Abstract
Two animal models of Down syndrome (human trisomy 21) with segmental trisomy for all (Ts65Dn) or part (Ts1Cje) of human chromosome 21-homologous region of mouse chromosome 16 have cognitive and behavioral abnormalities. To compare these trisomies directly and to assess the phenotypic contribution of the region of difference between them, Ts65Dn, Ts1Cje, and a new segmental trisomic (Ms1Ts65) for the region of difference (App to Sod1) have been generated as littermates and tested in parallel. Although the performance of Ts1Cje mice in the Morris water maze is similar to that of Ts65Dn mice, the reverse probe tests indicate that Ts65Dn is more severely affected. By contrast, the deficits of Ms1Ts65 mice are significantly less severe than those of Ts65Dn. Therefore, whereas triplication of Sod1 to Mx1 plays the major role in causing the abnormalities of Ts65Dn in the Morris water maze, imbalance of App to Sod1 also contributes to the poor performance. Ts65Dn mice are hyperactive and Ts1Cje mice are hypoactive; the activity of Ms1Ts65 mice is not significantly above normal. These findings indicate that genes in the Ms1Ts65 trisomic region must interact with others in the Ts1Cje region to produce hyperactivity in Ts65Dn mice.
Similar content being viewed by others
Main
Although the complex phenotype of DS is believed to result from the overexpression of a large number of genes present on human chromosome 21 (HSA 21) (1, 2), the analysis of individuals with segmental trisomies of HSA 21 has helped to localize some components of the phenotype to subregions of this chromosome (3–5). To permit a more detailed analysis of the relationship between the imbalance of individual genes or chromosome regions and specific phenotypic abnormalities, several animal models of human trisomy 21 have been developed (6–9). Mice with segmental trisomies of the HSA 21 homologous region on MMU 16 have demonstrated cognitive and behavioral abnormalities (6, 8) that are susceptible to genetic dissection. Several laboratories have shown that Ts65Dn mice, with a segmental trisomy extending from proximal of App to Mx1, do not perform as well as controls in the Morris water maze (6, 10, 11). This task, in which the subject has to use visual-spatial cues in the environment to navigate to a platform hidden under the surface of a tank of water, has been repeatedly demonstrated to be sensitive to lesions of the hippocampus in normal animals (12). Another segmentally trisomic mouse, Ts1Cje, is genetically similar to Ts65Dn but does not have triplication of the region from App to Sod1 (Fig. 1). Mice with this trisomy also show impaired performance in the Morris water maze, indicating that segmental trisomy for the distal part of the DS-homologous region of MMU 16 is responsible for hippocampal dysfunction (8). However, because Ts1Cje and Ts65Dn were bred on different genetic backgrounds and different protocols and apparatus were used for behavioral testing, it was not possible to determine whether the phenotype of Ts1Cje, with the smaller region of triplication, differed significantly from that of Ts65Dn. Therefore, we have generated Ts65Dn and Ts1Cje mice as littermates and, in the present article, describe the comparison of these segmental trisomics with the same behavioral tests. In addition, we have generated and examined a new mouse segmentally trisomic for the region by which Ts1Cje and Ts65Dn differ. These animals, designated Ms1Ts65, have permitted us to assess the contribution of the region from App to Sod1 to the overall Ts65Dn phenotype (Fig. 1).
Diagram drawn to scale (cM) of the triplicated regions in segmental trisomy 16 mice (Ms1Ts65, Ts1Cje, and Ts65Dn). The triplicated region in Ts1Cje spans from Sod1 to Mx1. However, Ts1Cje is not functionally triplicated for Sod1 because the Sod1 gene in the translocated segment has been inactivated by the insertion of the neomycin resistance sequence. App, amyloid beta (A4) precursor protein;Grik1, glutamate receptor, ionotropic, kainate1;Sod1, superoxide dismutase 1;Gart, phosphoribosylglycinamide formyltransferase;Sim1, simple-minded 2;Dryk1, dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1a;Ets2, E26 avian leukemia oncogene;Mx1, myxovirus (influenza virus) resistence-1.
METHODS
Mice.
T(12;16)1Cje animals that carry the CuZnSOD c allele (c/-) were generated by mating T(12;16)1Cje animals on a CD1 background (a/-) to homozygous (c/c) SOD1/Ei (JAX #JR1224) mice. Ts65Dn animals were generated by mating Ts65Dn female mice, which were originally obtained from M. Davisson (Jackson Laboratory), with (C57BL/6JEi × C3H/HeJ)F1 [JAX #JR1875] males (13). All animal breeding and studies were approved by the UCSF Committee on Animal Research.
