Abstract
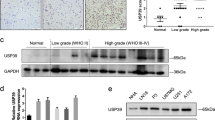
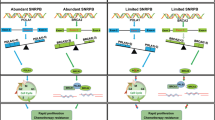
Serine/arginine-rich splicing factor 3 (SRSF3) likely has wide-ranging roles in gene expression and facilitation of tumor cell growth. SRSF3 knockdown induced G1 arrest and apoptosis in colon cancer cells (HCT116) in association with altered expression of 833 genes. Pathway analysis revealed ‘G1/S Checkpoint Regulation’ as the most highly enriched category in the affected genes. SRSF3 knockdown did not induce p53 or stimulate phosphorylation of p53 or histone H2A.X in wild-type HCT116 cells. Furthermore, the knockdown induced G1 arrest in p53-null HCT116 cells, suggesting that p53-dependent DNA damage responses did not mediate the G1 arrest. Real-time reverse transcription–polymerase chain reaction and western blotting confirmed that SRSF3 knockdown reduced mRNA and protein levels of cyclins (D1, D3 and E1), E2F1 and E2F7. The decreased expression of cyclin D and E2F1 likely impaired the G1-to-S-phase progression. Consequently, retinoblastoma protein remained hypophosphorylated in SRSF3 knockdown cells. The knockdown also induced apoptosis in association with reduction of BCL2 protein levels. We also found that SRSF3 knockdown facilitated skipping of 81 5′-nucleotides (27 amino acids) from exon 8 of homeodomain-interacting protein kinase-2 (HIPK2) and produced a HIPK2 Δe8 isoform. Full-length HIPK2 (HIPK2 FL) is constantly degraded through association with an E3 ubiquitin ligase (Siah-1), whereas HIPK2 Δe8, lacking the 27 amino acids, lost Siah-1-binding ability and became resistant to proteasome digestion. Interestingly, selective knockdown of HIPK2 FL induced apoptosis in various colon cancer cells expressing wild-type or mutated p53. Thus, these findings disclose an important role of SRSF3 in the regulation of the G1-to-S-phase progression and alternative splicing of HIPK2 in tumor growth.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Manley JL, Krainer AR . A rational nomenclature for serine/arginine-rich protein splicing factors (SR proteins). Genes Dev 2010; 24: 1073–1074.
Chen M, Manley JL . Mechanisms of alternative splicing regulation: insights from molecular and genomics approaches. Nat Rev Mol Cell Biol 2009; 10: 741–754.
Lin S, Coutinho-Mansfield G, Wang D, Pandit S, Fu X-D . The splicing factor SC35 has an active role in transcriptional elongation. Nat Struct Mol Biol 2008; 15: 819–826.
Lou H, Neugebauer KM, Gagel RF, Berget SM . Regulation of alternative polyadenylation by U1 snRNPs and SRp20. Mol Cell Biol 1998; 18: 4977–4985.
Sapra AK, Anko ML, Grishina I, Lorenz M, Pabis M, Poser I et al. SR protein family members display diverse activities in the formation of nascent and mature mRNPs in vivo. Mol Cell 2009; 34: 179–190.
Zhang Z, Krainer AR . Involvement of SR proteins in mRNA surveillance. Mol Cell 2004; 16: 597–607.
Sanford JR, Gray NK, Beckmann K, Caceres JF . A novel role for shuttling SR proteins in mRNA translation. Genes Dev 2004; 18: 755–768.
Anczuków O, Rosenberg AZ, Akerman M, Das S, Zhan L, Karni R et al. The splicing factor SRSF1 regulates apoptosis and proliferation to promote mammary epithelial cell transformation. Nat Struct Mol Biol 2012; 19: 220–228.
Ghigna C, Moroni M, Porta C, Riva S, Biamonti G . Altered expression of heterogeneous nuclear ribonucleoproteins and SR factors in human colon adenocarcinomas. Cancer Res 1998; 58: 5818–5824.
Ahn E-Y, DeKelver Russell C, Lo M-C, Nguyen Tuyet A, Matsuura S, Boyapati A et al. SON controls cell-cycle progression by coordinated regulation of RNA splicing. Mol Cell 2011; 42: 185–198.
Li X, Manley JL . Inactivation of the SR protein splicing factor ASF/SF2 results in genomic instability. Cell 2005; 122: 365–378.
Li X, Wang J, Manley JL . Loss of splicing factor ASF/SF2 induces G2 cell cycle arrest and apoptosis, but inhibits internucleosomal DNA fragmentation. Genes Dev 2005; 19: 2705–2714.
