Abstract
STING (Stimulator of Interferon Genes) is an endoplasmic reticulum-anchored adaptor of the innate immunity best known to trigger pro-inflammatory cytokine expression in response to pathogen infection. In cancer, this canonical pathway can be activated by intrinsic or drug-induced genomic instability, potentiating antitumor immune responses. Here we report that STING downregulation decreases cell survival and increases sensitivity to genotoxic treatment in a panel of breast cancer cell lines in a cell-autonomous manner. STING silencing impaired DNA Damage Response (53BP1) foci formation and increased DNA break accumulation. These newly identified properties were found to be independent of STING partner cGAS and of its canonical pro-inflammatory pathway. STING was shown to partially localize at the inner nuclear membrane in a variety of breast cancer cell models and clinical tumor samples. Interactomics analysis of nuclear STING identified several proteins of the DNA Damage Response, including the three proteins of the DNA-PK complex, further supporting a role of STING in the regulation of genomic stability. In breast and ovarian cancer patients that received adjuvant chemotherapy, high STING expression is associated with increased risk of relapse. In summary, this study highlights an alternative, non-canonical tumor-promoting role of STING that opposes its well-documented function in tumor immunosurveillance.
Similar content being viewed by others
Introduction
Stimulator of Interferon Genes (STING) is a key adaptor protein of the innate immune response to cytosolic DNA [1]. STING is anchored in the endoplasmic reticulum (ER) by its four N-terminal transmembrane domains. Its C-terminal tail resides in the cytoplasm and contains the cyclic dinucleotide (CDN)-binding domain and domains of interaction with downstream effectors [2, 3]. Upon infection, CDNs are directly secreted by pathogens or generated by the cyclic GMP-AMP synthase (cGAS) in response to cytosolic pathogen-derived DNA [4]. CDNs bind to and activate STING that recruits TANK-binding kinase 1 (TBK1) and the transcription factor IRF-3 (interferon [IFN] regulatory factor-3). TBK1 phosphorylates IRF-3, then phospho-IRF-3 homodimers translocate to the nucleus to induce the expression of IFNs [3]. NFκB signaling has also been shown to contribute to STING-mediated induction of pro-inflammatory genes but the molecular mechanisms involved are not fully elucidated [5,6,7,8].
STING is expressed in several cell types including cancer cells [9]. In cancer, STING signaling is best known to promote antitumor immune responses [10,11,12,13] which stimulated the development of therapeutic strategies involving STING agonists [14]. Otherwise, accumulating evidence suggests that the activation of the cGAS/STING/IFN pathway may in some instance promote cancer progression through chronic inflammation [15], enhanced DNA repair responses [16] and metastasis formation [7, 17].
We [18] and others (reviewed in ref. [19]) recently linked activation of STING inflammatory pathway to genotoxic stress in cancer. Genotoxic treatment of MCF7 breast cancer cells led to the accumulation of DNA in the cytoplasm and triggered the expression of IFNs and of several IFN-stimulated genes (ISGs) via the canonical STING/TBK1/IRF-3 pathway [18]. Similar observations were reported by others using several cell models treated with various DNA-damaging agents [10, 11, 13, 15, 20,21,22,23], or in the context of intrinsic genomic instability characteristic of cancer cells [21, 22, 24]. In particular, we showed that in vitro, i.e. in absence of a functional immune system, STING silencing potentiated chemotherapy-induced MCF7 cell death [18].
The emerging controversial functions of STING in cancer prompted us to investigate further its possible contribution to cancer cell survival. Here, we show that STING promotes survival and resistance to genotoxics of various immortalized and PDX-derived breast cancer cells. Loss of STING expression decreased 53BP1 foci formation and increased DNA instability independently of cGAS and IFN responses. We discovered that STING partially localizes inside breast cancer cell nucleus, and more particularly at the inner nuclear membrane (INM). Nuclear STING interactome analysis by mass spectrometry identified several proteins involved in the DNA damage response (DDR), including DNA-PK. In breast and ovarian cancer patients treated with chemotherapy, high STING expression predicts poor prognosis. Together, our data support that STING intrinsically promotes breast cancer cell survival via a non-canonical, cell-autonomous mechanism that opposes to its well-documented role in antitumor immunity.