Breeding the segmental trisomies.
Balanced carriers of the translocation designated T(12;16)1Cje have one each of normal chromosomes 12 and 16 and of translocation chromosomes 1216 and 1612 (Fig. 2a). The T(12;16) translocation breakpoint is proximal to Sod1 (Fig. 1). Ts65Dn mice have 41 chromosomes including the small 1716 translocation chromosome (Fig. 2a). Because Ts65Dn male mice are sterile, Ts65Dn females were bred to balanced T(12;16)1Cje males. Eight different genotypes, which include wild type and three different segmental trisomies (Fig. 2b), two early lethals, and two other viables (Table 1), are obtained from this cross. In addition to the segmental trisomiesTs65Dn and Ts1Cje, a new segmental trisomy, Ms1CjeTs65Dn (designated Ms1Ts65), is also generated (Figs. 1 and 2B).
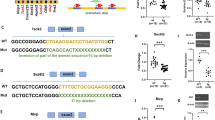
(a) Schematic diagram of the normal and translocation chromosomes used to generate the three segmental trisomies with the location/existence of Sod1 indicated. Note that Sod1 is missing from 1612 and that the inactivated form of Sod1, labeled Sod1-(neo +), is present on 1216. (b) Generation of the three segmental trisomies from the cross of Ts65Dn female and T(12;16)1Cje male mice. In addition to the three sequential trisomies and wild-type mice used for analysis, four other genotypes, two lethal and two viable, are also generated (See Table 1). The Sod1 allelic variants are identified as Sod1A, Sod1C, Sod1-(neo +) (functionally null). (c) Genotype identification of diploid and segmental trisomy 16 mice (PCR, upper; SOD gel, middle; cytogenetic analysis, bottom). Upper : A specific 176 bp fragment was amplified from the neomycin resistance gene (neo) and a 333 bp fragment was amplified from the Grik1 gene on chromosome 16 as an internal control (control). Offspring with a neo specific fragment have a 1216 chromosome. Middle : The AC heterodimeric SOD1 protein is present between the CC and AA homodimers. The presence of the 1612 chromosome is indicated by the detection of the AA homodimer without AC or CC. Bottom : The 1716 chromosome is directly detected by cytogenetic analysis as shown.
Genotyping.
Primers for PCR amplification of the neomycin resistance sequence (neo) were Neo3: 5′-ctcaccttgctcctgccgag-3′ and Neo4: 5′-ctgatgctcttcgtccagatcatc-3′, and those for internal controls were Grik1F2: 5′-ccccttagcataacgaccag-3′ and Grik1R2: 5′-ggcacgagacagacactgag-3′. PCR was performed using the following reaction conditions: “hot start” followed by 30 cycles of 94°C, 45 s; 55°C, 60 s in a 25 μL reaction mixture containing DNA (50–100 ng), 10 mM Tris-HCl (pH 8.3), 50 mM KCl, 1.5 mM MgCl2, 0.001% gelatin, 0.2 mM dNTPs, 0.4 μM of each primer, and 0.7 units of AmpliTaq polymerase (Applied Biosystems, Foster City, CA, U.S.A.). The CuZnSOD a and c isoforms were detected by staining for SOD activity from blood samples as described (14). Conventional karyotyping was performed using phytohemagglutinin-stimulated lymphocytes as described (15).
The 1216 chromosome is marked by a neomycin resistance sequence and is identified by PCR for neo (Fig. 2c, top). The 1612 chromosome lacks Sod1 and is identified by tracking the CuZnSOD protein polymorphism (AA, AC, or CC) (Fig. 2c, middle). The 1716 chromosome is identified as a small chromosome by cytogenetic analysis(Fig. 2c, bottom). Therefore, each genotype is identified by a combination of PCR, SOD gel staining, and cytogenetic analysis.
Morris water maze.
