of cell extracts
LCLs pellets (4 × 107 cells) were resuspended in 200 ul of lysis buffer (50 mM Tris–HCl, pH 7.5, 200 mM NaCl, 1 mM EDTA, 1 mM phenylmethylsulfonyl fluoride, 1 × protease inhibitor cocktail containing 25 μg/ml pestatin A, 50 μg/ml leupeptin and 0.2% aprotinin (Roche Diagnostic S.p.A, Monza, Italy) and 5% glycerol. Cells were frozen at −80 °C for 2 h and quickly thawed at 30 °C. Samples were then sonicated on ice four times for 10 s each. Cells were cleared by centrifugation at 14 000 r.p.m. for 30 min at 4 °C and the supernatant saved. Protein concentrations were quantified with Bradford reagent (Bio-Rad) and cell extracts aliquoted and stored at −80 °C.
DNA glycosylase assay
The 8-oxoG:A substrate was a 30-mer double-stranded oligonucleotide containing a single 8-oxodG modification at position 21 in the sequence 5′-CCTCTGTGTGCTCAAGGGGGoxoGCTATAAGTTCTTTGC-3′, annealed with the complementary strand, 3′ end labeled with 6-carboxyfluorescein, containing an A at position 16. The 8-oxoG:C substrate differed for the presence of an 8-oxoG:C mispair and was 3′ end labeled with 6-carboxyfluorescein at the oxidized strand. These high-performance liquid chromatography purified oligonucleotides were purchased from ThermoFisher Scientific Inc., GmbH., Ulm, Germany.
Dose response base excision repair reactions were performed by incubating the 30-bp duplex (150 fmol) with increasing amount of LCL cell extracts (30, 60 and 90 μg) in 10 μl of 20 mM Tris–HCl, pH 8, 100 mM NaCl, 1 mM EDTA, 1 mM dithiothreitol, 0.1 mg/ml bovine serum albumin buffer at 37 °C for 1 h. The comparison of the DNA glycosylase activity of wild-type and variant MUTYH proteins present in cell extracts was obtained at 90 μg, as measured by Bradford assay. Following the DNA glycosylase reaction, the samples were incubated with NaOH (80 mM) at 90 °C for 4 min to hydrolyze uncleaved abasic sites produced by MUTYH or OGG1. After the addition of a loading dye solution, samples were heated at 90 °C for 2 min. Reaction products were analyzed by 20% denaturing polyacrylamide gel electrophoresis in 1 × Tris-Borate-EDTA buffer at 500 V for 2 h and 30 min. Fluorescent bands, visualized by Typhoon 9200 Gel Imager (GE Healthcare), were quantified by using the public domain NIH ImageJ software (http://rsb.info.nih.gov/ij/).
Determination of mutations at the PIG-A gene
LCLs are stained on ice with a mixture of phycoerythrin-mouse anti-human antibodies specific for three GPI-linked proteins, anti-CD48 (BD Pharmingen, BD Biosciences, Franklin Lakes, NJ, USA), anti-CD55 (BD Pharmingen), and anti-CD59 (AbD Serotec, Oxford, UK), and with the fluorescein isothiocyanate mouse anti-human leukocyte antigen-DR, (BD Pharmingen), against a non-GPI-anchored transmembrane protein. Mutant cells (GPI-negative) are detected as human leukocyte antigen-DR positive but CD48, CD55 and CD59-negative events. Dead cells were excluded by propidium iodide staining (PI/FSC-A), whereas cells doublets were discriminated and excluded by comparing Area and Height signals of Forward scatter parameters (FSC-H/FSC-A). The mutant frequency (f) is calculated as the number of GPI-negative cells divided by the number of total events analyzed. To calculate the mutation rate, preexisting mutants are eliminated by flow sorting on a FACS Aria (BD Biosciences) from the population by collecting the upper 50th percentile of the distribution curve after staining with anti-CD59. Live cells were identified by light scatter characteristics and doublets excluded by pulse width.31 The collected GPI-positive cells are counted and the new culture re-started with at least 1 × 106 cells in order to minimize fluctuation effects. After expansion (5–7 cell divisions), the cells are again analyzed by flow cytometry using the mixture of all the GPI-linked proteins in order to avoid classifying cells as PIG-A mutant falsely. The mutation rate (μ) is calculated using the formula μ=f÷d, where d represents the number of cell divisions occurring in vitro after sorting.22
Statistical analysis
The effect of each mutation on MUTYH expression and 8-oxodG levels was contrasted, as for its statistical significance, with data relative to the wild-type BR77 and BR806 cultures. The control sample was made of different independent experiments performed in the two wild-type groups, whereas the ‘treatment’ sample was each time a different mutation. The used statistical test was Student’s t-test because the independence of each mutation prevented us to use an analysis of variance+post hoc testing that is a strongly context-dependent paradigm and must be based on a consistent source of variation. The analysis of PIG-A mutant frequency was performed by Student’s t-test having as groups the wild-type cultures contrasted with mutant LCLs considered as a whole.
