Abstract
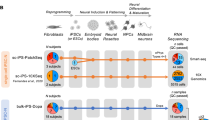
Parkinson’s disease (PD) is a progressive, neurodegenerative disease affecting over 1% of the population beyond 65 years of age. Although some PD cases are inheritable, the majority of PD cases occur in a sporadic manner. Risk factors comprise next to heredity, age, and gender also exposure to neurotoxins from for instance pesticides and herbicides. As PD is characterized by a loss of dopaminergic neurons in the substantia nigra, it is nearly impossible to access and extract these cells from patients for investigating disease mechanisms. The emergence of induced pluripotent stem (iPSC) technology allows differentiating and growing human dopaminergic neurons, which can be used for in vitro disease modeling. Here, we differentiated human iPSCs into dopaminergic neurons, and subsequently exposed the cells to increasing concentrations of the neurotoxin MPP+. Temporal transcriptomics analysis revealed a strong time- and dose-dependent response with genes over-represented across pathways involved in PD etiology such as “Parkinson’s Disease”, “Dopaminergic signaling” and “calcium signaling”. Moreover, we validated this disease model by showing robust overlap with a meta-analysis of transcriptomics data from substantia nigra from post-mortem PD patients. The overlap included genes linked to e.g. mitochondrial dysfunction, neuron differentiation, apoptosis and inflammation. Our data shows, that MPP+-induced, human iPSC-derived dopaminergic neurons present molecular perturbations as observed in the etiology of PD. Therefore we propose iPSC-derived dopaminergic neurons as a foundation for a novel sporadic PD model to study the pathomolecular mechanisms of PD, but also to screen for novel anti-PD drugs and to develop and test new treatment strategies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Data that support the findings of this study have been deposited in Gene Expression Omnibus with the accession code GSE196190.
References
Gonera EG, Van’t Hof M, Berger HJC, Van Weel C, Horstink MWIM. Symptoms and duration of the prodromal phase in Parkinson’s disease. Mov Disord. 1997;12:871–6.
Ray Dorsey E, Elbaz A, Nichols E, Abd-Allah F, Abdelalim A, Adsuar JC, et al. Global, regional, and national burden of Parkinson’s disease, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 2018;17:939–53.
Fearnley JM, Lees AJ. Ageing and parkinson’s disease: substantia nigra regional selectivity. Brain. 1991;114:2283–301.
Dauer W, Przedborski S. Parkinson’s disease: mechanisms and models. Neuron. 2003;39:889–909.
Moore TJ, Glenmullen J, Mattison DR. Reports of pathological gambling, hypersexuality, and compulsive shopping associated with dopamine receptor agonist drugs. JAMA Intern Med. 2014;174:1930–3.
Bastide MF, Meissner WG, Picconi B, Fasano S, Fernagut PO, Feyder M, et al. Pathophysiology of L-dopa-induced motor and non-motor complications in Parkinson’s disease. Prog Neurobiol. 2015;132:96–168.
Lashuel HA, Overk CR, Oueslati A, Masliah E. The many faces of α-synuclein: from structure and toxicity to therapeutic target. Nat Rev Neurosci. 2013;14:38–48.
Minakaki G, Menges S, Kittel A, Emmanouilidou E, Schaeffner I, Barkovits K, et al. Autophagy inhibition promotes SNCA/alpha-synuclein release and transfer via extracellular vesicles with a hybrid autophagosome-exosome-like phenotype. Autophagy. 2018;14:98–119.
Gao F, Yang J, Wang D, Li C, Fu Y, Wang H, et al. Mitophagy in Parkinson’s disease: pathogenic and therapeutic implications. Front Neurol. 2017;8:527.
Bose A, Beal MF. Mitochondrial dysfunction in Parkinson’s disease. J Neurochemistry. 2016;139:216–31.Suppl 1.
Kaur K, Gill JS, Bansal PK, Deshmukh R. Neuroinflammation - A major cause for striatal dopaminergic degeneration in Parkinson’s disease. J Neurological Sci. 2017;381:308–14.
Berry C, La Vecchia C, Nicotera P. Paraquat and parkinson’s disease. Cell Death Differ. 2010;17:1115–25.
Curtin K, Fleckenstein AE, Robison RJ, Crookston MJ, Smith KR, Hanson GR. Methamphetamine/amphetamine abuse and risk of Parkinson’s disease in Utah: A population-based assessment. Drug Alcohol Depend 2015;146:30–38.