Four groups of approximately equal numbers of male and female mice that were 20–24 wk of age were tested: diploid controls (n = 17, wt 30.9 ± 1.9 g, age 23.7 ± 1.0 wk), Ms1Ts65 (n = 16, wt 32.4 ± 1.6 g, age 21.4 ± 1.4 wk), Ts1Cje (n = 12, wt 30.6 ± 1.7 g, age 20.1 ± 1.3 wk), and Ts65Dn (n = 18, wt 34.8 ± 1.8 g, age 20.9 ± 0.8 wk). The Morris water maze tests were performed as described (7) using a 1.2-m diameter circular pool filled with water made opaque by the addition of white nontoxic powder paint and were monitored by a video camera connected to a digital tracking device (San Diego Instruments, San Diego, CA, U.S.A.). The distance of the pool from the walls of the room varied between 35 and 50 cm. The mice were first repeatedly tested for 2 d for their ability to locate a submerged platform (14.5 cm in diameter) that was marked with a flag (the visible platform test). The next week they were tested for 3 d for their ability to locate an unmarked platform that was submerged 1.5 cm beneath the surface of the water (the hidden platform test), followed by 2 d of testing in which the platform was switched to the opposite quadrant (the reverse hidden platform test). One hour following the final trial of each of the latter two tests, the platform was removed and the times spent in each quadrant (dwell) and the number of times the mice crossed the imaginary site of the platform (crossings) were recorded for 1 min (as the probe and reverse probe tests, respectively). Each training block represents four different trials in which the mice were released in a pseudorandom fashion from each of four quadrants. Three blocks of trials were performed per day. The mice were allowed to stay on the platform for 10 s before the next trial. Mice not finding the platform within 60 s were immediately placed on the platform for 60 s and the next trial was then begun.
Spontaneous locomotor activity.
Spontaneous activity was examined as described (7) several days before or after the Morris water maze test. Four groups of mice (diploid control n = 16, Ms1Ts65 n = 16, Ts1Cje n = 12, Ts65Dn n = 18) were placed in a cage (47 × 26 × 15 cm) for 1 h in the dark during their light phase and were monitored by equally spaced infrared beams along the long and short axes.
Data analysis.
Repeated measures ANOVA was used to analyze latencies and path lengths in the platform tests and spontaneous locomotor activity. This provided tests of an overall difference between four groups, differences between blocks, and group by block interaction. Repeated measures ANOVA was also used for pairwise comparisons of two groups. One-sample t tests were used separately for each group to test for shorter latencies and path lengths in the last block than in the first block and to test for >25% time spent in the trained quadrant in the probe dwell tests. They were also used to test for the significance of the preference scores, which were calculated as the number of crossings of the trained site minus the mean number of crossings of the equivalent sites in other quadrants (16). Two sample t tests were used to compare the groups with regard to the time spent in the trained quadrant, the numbers of crossings over the trained site, and the preference scores. For the distribution of live born progeny, the null hypothesis of equal probabilities of each genotype was tested using a likelihood ratio test for a multinominal distribution. All values in the text and figures are expressed as mean ± SEM.
RESULTS
Generation of mice with different segmental trisomies of MMU 16(Ms1CjeTs65Dn, Ts1Cje, Ts65Dn).
All of the expected viable genotypes were obtained in equal frequencies, and the lethal genotypes Ms1Cje (a segmental monosomy) and Ts1CjeTs65Dn (a segmental tetrasomy), were, as expected, not observed (Table 1).
Performance of the three segmental trisomics in the Morris watermaze.
The three segmental trisomics (Ms1Ts65, Ts1Cje, and Ts65Dn), along with diploid controls, were tested in the Morris water maze. In the visible platform test, in which the mice need only learn to associate the flag with the platform to solve the task and spatial information is irrelevant, all groups of mice showed significant reduction in latency (search time) over successive trials. Both Ts1Cje and Ts65Dn mice performed as efficiently as control mice (Fig. 3 and Table 2), although the reduction in latency for Ms1Ts65 mice was less than for diploid controls in the first trial block only (Fig. 3). Nevertheless, all groups of mice appeared to be normal with respect to vision, swimming ability, and motivation to escape from water.
The visible platform test of the Morris water maze for control, Ms1Ts65, Ts1Cje, and Ts65Dn mice. Mean latency: time to reach the platform. Performance improved over blocks (p < 0.0001). Although the effect of genotype was not statistically significant (p = 0.14) and there was no statistically significant block by genotype interaction (p = 0.098), there was a statistically significant difference between Ms1Ts65 and diploid control mice in overall performance (p = 0.039) with the latency being longer than controls in block 1 (p = 0.004).
In the hidden platform test, the mice must learn the spatial relationships between objects in the room and the position of the platform to escape from the water. Control mice showed significant reductions in latency over the nine blocks of training [3 per day for 3 d] (Fig. 4a). Ms1Ts65 and Ts1Cje mice also showed reduced latencies, although Ms1Ts65 mice took significantly longer than controls to locate the hidden platform in the middle blocks and Ts1Cje mice were significantly slower than controls over all nine blocks (Fig. 4a and Table 2). The performance of Ts65Dn was significantly impaired and was not improved over the blocks (Fig. 4a and Table 2). The findings for path lengths closely paralleled those of escape latencies, with path length shortening as latency decreased (Fig. 4, a and b). These findings indicate that decreases in latency are truly learning effects resulting from the use of spatial cues to shorten path length.