Results
STING promotes intrinsic breast cancer cell survival and resistance to genotoxic stress
To investigate the impact of STING on cell survival, we manipulated its level of expression in various breast cancer cell lines (Fig. S1a, b). Cells were treated with mafosfamide, an analog of cyclophosphamide commonly used for breast cancer therapy that induces DNA breaks by nucleotide cross-linking [25]. In short-term cell viability assays, stable STING knockdown (KD) using two different shRNAs increased mafosfamide-sensitivity of MCF7 cells (Fig. S1c). Similar effects were observed with non-cancerous HEK293 cells harboring CRISPR-mediated STING knockout (KO) (Fig. S1d). In both cell lines, STING silencing also potentiated the effect of etoposide, a topoisomerase II inhibitor that stabilizes transient DNA double-strand breaks (DSBs) (Fig. S1e, f). Although of mild amplitude, the robustness of STING protective effects in these short-term assays called for further investigation.
Long-term survival assays monitor the ability of adherent cells to survive and resume proliferation after genotoxic treatment [18]. In both MCF7 (Fig. 1a) and BT20 (Fig. 1b) cell lines, STING KD significantly reduced the regrowth of mafosfamide-resistant clones by ∼2 fold. Conversely, transient STING overexpression markedly enhanced resistance to treatment (Fig. 1c). Since STING is a negative regulator of cell cycle progression [6, 26], these effects cannot result from STING-dependent alteration of cell proliferation. To complement these observations, we performed classical clonogenic assays [27]. The most striking observation was that STING silencing per se, i.e. in the absence of drug, dramatically hampered colony formation. Both colony size and number were reduced (Fig. 1d), indicating cytotoxic and cytostatic effects of STING KD. This effect was also observed at low doses of mafosfamide. Dramatic cell death occurred at 3 µM (Fig. 1d), suggesting that the trypsinization step prevented the recovery of many damaged cells compared to long-term survival assays (Fig. 1a). To confirm the intrinsic effect of STING silencing on cell survival, we transiently silenced STING in various estrogen receptor-positive (ER+) and triple negative (TN) breast cancer models including MCF7, BT20, HCC1937, and three HBCx (Human Breast Cancer xenograft) cells generated from breast cancer Patient-Derived Xenografts (PDX). Cell viability was significantly reduced in all but one cell line, and the amplitude of the effect paralleled siSTING efficiency (Fig. 1e).
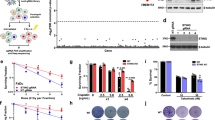
Long-term survival assays of MCF7 (a, c) and BT20 (b) cells showing the effect of STING KD (a, b) and STING overexpression (c) on survival and regrowth of adherent cells 20 or 40 days after exposure to mafosfamide (10 µM), as indicated. MCF7 cells were stably transduced with STING-targeting [shSTING] versus a non-targeting [shNT] shRNA prior to treatment (a, c). STING was overexpressed (shNT + STING, “STING”) or not (shNT + mock, “Mock”) (c) by the transient transfection of a STING-encoding plasmid the day of mafosfamide treatment. In BT20 cells (b), STING was silenced by siRNA 3 days before mafosfamide exposure. Representative wells are shown. Mean ± s.d. of n = 9 from three independent experiments, Student’s t-test. d Clonogenic assay showing the effect of STING KD on the ability of MCF7 cells to survive, adhere and regrow after exposure to a range of mafosfamide concentrations, as indicated. STING was silenced by siRNA before mafosfamide (or vehicle) exposure. Mean ± s.d. of n = 6 from three independent experiments, two-way ANOVA and post-hoc Sidak’s multiple comparisons test. e Viability of naïve MCF7 cells (mean ± s.d. of n = 9 from three independent experiments, Student’s t-test), of two immortalized and three PDX-derived breast cancer HBCx cells (mean ± s.d. of n = 6 from two independent experiments, Student’s t-test) 10 days after transfection with siSTING versus siNT. The molecular subtype (ER, TN) and the efficiency of siRNA to inhibit STING expression is indicated. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001. See also Fig. S1.
Together, these data demonstrate that STING promotes breast cancer cell survival in a cell-autonomous manner, under genotoxic stress but also at steady state.