References
David SS, O’Shea VL, Kundu S . Base-excision repair of oxidative DNA damage. Nature 2007; 447: 941–950.
van Loon B, Markkanen E, Hübscher U . Oxygen as a friend and enemy: How to combat the mutational potential of 8-oxo-guanine. DNA Repair (Amst) 2010; 9: 604–616.
Oka S, Nakabeppu Y . DNA glycosylase encoded by MUTYH functions as a molecular switch for programmed cell death under oxidative stress to suppress tumorigenesis. Cancer Sci 2011; 102: 677–682.
Slupska MM, Luther WM, Chiang JH, Yang H, Miller JH . Functional expression of hMYH, a human homolog of the Escherichia coli MutY protein. J Bacteriol 1999; 181: 6210–6213.
Al-Tassan N, Chmiel NH, Maynard J, Fleming N, Livingston AL, Williams GT et al. Inherited variants of MYH associated with somatic G:C-->T:A mutations in colorectal tumors. Nat Genet 2002; 30: 227–232.
Jones S, Emmerson P, Maynard J, Best JM, Jordan S, Williams GT et al. Biallelic germline mutations in MYH predispose to multiple colorectal adenoma and somatic G:C-->T:A mutations. Hum Mol Genet 2002; 11: 2961–2967.
Sieber OM, Lipton L, Crabtree M, Heinimann K, Fidalgo P, Phillips RK et al. Multiple colorectal adenomas, classic adenomatous polyposis, and germ-line mutations in MYH. N Engl J Med 2003; 348: 791–799.
Poulsen ML, Bisgaard ML . MUTYH associated polyposis (MAP). Curr Genomics 2008; 9: 420–435.
Jasperson KW, Tuohy TM, Neklason DW, Burt RW . Hereditary and familial colon cancer. Gastroenterology 2010; 138: 2044–2058.
Wang L, Baudhuin LM, Boardman LA, Steenblock KJ, Petersen GM, Halling KC et al. MYH mutations in patients with attenuated and classic polyposis and with young-onset colorectal cancer without polyps. Gastroenterology 2004; 127: 9–16.
Farrington SM, Tenesa A, Barnetson R, Wiltshire A, Prendergast J, Porteous M et al. Germline susceptibility to colorectal cancer due to base-excision repair gene defects. Am J Hum Genet 2005; 77: 112–119.
Cardoso J, Molenaar L, de Menezes RX, van Leerdam M, Rosenberg C, Möslein G et al. Chromosomal instability in MYH- and APC-mutant adenomatous polyps. Cancer Res 2006; 66: 2514–2519.
Cleary SP, Cotterchio M, Jenkins MA, Kim H, Bristow R, Green R et al. Germline MutY human homologue mutations and colorectal cancer: a multisite case-control study. Gastroenterology 2009; 136: 1251–1260.
Out AA, Tops CM, Nielsen M, Weiss MM, van Minderhout IJ, Fokkema IF et al. Leiden open variation database of the MUTYH gene. Hum Mutat 2010; 31: 1205–1215.