The hidden platform phases of the Morris water maze. Mean latency: time to reach the platform. Dwell: the proportion of time spent searching in each of the four quadrants of the pool. Crossings: the number of times the mice crossed regions of the pool marking the exact position of the former platform. Preference score: the number of crossings in the trained site minus the mean number of crossings from the other sites [Owen et al. (16)]. The trained sites were quadrants 1 in the probe tests. (a) Hidden platform test (latency). The effect of genotype was statistically significant (p = 0.001). Performance improved over blocks (p < 0.0001), and there was a statistically significant block and genotype interaction (p = 0.027). Diploid controls (p = 0.003), Ms1Ts65 (p = 0.026), and Ts1Cje (p = 0.021) showed significant decreases of latency, but Ts65Dn did not (p = 0.75). However, overall performances of Ms1Ts65 (p = 0.039), Ts1Cje (p = 0.0002), and Ts65Dn (p = 0.0002) were all impaired relative to controls. (b) Hidden platform test (path length). The effect of genotype was statistically significant (p = 0.008). Performance improved over blocks (p < 0.0001), and there was a statistically significant block and genotype interaction (p = 0.023). Diploid controls (p = 0.001), Ms1Ts65 (p = 0.0002), and Ts1Cje (p = 0.005) showed significant decreases of path length, but Ts65Dn (p = 0.24) did not. Overall performances of Ts1Cje (p = 0.004) and Ts65Dn (p = 0.004) were impaired compared with those of controls. However, that of Ms1Ts65 did not reach statistical significance (p = 0.14). (c) Probe test (dwell). All of groups (diploid control, p < 0.0001; Ms1Ts65, p < 0.0001; Ts1Cje, p = 0.005; Ts65Dn, p = 0.012) spent significantly more than 25% of the time in the trained quadrant. The time Ts65Dn mice spent in the trained quadrant was significantly less than that of controls (p = 0.008). The times for Ts1Cje were lower than controls, but this did not reach statistical significance (p = 0.070). There was no significant difference between Ms1Ts65 and controls (p = 0.22). (d) The probe test (crossings). Ts65Dn mice crossed the trained site significantly less frequently than controls (p = 0.003). Ts1Cje mice crossed the trained site less frequently than controls, but the difference just missed reaching statistical significance (p = 0.055). There was no significant difference between Ms1Ts65 and control mice (p = 0.34), but Ms1Ts65 mice were significantly better than Ts65Dn mice (p = 0.027). (e) The probe test preference score. All groups (diploid control, p < 0.0001; Ms1Ts65, p < 0.0001; Ts1Cje, p = 0.003; Ts65Dn, p = 0.0003) showed positive scores indicating that the mice crossed the trained platform more often than they crossed corresponding sites in other quadrants. The score for Ts65Dn mice is significantly lower than for controls (p = 0.019) and Ms1Ts65 (p = 0.048) mice. The score for Ts1Cje mice is also lower than for controls, but the difference did not reach statistical significance (p = 0.072). There is no difference between control and Ms1Ts65 mice (p = 0.51).
After the hidden platform test, the mice were tested in the probe trial. This test, which assesses spatial selectivity, is based on the premise that mice that had learned the location of the platform should search selectively where the platform had been located. All groups showed preference for the trained quadrant in terms of dwell (the time spent searching) (Fig. 4c) and crossings (the number of times the mouse crossed the position of the platform) (Fig. 4d), providing evidence for learning. However, Ts65Dn mice spent significantly less time in the trained quadrant and crossed the trained site significantly less frequently than did controls (Fig. 4, c and d, and Table 2). Although the preference of Ts1Cje mice for the trained site appeared to be reduced compared with control mice, the differences in dwell, crossings, and preference score just missed reaching statistical significance (Fig. 4, c–e, and Table 2). The performance of Ms1Ts65 mice was not significantly different from that of the controls (Fig. 4, c–e, and Table 2).
In the reverse platform test, the mice are required to learn a novel position for the hidden platform that has been moved to the quadrant opposite to its original location. Both Ts1Cje and Ms1Ts65 mice showed a decrease in latency over the six blocks of the test (Fig. 5a), but the rates of decrease were significantly less than for the controls (Fig. 5a and Table 2). Ts65Dn mice showed no decrease in latency (Fig. 5a). The findings for path lengths closely paralleled those for escape latencies (Fig. 5b). Although the difference in latency between Ms1Ts65 and Ts1Cje did not reach statistical significance, Ms1Ts65 was significantly better than Ts65Dn whereas Ts1Cje was not (Table 3).
