Abstract
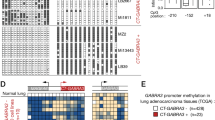
Aberrant expression of the DNA methyltransferases (DNMTs) and disruption of DNA methylation patterns are associated with carcinogenesis and cancer cell survival. The oncogenic MUC1-C protein is aberrantly overexpressed in diverse carcinomas; however, there is no known link between MUC1-C and DNA methylation. Our results demonstrate that MUC1-C induces the expression of DNMT1 and DNMT3b, but not DNMT3a, in breast and other carcinoma cell types. We show that MUC1-C occupies the DNMT1 and DNMT3b promoters in complexes with NF-κB p65 and drives DNMT1 and DNMT3b transcription. In this way, MUC1-C controls global DNA methylation as determined by analysis of LINE-1 repeat elements. The results further demonstrate that targeting MUC1-C downregulates DNA methylation of the CDH1 tumor suppressor gene in association with induction of E-cadherin expression. These findings provide compelling evidence that MUC1-C is of functional importance to induction of DNMT1 and DNMT3b and, in turn, changes in DNA methylation patterns in cancer cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Abbreviations
- DNMT:
-
DNA methyltransferase
- MUC1:
-
mucin 1
- EMT:
-
epithelial-mesenchymal transition
- ECAD:
-
E-cadherin
- LINE-1:
-
long interspersed nucleotide element-1.
References
Li E, Zhang Y . DNA methylation in mammals. Cold Spring Harb Perspect Biol 2014; 6: a019133.
Easwaran H, Tsai HC, Baylin SB . Cancer epigenetics: tumor heterogeneity, plasticity of stem-like states, and drug resistance. Mol Cell 2014; 54: 716–727.
Ahuja N, Easwaran H, Baylin SB . Harnessing the potential of epigenetic therapy to target solid tumors. J Clin Invest 2014; 124: 56–63.
Pradhan S, Bacolla A, Wells RD, Roberts RJ . Recombinant human DNA (cytosine-5) methyltransferase. I. Expression, purification, and comparison of de novo and maintenance methylation. J Biol Chem 1999; 274: 33002–33010.
Ko YG, Nishino K, Hattori N, Arai Y, Tanaka S, Shiota K . Stage-by-stage change in DNA methylation status of Dnmt1 locus during mouse early development. J Biol Chem 2005; 280: 9627–9634.
Ratnam S, Mertineit C, Ding F, Howell CY, Clarke HJ, Bestor TH et al. Dynamics of Dnmt1 methyltransferase expression and intracellular localization during oogenesis and preimplantation development. Dev Biol 2002; 245: 304–314.
Okano M, Bell DW, Haber DA, Li E . DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 1999; 99: 247–257.
Liang G, Chan MF, Tomigahara Y, Tsai YC, Gonzales FA, Li E et al. Cooperativity between DNA methyltransferases in the maintenance methylation of repetitive elements. Mol Cell Biol 2002; 22: 480–491.
Lin RK, Hsu HS, Chang JW, Chen CY, Chen JT, Wang YC . Alteration of DNA methyltransferases contributes to 5'CpG methylation and poor prognosis in lung cancer. Lung Cancer 2007; 55: 205–213.
Mizuno S, Chijiwa T, Okamura T, Akashi K, Fukumaki Y, Niho Y et al. Expression of DNA methyltransferases DNMT1, 3 A, and 3B in normal hematopoiesis and in acute and chronic myelogenous leukemia. Blood 2001; 97: 1172–1179.
Qu Y, Mu G, Wu Y, Dai X, Zhou F, Xu X et al. Overexpression of DNA methyltransferases 1, 3a, and 3b significantly correlates with retinoblastoma tumorigenesis. Am J Clin Pathol 2010; 134: 826–834.
Bakin AV, Curran T . Role of DNA 5-methylcytosine transferase in cell transformation by fos. Science 1999; 283: 387–390.
Bigey P, Ramchandani S, Theberge J, Araujo FD, Szyf M . Transcriptional regulation of the human DNA methyltransferase (dnmt1) gene. Gene 2000; 242: 407–418.