STING is a positive regulator of the DDR
In vitro, the canonical STING pathway can be triggered in breast cancer cells by various genotoxic stresses, leading to autocrine/paracrine IFN signaling [18, 19]. In the absence of such stress, however, IFN expression was virtually undetectable (Fig. S2a and ref. [18]). This suggested that the marked alteration of cell viability induced by STING KD (Fig. 1d, e) involved a non-canonical STING-dependent mechanism. Preliminary experiments suggested that STING silencing impeded the formation of 53BP1 foci upon genotoxic stress [28]. The formation of 53BP1 foci at DSBs sites is an early hallmark of the DDR, a network of cellular pathways that sense, signal and repair DNA lesions [29]. This observation prompted us to investigate further the potential involvement of STING in the maintenance of genomic stability.
In naïve MCF7 (Fig. 2a) and HEK293 (Fig. S2b) cells, STING depletion reduced the formation of 53BP1 foci. Not only the average number of foci, but also their distribution per cell was altered as the ratio of cells harboring ≤1 foci was increased in STING-deficient cells at the expense of cells harboring ≥3 foci (Figs. 2b and S2b). STING overexpression reversed these effects (Fig. 2a, b). The distribution of 53BP1 foci was independent of the cell cycle as G2/S/M and G1 cells displayed virtually identical patterns (Fig. S2c). This observation suggests that these foci are not ‘53BP1 bodies’ shown to be largely confined to G1 cells [30]. Neither STING depletion nor overexpression impacted the spatial distribution of 53BP1 foci within the nucleus (Fig. S2d).
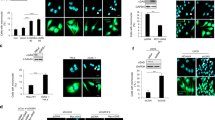
a Immunofluorescence and quantification of endogenous 53BP1 foci in MCF7 cells stably transduced with shNT or shSTING-2 and rescued or not for STING expression, as indicated. Nuclei were stained with DAPI. Mean ± s.d. of the average number of 53BP1 foci per cell from n = 3 independent experiments (one-way ANOVA and post-hoc Tukey’s multiple comparisons test). Between 500 and 1000 cells were counted per condition per experiment. b Distribution of the number of foci per cell in the various conditions described in a. Comet assays of untreated MCF7 and HEK 293 cells (c) and mafosfamide-treated MCF7 cells (d) including representative images of the most commonly observed comet tails in the indicated experimental conditions. MCF7 cells were stably transduced with shSTING-1 or -2 versus shNT. STING was overexpressed (STING) or not (mock) by the transient transfection of a STING-encoding plasmid, as indicated. Quantification of comet tail moment (length of the tail x percentage of DNA in the comet tail): in c, mean ± s.e.m of tail moment of n = 132 (MCF7-shNT), n = 118 (MCF7-shSTING-2) cells, n = 91 (HEK WT + mock), n = 89 (HEK-KO + mock) and n = 117 (HEK-KO + STING) cells (MCF7 cells: Student’s t-test, HEK293 cells: one-way ANOVA and post-hoc Tukey’s multiple comparison test); in d, mean ± s.e.m. of tail moment of n = 746 (shNT), n = 976 (shSTING-2), n = 1084 (shSTING-1), n = 776 (shNT + mock plasmid) and n = 605 (shSTING-2 + STING vector) cells from n = 3 independent experiments (one-way ANOVA and post-hoc Tukey’s multiple comparison test). See also Fig. S1 for relative STING expression levels in each cell line and Fig. S2.
In both MCF7 and HEK293 cells, STING deficiency increased the accumulation of genomic DNA damage as shown by the alkaline comet assay that accounts for both DSBs and single stranded DNA breaks (SSBs) (Fig. 2c). The stronger effect observed in HEK293 versus MCF7 cells may account for total versus partial STING silencing, respectively (Fig. S1a, b). The rescue of STING expression reversed this effect (Fig. 2c). The protective effect of STING on intrinsic DNA damage was maintained in genotoxic-treated MCF7 cells as STING KD increased the tail moment (irrespective of the shRNA used), and this effect was reversed by STING overexpression (Fig. 2d).
Together, these data demonstrate that STING contributes to genomic stability in steady state as well as under genotoxic stress conditions.