Xiao R, Sun Y, Ding J, Lin S, Rose D, Rosenfeld M et al. Splicing regulator SC35 is essential for genomic stability and cell proliferation during mammalian organogenesis. Mol Cell Biol 2007; 27: 5393–5402.
Moore MJ, Wang Q, Kennedy CJ, Silver PA . An alternative splicing network links cell-cycle control to apoptosis. Cell 2010; 142: 625–636.
Srebrow A, Kornblihtt A . The connection between splicing and cancer. J Cell Sci 2006; 119: 2635–2641.
He X, Ee P, Coon J, Beck W . Alternative splicing of the multidrug resistance protein 1/ATP binding cassette transporter subfamily gene in ovarian cancer creates functional splice variants and is associated with increased expression of the splicing factors PTB and SRp20. Clin Cancer Res 2004; 10: 4652–4660.
He X, Arslan AD, Pool MD, Ho TT, Darcy KM, Coon JS et al. Knockdown of splicing factor SRp20 causes apoptosis in ovarian cancer cells and its expression is associated with malignancy of epithelial ovarian cancer. Oncogene 2011; 30: 356–365.
Stickeler E, Kittrell F, Medina D, Berget SM . Stage-specific changes in SR splicing factors and alternative splicing in mammary tumorigenesis. Oncogene 1999; 18: 3574–3582.
Jumaa H, Wei G, Nielsen PJ . Blastocyst formation is blocked in mouse embryos lacking the splicing factor SRp20. Curr Biol 1999; 9: 899–902.
Änkö ML, Morales L, Henry I, Beyer A, Neugebauer KM . Global analysis reveals SRp20- and SRp75-specific mRNPs in cycling and neural cells. Nat Struct Mol Biol 2010; 17: 962–970.
Hailemariam K, Iwasaki K, Huang BW, Sakamoto K, Tsuji Y . Transcriptional regulation of ferritin and antioxidant genes by HIPK2 under genotoxic stress. J Cell Sci 2010; 123: 3863–3871.
Hofmann TG, Moller A, Sirma H, Zentgraf H, Taya Y, Droge W et al. Regulation of p53 activity by its interaction with homeodomain-interacting protein kinase-2. Nat Cell Biol 2002; 4: 1–10.
D’Orazi G, Cecchinelli B, Bruno T, Manni I, Higashimoto Y, Saito S et al. Homeodomain-interacting protein kinase-2 phosphorylates p53 at Ser 46 and mediates apoptosis. Nat Cell Biol 2002; 4: 11–19.
Wei G, Ku S, Ma GK, Saito S, Tang AA, Zhang J et al. HIPK2 represses β-catenin-mediated transcription, epidermal stem cell expansion, and skin tumorigenesis. Proc Natl Acad Sci USA 2007; 104: 13040–13045.
Winter M, Sombroek D, Dauth I, Moehlenbrink J, Scheuermann K, Crone J et al. Control of HIPK2 stability by ubiquitin ligase Siah-1 and checkpoint kinases ATM and ATR. Nat Cell Biol 2008; 10: 812–824.
Rinaldo C, Prodosmo A, Mancini F, Iacovelli S, Sacchi A, Moretti F et al. MDM2-regulated degradation of HIPK2 prevents p53Ser46 phosphorylation and DNA damage-induced apoptosis. Mol Cell 2007; 25: 739–750.
Choi DW, Seo YM, Kim EA, Sung KS, Ahn JW, Park SJ et al. Ubiquitination and degradation of homeodomain-interacting protein kinase 2 by WD40 repeat/SOCS box protein WSB-1. J Biol Chem 2008; 283: 4682–4689.
Calzado MA, de la Vega L, Moller A, Bowtell DD, Schmitz ML . An inducible autoregulatory loop between HIPK2 and Siah2 at the apex of the hypoxic response. Nat Cell Biol 2009; 11: 85–91.
Kim SY, Choi DW, Kim EA, Choi CY . Stabilization of HIPK2 by escape from proteasomal degradation mediated by the E3 ubiquitin ligase Siah1. Cancer Lett 2009; 279: 177–184.
Sombroek D, Hofmann TG . How cells switch HIPK2 on and off. Cell Death Differ 2009; 16: 187–194.