Bohler S, Krauskopf J, Espín-Pérez A, Gebel S, Palli D, Rantakokko P, et al. Genes associated with Parkinson’s disease respond to increasing polychlorinated biphenyl levels in the blood of healthy females. Environ Pollut 2019;250:107–17.
Ke M, Chong CM, Su H. Using induced pluripotent stem cells for modeling Parkinson’s disease. World J Stem Cells. 2019;11:634–49.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663–76.
Kopin IJ. MPTP: An industrial chemical and contaminant of illicit narcotics stimulates a new era in research on Parkinson’s disease. Environ Health Perspect. 1987;75:45–51.
Glaab E, Schneider R. Comparative pathway and network analysis of brain transcriptome changes during adult aging and in Parkinson’s disease. Neurobiol Dis 2015;74:1–13.
Piñero J, Bravo À, Queralt-Rosinach N, Gutiérrez-Sacristán A, Deu-Pons J, Centeno E, et al. DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res. 2017;45:D833–D839.
Russell AC, Šimurina M, Garcia MT, Novokmet M, Wang Y, Rudan I, et al. The N-glycosylation of immunoglobulin G as a novel biomarker of Parkinson’s disease. Glycobiology. 2017;27:501–10.
Sandor C, Robertson P, Lang C, Heger A, Booth H, Vowles J, et al. Transcriptomic profiling of purified patient-derived dopamine neurons identifies convergent perturbations and therapeutics for Parkinson’s disease. Hum Mol Genet. 2017;26:552–66.
Fernandes HJR, Patikas N, Foskolou S, Field SF, Park JE, Byrne ML, et al. Single-cell transcriptomics of parkinson’s disease human in vitro models reveals dopamine neuron-specific stress responses. Cell Rep. 2020;33:108263.
Sacchetti P, Carpentier R, Ségard P, Olivé-Cren C, Lefebvre P. Multiple signaling pathways regulate the transcriptional activity of the orphan nuclear receptor NURR1. Nucleic Acids Res. 2006;34:5515–27.
Takahashi M, Suzuki M, Fukuoka M, Fujikake N, Watanabe S, Murata M, et al. Normalization of overexpressed α-synuclein causing Parkinson’s disease by a moderate gene silencing with RNA interference. Mol Ther - Nucleic Acids. 2015;4:e241.
Smirnova L, Harris G, Delp J, Valadares M, Pamies D, Hogberg HT, et al. A LUHMES 3D dopaminergic neuronal model for neurotoxicity testing allowing long-term exposure and cellular resilience analysis. Arch Toxicol. 2016;90:2725–43.
Pamies D, Wiersma D, Katt ME, Zhao L, Burtscher J, Harris G, et al. Human IPSC 3D brain model as a tool to study chemical-induced dopaminergic neuronal toxicity. Neurobiol Dis. 2022;169:105719.
Kriks S, Shim JW, Piao J, Ganat YM, Wakeman DR, Xie Z, et al. Dopamine neurons derived from human ES cells efficiently engraft in animal models of Parkinson’s disease. Nature. 2011;480:547–51.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–20.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinforma. 2011;12:323.
Radley AH, Schwab RM, Tan Y, Kim J, Lo EKW, Cahan P. Assessment of engineered cells using CellNet and RNA-seq. Nat Protoc 2017;12:1089–102.
Soneson C, Love MI, Robinson, MD. Differential analyses for RNA-seq: Transcript-level estimates improve gene-level inferences [version 2; referees: 2 approved]. F1000Research. 2016;4:1521.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple. Test J R Stat Soc Ser B. 1995;57:289–300.
Gu Z, Eils R, Schlesner M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics. 2016;32:2847–9.
Author information
Authors and Affiliations
Contributions
JCK and CV designed the experiments. KE, RFMC, SB, DH, and FC performed and organized the experimental work. JK analyzed the data. Advice and supervision were from JCK, CV, TMK and FC. JK wrote the manuscript and all authors reviewed and approved the final version of this article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Krauskopf, J., Eggermont, K., Madeiro Da Costa, R.F. et al. Transcriptomics analysis of human iPSC-derived dopaminergic neurons reveals a novel model for sporadic Parkinson’s disease. Mol Psychiatry 27, 4355–4367 (2022). https://doi.org/10.1038/s41380-022-01663-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-022-01663-y
This article is cited by
-
Induced pluripotent stem cells (iPSCs): molecular mechanisms of induction and applications
Signal Transduction and Targeted Therapy (2024)