STING impact on DNA integrity and cancer cell survival is independent of its canonical pro-inflammatory pathway
To further assess the independence of these effects from the canonical cGAS/STING inflammatory pathway, we performed a series of additional experiments. First, we showed that STING and/or cGAS silencing had no effect on basal levels of IFN expression and STAT1 expression and phosphorylation (Figs. S2a and 3a), confirming that this pathway is not activated in naïve MCF7 and HEK293 cells. Second, we showed that HEK293 cells lacked detectable cGAS expression (Fig. 3a). Third, we activated the cGAS/STING/TBK1/IRF-3/IFN pathway in MCF7 cells using cGAMP, a typical STING agonist [4]. While the treatment triggered IFN signaling as reflected by the upregulation of typical ISGs (Fig. 3b), it had no impact on 53BP1 foci formation (Fig. 3c). Fourth, in contrast to STING KD (Fig. 2a), TBK1 silencing (Fig. 3d) did not impair the formation of 53BP1 foci, that was even slightly increased (Fig. 3e). Fifth, a naturally occurring C-terminally truncated STING isoform has been shown to act as a dominant-negative (DN) of full-length STING on IFN induction due to its inability to interact with TBK1/IRF3 complex [31]. Expression of STING-DN in a shRNA-mediated-STING-deficient background was sufficient to protect MCF7 cells from DNA damage accumulation as revealed by the comet assay (Fig. 3f). Sixth, as opposed to STING silencing (Fig. 1), IFN receptor (IFNAR1) silencing (Fig. 3g) had no effect on MCF7 cell survival in steady-state and on its sensitivity to the DNA-damaging agent mafosfamide, i.e. when IFN production is triggered [18] (Fig. 3h).
a Immunoblot of phosphoSer-STAT1, total STAT1 and cGAS in MCF7 and HEK293 cells silenced or not for cGAS. STING was KD by siRNA (MCF7) or KO by CRISPR-Cas9 (HEK293) b RT-qPCR analysis of IFNβ, IFIT1, and OAS2 expression in MCF7 cells 6 h after exposure to 50 µg/mL of the CDN cGAMP. Mean ± s.d. of n = 3 independent experiments, Student’s t-test. c Effect of cGAMP treatment on the formation of 53BP1 foci in MCF7 cells as determined by immunofluorescence. Mean ± s.e.m. of the number of foci per cell in n = 3134 (vehicle) and n = 3204 (agonist) cells from n = 3 independent experiments (Student’s t-test). d RT-qPCR analysis of endogenous TBK1 in MCF7 cells transfected with siTBK1 versus siNT (Student’s t-test, n = 2 independent experiments). e Effect of TBK1 silencing on the formation of 53BP1 foci in MCF7 cells as determined by immunofluorescence. Mean ± s.e.m. of the number of foci per cell in n = 2640 (shNT) and n = 2340 (siTBK1) cells from n = 3 independent experiments (Student’s t-test). f Tail moment of MCF7 stably transduced with shNT or shSTING-2 transiently transfected or not with a shRNA-resistant STING-DN expression vector. Mean ± s.e.m. of tail moment of n = 746 (shNT), n = 976 (shSTING-2), and n = 871 (shSTING-2 + STING-DN plasmid, STING-DN) cells from n = 3 independent experiments (one-way ANOVA and post-hoc Tukey’s multiple comparison test). g RT-qPCR analysis of endogenous IFNAR1 expression in MFC7 cells transfected with siIFNAR1 versus a siNT (Student’s t-test, n = 2 independent experiments). h Viability of MCF7 cells 10 days after transfection with siSTING or siIFNAR1 versus siNT exposed (right) or not (left) to mafosfamide (10 µM) 3 days after transfection. Mean ± s.d. of n = 9 from three independent experiments, two-way ANOVA and post-hoc Dunnett’s multiple comparison test.
Together, these data demonstrate that STING promotes the DDR and cancer cell survival in a cell-autonomous and cGAS/STING/IFN-independent manner.
STING partly resides in the nucleus of breast cancer cells
The canonical inflammatory pathway-independence of STING contribution to DNA integrity raised the question of the alternative mechanism involved. Preliminary cell fractionation experiments showed that STING partially resided in the nuclear fraction of MCF7 cells, irrespective of mafosfamide treatment [18]. Interestingly, the cytosolic DNA sensor cGAS has been recently shown to translocate to the nucleus upon DNA damage where it contributes to the DDR [32]. Conversely, several nuclear proteins involved in the DDR (e.g. DNA-PKcs, Ku70, MRE11) have been shown to act as cytosolic DNA sensor to activate inflammatory responses [33,34,35,36]. The dual subcellular localization and function of these factors called for investigating further the nuclear localization of STING in various breast cancer models.