The reverse hidden platform phases of the Morris water maze. The trained site was quadrant 3 in the reverse probe tests. (a) Reverse hidden platform test (latency). The effect of genotype was statistically significant (p < 0.0001). Performance improved over blocks (p < 0.0001), but there was no statistically significant block by genotype interaction (p = 0.79). Diploid controls (p < 0.0001), Ms1Ts65 (p = 0.002), and Ts1Cje (p = 0.022) showed statistically significant decreases of latency, but the decreases for Ts65Dn did not reach statistical significance (p = 0.087). However, overall performances of Ms1Ts65 (p = 0.048), Ts1Cje (p = 0.002), and Ts65Dn (p < 0.0001) were all impaired compared with those of controls. There was also a significant difference between Ms1Ts65 and Ts65Dn mice in overall performance (p = 0.001). (b) Reverse hidden platform test (path length). The effect of genotype was statistically significant (p = 0.004). Performance improved over blocks (p < 0.0001), but there was no statistically significant block by genotype interaction (p = 0.79). All groups (diploid controls, p < 0.0001; Ms1Ts65, p < 0.0001; Ts1Cje, p = 0.0002; Ts65Dn, p = 0.007) showed statistically significant decreases of path length. The overall performance of Ts65Dn (p = 0.0006) was impaired compared with that of controls, but comparisons of Ms1Ts65 (p = 0.22) and of Ts1Cje (p = 0.092) with controls did not reach statistical significance. There was a significant difference between Ms1Ts65 and Ts65Dn mice in overall performance (p = 0.005). (c) Reverse probe test (dwell). Only diploid control mice spent significantly more than 25% of the time in the trained quadrant (p = 0.005). The difference in the time spent in the trained quadrant compared with controls was statistically significant for Ts65Dn mice (p = 0.001), but not for Ms1Ts65 (p = 0.19) nor for Ts1Cje mice (p = 0.71). Ts65Dn mice were also significantly poorer than Ts1Cje mice on this measure (p = 0.045). Ts65Dn mice spent significantly more than 25% of the time in quadrant 1, the quadrant in which the platform was previously located in the hidden platform test (p = 0.0008), but this did not reach statistical significance for the Ts1Cje mice (p = 0.086). The difference in the time spent in quadrant 1 compared with controls was statistically significant for Ts65Dn (p = 0.012) but not for Ts1Cje (p = 0.24) mice. (d) Reverse probe test (crossings). Ts65Dn mice crossed the trained site significantly less frequently than diploid controls (p = 0.0004), but this difference was not significant for the Ms1Ts65 (p = 0.11) nor for Ts1Cje (p = 0.17) mice. Ts65Dn mice were significantly poorer than Ms1Ts65 mice (p = 0.022), but the comparison with Ts1Cje did not reach significance (p = 0.072). (e) Reverse probe test (preference score). Diploid control (p < 0.0001), Ms1Ts65 (p = 0.0008), and Ts1Cje (p = 0.010) crossed the trained platform more often than other corresponding sites in other quadrants, but this did not quite reach statistical significance for the Ts65Dn mice (p = 0.067). The score for Ts65Dn mice (p = 0.004) but not Ms1Ts65 (p = 0.068) nor Ts1Cje (p = 0.30) was significantly lower than that for controls. The score in Ts65Dn mice was not significantly lower than for Ms1Ts65 and Ts1Cje mice (p = 0.18 and p = 0.15, respectively).
In the reverse probe dwell test, there was no preference for the trained quadrant in any of the trisomic mice (Fig. 5c and Table 2). Interestingly, Ts65Dn mice continued to show a preference for the initial trained site in quadrant 1 (Fig. 5c and Table 2). In the reverse probe crossings test and preference score, only Ts65Dn mice failed to show a preference for the trained site (Fig. 5, d and e, and Table 2). Ms1Ts65 and Ts1Cje did significantly or close to significantly better than Ts65Dn in reverse crossings and dwell (Table 3).
Spontaneous locomotor activity.
The total locomotor activity of all groups decreased over the period of testing, indicating habituation to a novel environment (Fig. 6). There were no significant differences in any of the trisomy groups compared with diploid controls (Fig. 6 and Table 2). However, there was a significant difference between Ts65Dn and Ts1Cje mice resulting from hyperactivity in Ts65Dn and from hypoactivity in Ts1Cje (Fig. 6 and Table 3).