Jinawath A, Miyake S, Yanagisawa Y, Akiyama Y, Yuasa Y . Transcriptional regulation of the human DNA methyltransferase 3 A and 3B genes by Sp3 and Sp1 zinc finger proteins. Biochem J 2005; 385: 557–564.
Kishikawa S, Murata T, Kimura H, Shiota K, Yokoyama KK . Regulation of transcription of the Dnmt1 gene by Sp1 and Sp3 zinc finger proteins. Eur J Biochem 2002; 269: 2961–2970.
Lin RK, Wang YC . Dysregulated transcriptional and post-translational control of DNA methyltransferases in cancer. Cell Biosci 2014; 4: 46.
Rhee I, Jair KW, Yen RW, Lengauer C, Herman JG, Kinzler KW et al. CpG methylation is maintained in human cancer cells lacking DNMT1. Nature 2000; 404: 1003–1007.
Rhee I, Bachman KE, Park BH, Jair KW, Yen RW, Schuebel KE et al. DNMT1 and DNMT3b cooperate to silence genes in human cancer cells. Nature 2002; 416: 552–556.
Kufe D . Mucins in cancer: function, prognosis and therapy. Nat Rev Cancer 2009; 9: 874–885.
Kufe D . MUC1-C oncoprotein as a target in breast cancer: activation of signaling pathways and therapeutic approaches. Oncogene 2013; 32: 1073–1081.
Takahashi H, Jin C, Rajabi H, Pitroda S, Alam M, Ahmad R et al. MUC1-C activates the TAK1 inflammatory pathway in colon cancer. Oncogene 2015; 34: 5187–5197.
Ahmad R, Raina D, Trivedi V, Ren J, Rajabi H, Kharbanda S et al. MUC1 oncoprotein activates the IκB kinase β complex and constitutive NF-κB signaling. Nat Cell Biol 2007; 9: 1419–1427.
Ahmad R, Raina D, Joshi MD, Kawano T, Kharbanda S, Kufe D . MUC1-C oncoprotein functions as a direct activator of the NF-κB p65 transcription factor. Cancer Res 2009; 69: 7013–7021.
Rajabi H, Alam M, Takahashi H, Kharbanda A, Guha M, Ahmad R et al. MUC1-C oncoprotein activates the ZEB1/miR-200c regulatory loop and epithelial-mesenchymal transition. Oncogene 2014; 33: 1680–1689.
Alam M, Rajabi H, Ahmad R, Jin C, Kufe D . Targeting the MUC1-C oncoprotein inhibits self-renewal capacity of breast cancer cells. Oncotarget 2014; 5: 2622–2634.
Raina D, Kosugi M, Ahmad R, Panchamoorthy G, Rajabi H, Alam M et al. Dependence on the MUC1-C oncoprotein in non-small cell lung cancer cells. Mol Cancer Ther 2011; 10: 806–816.
Shen N, Yan F, Pang J, Wu LC, Al-Kali A, Litzow MR et al. A nucleolin-DNMT1 regulatory axis in acute myeloid leukemogenesis. Oncotarget 2014; 5: 5494–5509.
Berger N, Ben Bassat H, Klein BY, Laskov R . Cytotoxicity of NF-kappaB inhibitors Bay 11-7085 and caffeic acid phenethyl ester to Ramos and other human B-lymphoma cell lines. Exp Hematol 2007; 35: 1495–1509.
Yang AS, Estecio MR, Doshi K, Kondo Y, Tajara EH, Issa JP . A simple method for estimating global DNA methylation using bisulfite PCR of repetitive DNA elements. Nucleic Acids Res 2004; 32: e38.
Weisenberger DJ, Campan M, Long TI, Kim M, Woods C, Fiala E et al. Analysis of repetitive element DNA methylation by MethyLight. Nucleic Acids Res 2005; 33: 6823–6836.