STING protein expression was detected in all HBCx models we tested (Fig. 4a) at a level that was much higher than in MCF7 cells (Figs. 4b and S1b), but nevertheless variable among samples, as previously reported for colorectal adenocarinoma [37]. STING was detected in the nuclear fraction of the three cell lines analyzed (Fig. 4c,f). To circumvent the sensitivity and specificity issues previously encountered in immunofluorescence experiments using commercial anti-STING antibodies [28], we generated tagged STING constructs containing C-terminal hemagglutinin (HA) tag with/without N-terminal Flag tag. All constructs, including untagged STING, displayed similar fractionation profile as endogenous STING (Fig. 4c, left) when ectopically expressed in MCF7 cells (Fig. 4d) and in STING-KO HEK293 cells (Fig. 4e). Overall, the ratio of STING present in the nuclear fraction ranged from 28 to 65% (Fig. 4f) and this was not correlated to the total amount of STING expressed, suggesting cell-type-specific regulatory mechanisms. Of note, MCF7 cells expressing ectopic Flag-STING-HA exhibited similar nuclear/cytoplasmic ratio as HBCx-39 cells expressing STING endogenously (Fig. 4f). In these STING immunoblot experiments, a band of higher electrophoretic mobility was detected by the various antibodies in some, but not all experiments. No consistent correlation could be made with the expression level of STING (e.g. HBCx-14 vs HBCx-19 in Fig. 4a), the type of construct (Fig. 4e) or the subcellular fraction (e.g. HBCx-3 nuclear vs cytoplasmic fractions, Fig. 4b). Its biochemical nature is uncertain.
a Immunoblot of endogenous STING in various PDX-derived breast cancer cell lines (named HBCx). b Immunoblot of endogenous STING and γH2AX in HBCx-3 and HBCx-39 versus MCF7 cells 48 h after treatment with (+) or without (−) mafosfamide (10 µM). Biochemical fractionation showing the amounts of endogenous (c) and ectopically expressed (d, e) STING present in the cytoplasmic (C) versus nuclear (N) fractions from various cell lines, as indicated. Lamin A/C was used as a nuclear marker and the ER-resident Protein Disulfide Isomerase (PDI) as the cytoplasmic marker. For MCF7 cell samples, the time of anti-STING blot exposure was adjusted to the expression level of STING, as indicated. In d, STING constructs were stably expressed in parental MCF7 cells. In e, the different tagged STING constructs were transiently expressed in STING-KO HEK293 cells. f Quantification of the nuclear (gray)/cytoplasmic (black) amounts of STING in cell lines expressing different amount of STING (normalized to that of parental MCF7 cells). The numbers represent the percentage of nuclear STING for each cell line.
Together, these data show that STING intrinsically resides in the nuclear fraction of various breast cancer cell models.
STING co-localizes with the lamina in breast cancer cells
We further characterized STING subcellular localization using immunofluorescence. As expected [38], STING was uniformly spread within the cytoplasm of MCF7 cells and co-localized with ER (calnexin) and Golgi (GM130) markers (Fig. 5a). ImageJ treatment of immunofluorescence images was performed to better display co-localization (appearing in white). Strikingly, STING also co-localized with lamin B1, a component of the nuclear lamina (Fig. 5a). The lamina is a fibrillary network underlying the INM that serves as anchoring point for INM proteins, chromatin, and transcription factors [39]. To strengthen this finding, we performed a pre-fixation ribonuclease- and detergent-based cell extraction that preferentially retains cytoskeleton, nuclear matrix, and chromatin, at the expense of soluble/loose structures [40, 41] (Fig. S3). After such a treatment, STING strongly co-localized with lamin B1 (Figs. 5b and S3, right) and lamin A/C (Fig. S3, right) at the nuclear rim (white arrowheads) and occasionally at small intra-nuclear structures (white arrows) presumably corresponding to nuclear membrane invaginations.
a Immunofluorescence of Flag-STING-HA transiently expressed in MCF7 cells (versus empty vector, mock), using anti-HA or anti-Flag antibodies (according to the species of the antibody directed against subcellular markers) as indicated (upper panels). Middle panels show immunofluorescence of ER (calnexin), Golgi (GM130), and nuclear lamina (lamin B1) markers. Lower panels display merged images, processed or not by Image J software to emphasize co-localization (appearing in white), as indicated. b Immunofluorescence experiment performed after pre-extraction of MCF7 cells stably transfected with untagged STING (versus mock vector), using anti-STING and anti-lamin B1 antibodies. Symbols: white arrowheads point to co-localization of STING at the lamina rim, and white arrows to intra-nuclear staining. Nuclei were stained with DAPI. See also Fig. S3.