Spontaneous locomotor activity of four groups (diploid control, Ms1Ts65, Ts1Cje, and Ts65Dn). The activity is shown in 12 sequential blocks of 5 min each. The effect of genotype was not quite statistically significant (p = 0.081). Activity decreased over blocks (p < 0.0001), but the block by genotype interaction did not quite reach statistical significance (p = 0.083). However, there is a significant difference between Ts65Dn and Ts1Cje mice in overall activity (p = 0.022).
DISCUSSION
Ts65Dn and Ts1Cje mice are generated as littermates when Ts65Dn and T(12;16)1Cje mice, used to generate two previously described segmental trisomy 16 models of DS, are crossed. In addition, a new segmental trisomy, Ms1Ts65, which involves the region of difference between the two original partial trisomies, is also generated (Fig. 2). This permits direct comparisons among the three trisomics and enables us to assess the contributions of the regions above and below Sod1 to the learning and behavioral phenotype.
To maintain fertility and viability, it is necessary to carry the segmentally trisomic strains on mixed genetic backgrounds. Therefore, it is not possible to breed them on inbred backgrounds. As a result, the genetic background on which the different segmental trisomics are bred cannot be precisely defined. Nevertheless, by generating the wild-type and three different segmentally trisomic mice as littermates, we have attempted to randomize the distribution of the different background alleles across the study population. Although the effect, if anything, of this lack of precise uniformity would be to obscure differences among the several strains of mice, we were able to demonstrate significant differences. Furthermore, although, as noted below, there were small differences between the results reported here and previous reports, our results were in general agreement with our own earlier findings for Ts65Dn and Ts1Cje and with the findings of other investigators for Ts65Dn.
Given the fact the differences in the hidden platform probe findings just barely fail to reach statistical significance (Table 2), the results for Ts1Cje relative to controls are quite similar to those previously reported for Ts1Cje and control mice on a different genetic background (8). Ts65Dn mice show statistically significant deficits in all the tests in the Morris water maze except for the visible platform test (6, 10, 11). These findings are consistent with previous reports except for the present observation of normal visible platform tests. The latter result suggests that the performance of Ts65Dn mice on the visible platform test may be altered by genetic background. Except in the reverse probe tests, the learning deficits of Ts1Cje mice in the Morris water maze are similar to those of Ts65Dn (Tables 2 and 3). These findings indicate that an important gene or genes involved in these deficits lie in the region of overlap between these mice, the region from Sod1 to Mx1 that contains the so-called “DS region” of human chromosome segment 21q22.
Ms1Ts65 mice show statistically significant deficits compared with controls in the latencies of the visible, hidden, and reverse hidden platform tests, but not in path length nor any of the probe tests (Table 2). The path length and probe results indicate that, although their performance does not match controls, they had little impairment in learning the task. Examination of the pairwise comparison between Ms1Ts65 and Ts65Dn demonstrates that the deficits of Ms1Ts65 mice in the Morris water maze are significantly less severe than those of Ts65Dn. Therefore, we conclude that, whereas triplication of the region from Sod1 to Mx1 plays the major role in causing the abnormalities of Ts65Dn in the Morris water maze, imbalance of the region from App to Sod1 also contributes to the poor performance.
Deficits on tasks that have repeatedly been demonstrated to depend upon a particular brain region in normal animals cannot be assumed to indicate malfunction of that region in genetically manipulated animals because we do not directly measure the underlying process. Rather, we examine performance of a task (e.g. spatial learning in the Morris maze) indicative of that process. A variety of factors, such as impaired sensory or motor function and differences in emotionality or motivation, can affect performance. These factors must be ruled out before conclusions can be made about the underlying processes. In the current study, the normal performance of all three segmental trisomics in the visible task of the Morris water maze rules out performance factors as accounting for impairment in the other tasks. This allows us to conclude that the deficits in hidden platform task performance indicate impairment of hippocampal function. Most measures also indicate that the larger the trisomic segment, the larger the deficit on this task. However, it is important to note even the most impaired animals learned the hidden platform task, indicating that although hippocampal function is impaired, it is not eliminated in any of the segmental trisomies. This is supported by other studies that show that extensive training improves the performance of both Ts1Cje (8) and Ts65Dn mice (17).