Puca R, Nardinocchi L, Givol D, D’Orazi G . Regulation of p53 activity by HIPK2: molecular mechanisms and therapeutical implications in human cancer cells. Oncogene 2010; 29: 4378–4387.
Zhong XY, Wang P, Han J, Rosenfeld MG, Fu XD . SR proteins in vertical integration of gene expression from transcription to RNA processing to translation. Mol Cell 2009; 35: 1–10.
Loomis RJ, Naoe Y, Parker JB, Savic V, Bozovsky MR, Macfarlan T et al. Chromatin binding of SRp20 and ASF/SF2 and dissociation from mitotic chromosomes is modulated by histone H3 serine 10 phosphorylation. Mol Cell 2009; 33: 450–461.
Schwerk C, Schulze-Osthoff K . Regulation of apoptosis by alternative pre-mRNA splicing. Mol Cell 2005; 19: 1–13.
Kim YH, Choi CY, Lee SJ, Conti MA, Kim Y . Homeodomain-interacting protein kinases, a novel family of co-repressors for homeodomain transcription factors. J Biol Chem 1998; 273: 25875–25879.
Giraud S, Diaz-Latoud C, Hacot S, Textoris J, Bourette RP, Diaz JJ . US11 of herpes simplex virus type 1 interacts with HIPK2 and antagonizes HIPK2-induced cell growth arrest. J Virol 2004; 78: 2984–2993.
Zhang Q, Yoshimatsu Y, Hildebrand J, Frisch SM, Goodman RH . Homeodomain interacting protein kinase 2 promotes apoptosis by downregulating the transcriptional corepressor CtBP. Cell 2003; 115: 177–186.
Hofmann TG, Stollberg N, Schmitz ML, Will H . HIPK2 regulates transforming growth factor-beta-induced c-Jun NH2-terminal kinase activation and apoptosis in human hepatoma cells. Cancer Res 2003; 63: 8271–8277.
Aikawa Y, Nguyen LA, Isono K, Takakura N, Tagata Y, Schmitz ML et al. Roles of HIPK1 and HIPK2 in AML1- and p300-dependent transcription, hematopoiesis and blood vessel formation. EMBO J 2006; 25: 3955–3965.
Hikasa H, Ezan J, Itoh K, Li X, Klymkowsky MW, Sokol SY . Regulation of TCF3 by Wnt-dependent phosphorylation during vertebrate axis specification. Dev Cell 2010; 19: 521–532.
Lazzari C, Prodosmo A, Siepi F, Rinaldo C, Galli F, Gentileschi M et al. HIPK2 phosphorylates ΔNp63α and promotes its degradation in response to DNA damage. Oncogene 2011; 30: 4802–4813.
D’Orazi G, Rinaldo C, Soddu S . Updates on HIPK2: a resourceful oncosuppressor for clearing cancer. J Exp Clin Cancer Res 2012; 31: 63.
Kuwano Y, Kamio Y, Kawai T, Katsuura S, Inada N, Takaki A et al. Autism-associated gene expression in peripheral leucocytes commonly observed between subjects with autism and healthy women having autistic children. PLoS One 2011; 6: e24723.
Kurokawa K, Kuwano Y, Tominaga K, Kawai T, Katsuura S, Yamagishi N et al. Brief naturalistic stress induces an alternative splice variant of SMG-1 lacking exon 63 in peripheral leukocytes. Neurosci Lett 2010; 484: 128–132.
Acknowledgements
This work was funded by the Grants-in-Aid for Scientific Research from Japan Society for the Promotion of Science (to KR).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Kurokawa, K., Akaike, Y., Masuda, K. et al. Downregulation of serine/arginine-rich splicing factor 3 induces G1 cell cycle arrest and apoptosis in colon cancer cells. Oncogene 33, 1407–1417 (2014). https://doi.org/10.1038/onc.2013.86
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2013.86
Keywords
This article is cited by
-
Panomics reveals patient individuality as the major driver of colorectal cancer progression
Journal of Translational Medicine (2023)
-
The involvement of E3 ubiquitin ligases in the development and progression of colorectal cancer
Cell Death Discovery (2023)
-
RNA splicing dysregulation and the hallmarks of cancer
Nature Reviews Cancer (2023)
-
Homeobox B9 Promotes Colon Cancer Progression by Targeting SRSF3
Digestive Diseases and Sciences (2023)
-
A novel SRSF3 inhibitor, SFI003, exerts anticancer activity against colorectal cancer by modulating the SRSF3/DHCR24/ROS axis
Cell Death Discovery (2022)