Together, these data demonstrate that in breast cancer cells, the nuclear fraction of the STING pool co-localizes with the lamina at the nuclear periphery.
Identification of STING at the INM by electronic microscopy
Considering the transmembrane nature of STING, its co-localization with the lamina, and the resistance of INM proteins to pre-extraction [42], we hypothesized that nuclear STING localizes at the INM. To explore this further, we monitored STING localization by immunogold labeling in immunoelectron microscopy (EM) (Fig. 6a). As shown in Fig. 6b–f (see Fig. S4a for negative controls), black dots corresponding to anti-Flag-bound gold particles were observed at cytoplasmic vesicle structures and, in agreement with STING/calnexin co-localization, at perinuclear ER membranes. Consistent with immunofluorescence observations, STING was also detected at the periphery of the nucleus, mainly at proximity of the INM at a distance compatible with immunogold staining of a transmembrane protein (Fig. 6c–f). Furthermore, STING was frequently observed at both sides of nuclear membrane invaginations (Fig. 6f) and sometimes appeared as gold dot doublets (Fig. 6e, f) presumably reflecting distinct quaternary structures [43].
a Schematic representation of the anti-Flag immunogold staining procedure of MCF7 cells stably expressing the Flag-STING-HA construct. b–f Five representative immunoelectron microscopy images illustrating typical features of STING subcellular localization identified using symbols displayed in the bottom left box. Negative control involved parental cells (Fig. S3a). Size bars are indicated in each panel. See Fig. S4 for negative controls (non-transfected cells).
Cells analyzed 48 h after mafosfamide treatment showed several signs of stress including nuclei with irregular shape and large invaginations, picnotic nuclei, dilated ER with dramatically enlarged lumen and large vesicles filled with cell debris (Fig. S4b, bottom panels). As reported [44], many cytoplasmic vesicles were positive for STING (Fig. S4b, upper left panel). The various localizations of STING described above for naïve cells were also observed in mafosfamide-treated cells (Fig. S4b, upper panels).
Together, these data demonstrate that in breast cancer cells, the nuclear STING pool mainly resides at the nucleus periphery and more specifically at the INM.
Determination of nuclear STING interactome using mass spectrometry
We next performed an interactomics analysis to determine the nuclear STING protein network. First, we confirmed that immunoprecipitates were significantly enriched for STING and nuclear proteins (Fig. S5a, b and Table S1). Remarkably, none of the canonical STING interactors (e.g. TBK1, IRF3, MAVS, STAT6, TRAF6) [2, 3, 8, 45] could be identified in nuclear STING interactome (Table S1). Functional analysis (STRING database) of the nuclear proteins specifically immunoprecipitated by STING revealed the enrichment of three functional networks: mRNA splicing, eukaryotic translation elongation, and DNA-repair (Fig. 7a). The latter contains many proteins involved in chromatin remodeling complexes that have been shown to facilitate the efficacy of DNA damage signaling and/or repair, e.g. SMARCA5 [46], ACTL6A [47], SUPT16H [48], RUVBL1/2 [49], SMC3 [50], and HMGB2 [51] (Fig. 7a, b). The most striking observation was the identification of the three core proteins forming the DNA-dependent protein kinase (DNA-PK) complex as part of STING interactome: DNA-PK catalytic subunit (DNA-PKcs), Ku70 (aka XRCC6), and Ku80 (aka XRCC5) (Fig. 7a,b). DNA-PK, together with ATM and ATR, is a master regulator of the DDR [52].