Lisanti S, Omar WA, Tomaszewski B, De Prins S, Jacobs G, Koppen G et al. Comparison of methods for quantification of global DNA methylation in human cells and tissues. PLoS ONE 2013; 8: e79044.
Sharma S, Kelly TK, Jones PA . Epigenetics in cancer. Carcinogenesis 2010; 31: 27–36.
De Carvalho DD, Sharma S, You JS, Su SF, Taberlay PC, Kelly TK et al. DNA methylation screening identifies driver epigenetic events of cancer cell survival. Cancer Cell 2012; 21: 655–667.
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J et al. Initial sequencing and analysis of the human genome. Nature 2001; 409: 860–921.
Rodic N, Sharma R, Sharma R, Zampella J, Dai L, Taylor MS et al. Long interspersed element-1 protein expression is a hallmark of many human cancers. Am J Pathol 2014; 184: 1280–1286.
Iskow RC, McCabe MT, Mills RE, Torene S, Pittard WS, Neuwald AF et al. Natural mutagenesis of human genomes by endogenous retrotransposons. Cell 2010; 141: 1253–1261.
Tubio JM, Li Y, Ju YS, Martincorena I, Cooke SL, Tojo M et al. Mobile DNA in cancer. Extensive transduction of nonrepetitive DNA mediated by L1 retrotransposition in cancer genomes. Science 2014; 345: 1251343.
Gnemmi V, Bouillez A, Gaudelot K, Hemon B, Ringot B, Pottier N et al. MUC1 drives epithelial-mesenchymal transition in renal carcinoma through Wnt/beta-catenin pathway and interaction with SNAIL promoter. Cancer Lett 2014; 346: 225–236.
Zrihan-Licht S, Weiss M, Keydar I, Wreschner DH . DNA methylation status of the MUC1 gene coding for a breast-cancer-associated protein. Int J Cancer 1995; 62: 245–251.
Yamada N, Nishida Y, Tsutsumida H, Hamada T, Goto M, Higashi M et al. MUC1 expression is regulated by DNA methylation and histone H3 lysine 9 modification in cancer cells. Cancer Res 2008; 68: 2708–2716.
Hasegawa M, Takahashi H, Rajabi H, Alam M, Suzuki Y, Yin L et al. Functional interactions of the cystine/glutamate antiporter, CD44v and MUC1-C oncoprotein in triple-negative breast cancer cells. Oncotarget 2016; 7: 11756–11769.
Acknowledgements
Research reported in this paper was supported by the National Cancer Institute of the National Institutes of Health under award numbers CA97098 and CA166480, and by the Lung Cancer Research Foundation.
Author contributions
HR proposed the studies, generated the promoter-reporter vectors and performed ChIP, DNA methylation ELISA and MeDIP assays. AT, MA, AB and YS performed ChIP, promoter-reporter and immunoblotting assays. SP performed bioinformatics analysis. DK designed the studies and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
DK holds equity in Genus Oncology and is a consultant to the company. The remaining authors disclosed no potential conflicts of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Rajabi, H., Tagde, A., Alam, M. et al. DNA methylation by DNMT1 and DNMT3b methyltransferases is driven by the MUC1-C oncoprotein in human carcinoma cells. Oncogene 35, 6439–6445 (2016). https://doi.org/10.1038/onc.2016.180
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.180
This article is cited by
-
DNA methyltransferase 1 inhibits microRNA-497 and elevates GPRC5A expression to promote chemotherapy resistance and metastasis in breast cancer
Cancer Cell International (2022)
-
DNMT1 facilitates growth of breast cancer by inducing MEG3 hyper-methylation
Cancer Cell International (2022)
-
Distinct global DNA methylation and NF-κB expression profile of preimplantation biopsies from ideal and non-ideal kidneys
Journal of Nephrology (2022)
-
GATA6 promotes epithelial-mesenchymal transition and metastasis through MUC1/β-catenin pathway in cholangiocarcinoma
Cell Death & Disease (2020)
-
DREAM complex suppresses DNA methylation maintenance genes and precludes DNA hypermethylation
Nature Plants (2020)