Gismondi V, Meta M, Bonelli L, Radice P, Sala P, Bertario L et al. Prevalence of the Y165C, G382D and 1395delGGA germline mutations of the MYH gene in Italian patients with adenomatous polyposis coli and colorectal adenomas. Int J Cancer 2004; 109: 680–684.
Isidro G, Laranjeira F, Pires A, Leite J, Regateiro F, Castro e Sousa F et al. Germline MUTYH (MYH) mutations in Portuguese individuals with multiple colorectal adenomas. Hum Mutat 2004; 24: 353–354.
Cheadle JP, Sampson JR . MUTYH-associated polyposis--from defect in base excision repair to clinical genetic testing. DNA Repair (Amst) 2007; 6: 274–279.
Molatore S, Russo MT, D’Agostino VG, Barone F, Matsumoto Y, Albertini AM et al. MUTYH mutations associated with familial adenomatous polyposis: functional characterization by a mammalian cell-based assay. Hum Mutat 2010; 31: 159–166.
Pin E, Pastrello C, Tricarico R, Papi L, Quaia M, Fornasarig M et al. MUTYH c. 933+3A>C, associated with a severely impaired gene expression, is the first Italian founder mutation in MUTYH-associated polyposis. Int J Cancer (e-pub ahead of print 3 August 2012; doi:10.1002/ijc.27761).
Russo MT, De Luca G, Casorelli I, Degan P, Molatore S, Barone F et al. Role of MUTYH and MSH2 in the control of oxidative DNA damage, genetic instability, and tumorigenesis. Cancer Res 2009; 69: 4372–4379.
Araten DJ, Golde DW, Zhang RH, Thaler HT, Gargiulo L, Notaro R et al. A quantitative measurement of the human somatic mutation rate. Cancer Res 2005; 65: 8111–8117.
Peruzzi B, Araten DJ, Notaro R, Luzzatto L . The use of PIG-A as a sentinel gene for the study of the somatic mutation rate and of mutagenic agents in vivo. Mutat Res 2010; 705: 3–10.
Alhopuro P, Parker AR, Lehtonen R, Enholm S, Mecklin JP, Karhu A et al. A novel functionally deficient MYH variant in individuals with colorectal adenomatous polyposis. Hum Mutat 2005; 26: 393–402.
Parker AR, Sieber OM, Shi C, Hua L, Takao M, Tomlinson IP et al. Cells with pathogenic biallelic mutations in the human MUTYH gene are defective in DNA damage binding and repair. Carcinogenesis 2005; 26: 2010–2018.
Gu Y, Lu AL . Differential DNA recognition and glycosylase activity of the native human MutY homolog (hMYH) and recombinant hMYH expressed in bacteria. Nucleic Acids Res 2001; 29: 2666–2674.
Yanaru-Fujisawa R, Matsumoto T, Ushijima Y, Esaki M, Hirahashi M, Gushima M et al. Genomic and functional analyses of MUTYH in Japanese patients with adenomatous polyposis. Clin Genet 2008; 73: 545–553.
Ali M, Heja K, Cleary S, Cupples C, Galllinger S, Bristow R . Characterization of mutant MUTYH proteins associated with familial colorectal cancer. Gastroenterology 2008; 135: 499–507.
Kundu S, Brinkmeyer MK, Livingston AL, David SS . Adenine removal activity and bacterial complementation with the human MutY homologue (MUTYH) and Y165C, G382D, P391L and Q324R variants associated with colorectal cancer. DNA Repair (Amst) 2009; 8: 1400–1410.
Goto M, Shinmura K, Nakabeppu Y, Tao H, Yamada H, Tsuneyoshi T et al. Adenine DNA glycosylase activity of 14 human MutY homolog (MUTYH) variant proteins found in patients with colorectal polyposis and cancer. Hum Mutat 2010; 31: E1861–E1874.
D'Agostino VG, Minoprio A, Torreri P, Marinoni I, Bossa C, Petrucci TC et al. Functional analysis of MUTYH mutated proteins associated with familial adenomatous polyposis. DNA Repair (Amst) 2010; 9: 700–707.