a, b Interactomics analysis of nuclear STING. Flag-STING-HA was immunoprecipitated using anti-Flag antibodies from stably transfected MCF7 cell nuclear extracts. a Functional networks of nuclear STING interactome using STRING database (interaction score high evidence = 0,700). Proteins involved in mRNA splicing (FDR = 1.22e−18) and Eukaryotic Translation Elongation (FDR = 0.00025) pathways, as per Reactome database, are highlighted in blue and green, respectively. Proteins involved in DNA repair according to GO terms (FDR = 0.00016) are highlighted in red. b Volcano plot of –log(p value) versus fold change (expressed in log2) of proteins present in anti-STING immunoprecipitates versus negative control as detected by mass spectrometry. Proteins previously reported to be involved in DNA repair (Gene Ontology Cell Component database) are colored in magenta. c Immunoblots (IB) of DNA-PKcs in immunoprecipitates recovered using three antibodies mapping distinct regions of Flag-STING-HA stably expressed in MCF7 cells. Immunoblots (IB) of proteins constituting the DNA-PK complex (DNA-PKcs, Ku80, Ku70) in immunopecipitates of untagged STING stably expressed in MCF7 cells (d) and of endogenous STING expressed in HBCx-3 (e) and HEK293 (f). d, f Show the reverse co-immunoprecipitations using anti-DNA-PKcs antibodies. In c–f, the negative control involved beads only. g Immunoblot of proteins constituting the DNA-PK complex (DNA-PKcs, Ku80, Ku70) in the chromatin-nuclear matrix fraction (lanes 1–4) versus whole cell lysates (lanes 5–8) of MCF7 cells stably overexpressing untagged STING or not (mock) and treated (+) or not (−) with mafosfamide (10 µM) for 48 h. Lamin B1 is used as a loading control. See also Fig. S5.
The STING/DNA-PK interaction was confirmed by the enrichment of DNA-PK subunits in STING immunoprecipitates performed using various antibodies and nuclear extracts involving tagged and untagged STING ectopically (Fig. 7c, d) or endogenously (Fig. 7e, f) expressed in MCF7, HBCx-3, and HEK293 cells. Reciprocal co-immunoprecipitation was less efficient, possibly due to epitope overlap, but nevertheless showed consistent enrichment of STING (1.7-fold) in DNA-PKcs immunoprecipitates from MCF7-STING and HEK293 cells (Fig. 7d, f). TBK1 silencing did not impair STING/DNA-PK interaction (Fig. S5c), further arguing that the nuclear STING DDR pathway is distinct from the canonical cytoplasmic inflammatory pathway.
To address whether STING impacts DNA-PK complex subcellular localization, we performed a pre-extraction that enriches chromatin, nuclear matrix (e.g. Lamin B1) and cytoskeleton [40,41,42, 53]. As shown in Fig. 7g, STING overexpression in untreated cells markedly enhanced the amount of pre-extraction resistant DNA-PK complex proteins (lane 2 vs 1) without affecting their expression at the cellular level (lane 6 vs 5). The presence of DNA-PK complex in this chromatin/nuclear matrix fraction was increased upon mafosfamide treatment, and this was further enhanced in the context of STING overexpression (lane 4).
Together, these data demonstrate that nuclear STING exhibits a specific interactome enriched in various DDR regulators among which DNA-PK appears as a promising candidate to support a role of nuclear STING in regulating DNA stability.
Clinical relevance of STING co-localization with the lamina
To address the clinical relevance of these findings, we investigated the subcellular localization of STING in malignant breast tumors. We first validated the reliability of our STING immunostaining settings by analyzing 4 breast cancer PDXs exhibiting different tissue levels of STING mRNA expression as determined by RT-qPCR (data not shown). Accordingly, different levels of STING protein immunostaining were detected (Fig. S6a). In high STING-expressing samples (HBCx-15 and HBCx-19), STING was detected in virtually all tumor cells and a clear co-localization of STING with lamin B1 was observed at the nuclear rim (Figs. 8a and S6b). A few lamin-negative (i.e. most likely non-tumoral) cells exhibiting strong STING staining were also detected. These cells were the only STING-positive cells detected in low/medium STING-expressing samples (e.g. HBCx-24).