Overall, the performance of Ts1Cje mice in the Morris water maze is similar to that of Ts65Dn mice, but there is a significant difference between the two groups in the reverse probe dwell test and a nearly significant difference in reverse probe crossings (Table 3). Reversal learning involves two processes: inhibiting the previously learned response and acquiring the new response. The probe tests indicate that the Ts65Dn mice are most severely impaired on this task and that the deficit consists of failure to inhibit search in the initial platform position and consequent failure to develop any preference for the new platform position. Together with hyperactivity, which can be interpreted as a failure of inhibitory control, difficulty in reversal learning indicates deficits in function of the prefrontal cortex. Failure of Ts65Dn mice to inhibit exploration of the open arms of the elevated plus maze (10, 18, 19) supports this conclusion, as does the difficulty in shifting from a thigmotaxic strategy in the visible platform task (10). Thus, as in DS, there is evidence for mild to moderate impairment of hippocampal and prefrontal cortex function. Tasks that require both of these regions, such as the reversal of spatial learning in the reverse probe tests, are especially impaired.
There is a significant difference between Ts1Cje and Ts65Dn in spontaneous locomotor activity (Table 3). Ts65Dn mice tend to be hyperactive and Ts1Cje mice tend to be hypoactive (Fig. 6). Several groups have reported that Ts65Dn mice are hyperactive, probably because of deficits in controlling and inhibiting behavior (6, 11, 18, 19). Our previous study suggested that Ts1Cje mice are hypoactive because of reduced exploratory activity (8). It might be inferred that the genes involved in producing hyperactivity in the Ts65Dn mice are in the region between App and Sod1, as Ts1Cje mice are hypoactive. However, the Ms1Ts65 mice are not significantly above normal in activity, indicating that genes in the Sod1-App region must interact with others in the Ts65Dn region to produce hyperactivity in the Ts65Dn mice.
The principal reason for generating Ms1Ts65 mice was to assess the contribution of the segment of MMU 16 from proximal to App to Sod1 to the phenotype of Ts65Dn as the first step in carrying out a genetic dissection of the phenotype of this model of human trisomy 21. Our previous results with Ts1Cje indicated that virtually all of the behavioral deficits found in Ts65Dn are also exhibited by animals with the shorter region of segmental trisomy. Our present findings, which permit the simultaneous comparison of Ts1Cje and Ts65Dn with one another and with Ms1Ts65, confirm the earlier result and show that the overall contribution of App-Sod1 to the Ts65Dn phenotype is much smaller than that of Sod1-Mx1. Nevertheless, Ms1Ts65 animals are not normal, indicating that the imbalance of the App-Sod1 region is not without effect on the function of the brain.
A full genetic dissection of the behavioral phenotype of Ts65Dn will require a much more extensive series of testing paradigms than were used in these studies. Nevertheless, the present results indicate that such a dissection, which will make use of the systematic deletion of one or several genes from the Ts65Dn and Ts1Cje trisomic regions, will be feasible.
Abbreviations
- DS:
-
Down syndrome
- MMU 16:
-
mouse chromosome 16
References
Epstein CJ 1986 The Consequences of Chromosome Imbalance; Principles, Mechanisms, and Models. Cambridge University Press, New York, 253–323.
Epstein CJ 1988 Mechanisms of the effects of aneuploidy in mammals. Ann Rev Genet 22: 51–75.
McCormick MK, Schinzel A, Petersen MB, Stetten G, Driscoll DJ, Cantu ES, Tranebjaerg L, Mikkelsen M, Watkins PC, Antonarakis SE 1989 Molecular genetic approach to the characterization of the “Down syndrome region” of chromosome 21. Genomics 5: 325–331.
Korenberg JR, Bradley C, Disteche CM 1992 Down syndrome: molecular mapping of the congenital heart disease and duodenal stenosis. Am J Hum Genet 50: 294–302.
Korenberg JR, Chen XN, Schipper R, Sun Z, Gonsky R, Gerwehr S, Carpenter N, Daumer C, Dignan P, Disteche C, Graham JM Jr, Hudgins L, McGillivray B, Miyazaki J, Ogasawara N, Park JP, Pagon R, Pueschel S, Sack G, Say B, Schuttenhauer S, Soukap S, Yamanaka T 1994 Down syndrome phenotypes: the consequences of chromosomal imbalance. Proc Natl Acad Sci U S A 91: 4997–5001.
Reeves RH, Irving NG, Moran TH, Wohn A, Kitt C, Sisodia SS, Schmidt C, Bronson RT, Davisson MT 1995 A mouse model for Down syndrome exhibits learning and behaviour deficits. Nat Genet 11: 177–184.
Smith DJ, Stevens ME, Sudanagunta SP, Bronson RT, Makhinson M, Watabe AM, O'Dell TJ, Fung J, Weier HUG, Chen JF, Rubin EM 1997 Functional screening of 2 Mb of human chromosome 21q22.2 in transgenic mice implicates minibrain in learning defects associated with Down syndrome. Nat Genet 16: 28–36.