Bai H, Grist S, Gardner J, Suthers G, Wilson TM, Lu AL . Functional characterization of human MutY homolog (hMYH) missense mutation (R231L) that is linked with hMYH-associated polyposis. Cancer Lett 2007; 250: 74–81.
Shi G, Chang DY, Cheng CC, Guan X, Venclovas C, Lu AL . Physical and functional interactions between MutY glycosylase homologue (MYH) and checkpoint proteins Rad9-Rad1-Hus1. Biochem J 2006; 400: 53–62.
Yi F, Saha A, Murakami M, Kumar P, Knight JS, Cai Q et al. Epstein-Barr virus nuclear antigen 3C targets p53 and modulates its transcriptional and apoptotic activities. Virology 2009; 388: 236–247.
Hirano S, Tominaga Y, Ichinoe A, Ushijima Y, Tsuchimoto D, Honda-Ohnishi Y et al. Mutator phenotype of MUTYH-null mouse embryonic stem cells. J Biol Chem 2003; 278: 38121–38124.
Lipton L, Halford SE, Johnson V, Novelli MR, Jones A, Cummings C et al. Carcinogenesis in MYH-associated polyposis follows a distinct genetic pathway. Cancer Res 2003; 63: 7595–7599.
van Puijenbroek M, Nielsen M, Tops CM, Halfwerk H, Vasen HF, Weiss MM et al. Identification of patients with (atypical) MUTYH-associated polyposis by KRAS2 c.34G>T prescreening followed by MUTYH hotspot analysis in formalin-fixed paraffin-embedded tissue. Clin Cancer Res 2008; 14: 139–142.
Glaab WE, Risinger JI, Umar A, Barrett JC, Kunkel TA, Tindall KR . Resistance to 6-thioguanine in mismatch repair-deficient human cancer cell lines correlates with an increase in induced mutations at the HPRT locus. Carcinogenesis 1998; 19: 1931–1937.
Xu XS, Narayanan L, Dunklee B, Liskay RM, Glazer PM . Hypermutability to ionizing radiation in mismatch repair-deficient, Pms2 knockout mice. Cancer Res 2001; 61: 3775–3780.
Sansom OJ, Bishop SM, Court H, Dudley S, Liskay RM, Clarke AR . Apoptosis and mutation in the murine small intestine: loss of Mlh1- and Pms2-dependent apoptosis leads to increased mutation in vivo. DNA Repair (Amst) 2003; 2: 1029–1039.
Bignami M, O'Driscoll M, Aquilina G, Karran P . Unmasking a killer: DNA O(6)-methylguanine and the cytotoxicity of methylating agents. Mutat Res 2000; 462: 71–82.
Chiera F, Meccia E, Degan P, Aquilina G, Pietraforte D, Minetti M et al. Overexpression of human NOX1 complex induces genome instability in mammalian cells. Free Radic Biol Med 2008; 44: 332–342.
Acknowledgements
The authors are grateful to Drs M Fornasarig and D Barana for patient recruitment. This work has been supported by grants to MB from NIH/ISS, AIRC and by Ministero della Salute Alleanza contro il Cancro to AV and MB.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Ruggieri, V., Pin, E., Russo, M. et al. Loss of MUTYH function in human cells leads to accumulation of oxidative damage and genetic instability. Oncogene 32, 4500–4508 (2013). https://doi.org/10.1038/onc.2012.479
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2012.479
Keywords
This article is cited by
-
8-Oxoguanine: from oxidative damage to epigenetic and epitranscriptional modification
Experimental & Molecular Medicine (2022)
-
MUTYH-associated tumor syndrome: The other face of MAP
Oncogene (2022)
-
Increased prevalence of Barrett’s esophagus in patients with MUTYH-associated polyposis (MAP)
Familial Cancer (2020)
-
Novel variant of unknown significance in MUTYH in a patient with MUTYH-associated polyposis: a case to reclassify
Clinical Journal of Gastroenterology (2018)
-
Type and frequency of MUTYH variants in Italian patients with suspected MAP: a retrospective multicenter study
Journal of Human Genetics (2017)