Sago H, Carlson EJ, Smith DJ, Kilbridge J, Rubin EM, Mobley WC, Epstein CJ, Huang T-T 1998 Ts1Cje, a partial trisomy 16 mouse models for Down syndrome, exhibits learning and behavioral abnormalities. Proc Natl Acad Sci U S A 95: 6256–6261.
Kola I, Hertzog P 1998 Down syndrome and mouse models. Curr Opin Genet Dev 8: 316–321.
Escorihuela RM, Fernandez-Teruel A, Vallina IF, Baamonde C, Lumbreras MA, Dierssen M, Tobena A, Florez J 1995 A behavioral assessment of Ts65Dn mice: a putative Down syndrome model. Neurosci Lett 199: 143–146.
Holtzman DM, Santucci D, Kilbridge J, Chua-Couzens J, Fontana DJ, Daniels SE, Johnson RM, Chen K, Sun Y, Carlson E, Alleva E, Epstein CJ, Mobley WC 1996 Developmental abnormalities and age-related neurodegeneration in a mouse model of Down syndrome. Proc Natl Acad Sci U S A 93: 13333–13338.
Morris RG, Garrud P, Rawlins JN, O'Keefe J 1982 Place navigation impaired in rats with hippocampal lesions. Nature 297: 681–683.
Davisson MT, Schmidt C, Reeves RH, Irving NG, Akeson EC, Harris BS, Bronson RT 1993 Segmental trisomy as a mouse model for Down syndrome. Prog Clin Biol Res 384: 117–133.
Luche RM, Maiwald R, Carlson EJ, Epstein CJ 1997 Novel mutations in an otherwise strictly conserved domain of CuZn superoxide dismutase. Mol Cell Biochem 168: 191–194.
Davisson MT, Akeson EC 1987 An improved method for preparing G-banded chromosomes from mouse peripheral blood. Cytogenet Cell Genet 45: 70–74.
Owen EH, Logue SF, Rasmussen DL, Wehner JM 1997 Assessment of learning by the Morris water task and fear conditioning in inbred mouse strains and F1 hybrids: implications of genetic background for single gene mutations and quantitative trait loci analyses. Neuroscience 80: 1087–1099.
Escorihuela RM, Vallina IF, Martinez-Cue C, Baamonde C, Dierssen M, Tobena A, Florez J, Fernandez-Teruel A 1998 Impaired short- and long-term memory in Ts65Dn mice, a model for Down syndrome. Neurosci Lett 247: 171–174.
Coussons-Read ME, Crnic LS 1996 Behavioral assessment of the Ts65Dn mouse, a model for Down syndrome: altered behavior in the elevated plus maze and open field. Behav Genet 26: 7–13.
Demas GE, Nelson RJ, Kruger BK, Yarowsky PJ 1996 Spatial working memory deficits in segmental trisomic Ts65Dn mice. Behav Brain Res 82: 85–92.
Acknowledgements
The authors thank R. Gacayan and M. Doan for excellent animal care, A.M. Gillespie for technical assistance, M. Davisson for the original strain of Ts65Dn mice, and P. Bacchetti for statistical analysis.
Author information
Authors and Affiliations
Additional information
This work was supported by grants from the National Institutes of Health (AG08938, HD04024, HD17499, HD31498, and HL18574) and from the Adler Foundation and the McGowan Charitable Trust.
Dr. Charles J. Epstei, Department of Pediatrics, University of California, Box 0748, San Francisco, CA 94143-0748, U.S.A.
Rights and permissions
About this article
Cite this article
Sago, H., Carlson, E., Smith, D. et al. Genetic Dissection of Region Associated with Behavioral Abnormalities in Mouse Models for Down Syndrome. Pediatr Res 48, 606–613 (2000). https://doi.org/10.1203/00006450-200011000-00009
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/00006450-200011000-00009
This article is cited by
-
Impairment of spatial memory accuracy improved by Cbr1 copy number resumption and GABAB receptor-dependent enhancement of synaptic inhibition in Down syndrome model mice
Scientific Reports (2020)
-
Autonomous trisomic rescue of Down syndrome cells
Laboratory Investigation (2019)
-
Differential Brain, Cognitive and Motor Profiles Associated with Partial Trisomy. Modeling Down Syndrome in Mice
Behavior Genetics (2017)
-
Distinct Defects in Spine Formation or Pruning in Two Gene Duplication Mouse Models of Autism
Neuroscience Bulletin (2017)
-
Altered Distribution of Hippocampal Interneurons in the Murine Down Syndrome Model Ts65Dn
Neurochemical Research (2015)