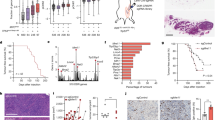
a Immunofluorescence of endogenous STING and lamin B1 in the high STING-expressing HBCx-19 PDX. Right panels display merged images without or with image treatment by Image J software to emphasize co-localization (appearing in white). b The left image shows one representative area of a patient tumor immunostained for STING and lamin B1. The four right panels show higher magnification of the squared area for STING, lamin B1 and merged staining without and with ImageJ treatment, as indicated. Nuclei were stained with DAPI. White arrowheads: examples of STING/lamin B1 co-localization; green arrowheads identify the few STING-positive/lamin-negative cells; orange arrows: cells in mitosis. c High STING levels correlate with resistance to chemotherapy in breast cancer. Kaplan–Meier plotter analysis of RFS in function of STING mRNA expression for all cases (left panel) and all patients receiving chemotherapy (right panel). d Same analysis for ovarian cancer who all received adjuvant chemotherapy. See also Fig. S6.
Then, we analyzed samples from 6 TN breast cancers resistant to neoadjuvant treatment containing cyclophosphamide (representative examples are shown in Figs. 8b and S6c). STING was detected in all tumor samples analyzed, and in each sample, the staining for STING was present in most, if not all, tumor cells, albeit with various intensities. Again, the STING/lamin B1 co-localization was observed at the nuclear rim of tumor cells (Figs. 8b and S6c) including in cells that were in mitosis (Fig. 8b).
High STING expression is associated with increased risk of relapse in patients receiving adjuvant chemotherapy
Finally, we interrogated the Kaplan–Meier Plotter database, which contains gene expression and clinical data for more than 4000 breast cancer patients, and looked at STING expression level association with patient relapse-free survival (RFS). In agreement with the well-known role of STING in immunosurveillance, STING expression in the whole patient cohort was positively correlated to RFS when all breast cancer cases were considered (Fig. 8c, left, p = 1.e−16). However, breast cancer is a heterogeneous disease, and patients are enrolled at diagnosis in different clinical protocols that combine surgery and treatments, including chemotherapy. When the analysis of STING expression was run only with data from patients that received adjuvant chemotherapy after surgery, an inverse correlation was observed (Fig. 8c, right). It was close to significance when all molecular and histological subtypes were pooled (p = 0.057), and more marked for HER2+ (p = 0.015), TN (p = 0.012) and more generally ER− (p = 0.018) tumors (Fig. S6d). Of interest, an even more significant negative correlation with progression-free survival was found in a cohort of ovarian cancer patients who all received chemotherapy after surgery (p = 0.00039; Fig. 8d).
Discussion
STING is broadly considered as an antitumor protein that activates IFN signaling driving cancer immunosurveillance. Several studies have reported that IFN signaling is frequently suppressed in cancer cell lines [6, 37, 54, 55]. This has led to the widespread notion that STING itself is frequently lost in cancer. Yet, data in this regard are conflicting since studies have reported increased or decreased tumoral STING expression depending on cancer types [56,57,58,59]. Genomic data also argue against massive STING loss in cancer and analyses of STING copy number variation in tumors indicate amplifications rather than deletions [55, 60]. In fact, the apparent lack of STING signaling in cancer cells may reflect the deletion of IFN genes and/or the activation of IFN signaling inhibitor pathways [60,61,62,63]. In line with this notion, STING expression was detected in all breast cancer cell models and tumor samples investigated in this study, albeit at variable levels.
While former studies have shown that STING inflammatory pathway is triggered by genomic instability [18, 19], this is to our knowledge the first report demonstrating that STING also protects cells from genomic instability in a manner that is independent of cGAS and IFN responses. Wu and colleagues recently reported that STING promotes DDR in genotoxic-treated cells [16]. However, this function was mediated by the activation of the cGAS/STING/IFN pathway and the subsequent expression of ISGs, including several PARP family genes. Interestingly, three independent studies recently reported that cGAS regulates genomic stability in an IFN-independent manner [32, 64, 65]. The first one showed that cGAS promoted DNA compaction in a higher-ordered state resistant to RAD51 strand invasion [64]. The second one reported that cGAS stabilized replication forks [65]. Finally, the third study showed that cGAS interacted with PARP1 impeding the PARP1-Timeless complex formation necessary to HR completion [32]. When investigated [64, 65], these new cGAS functions were shown to be independent of STING. Together with our observations, these data strongly suggest that the regulation of genomic stability by cGAS and STING involves distinct pathways.
Our proteomic analysis revealed that nuclear STING interactome contains several proteins involved in DSB DNA repair (Fig. 7b). The most remarkable finding was that nuclear STING interacts with the DNA-PK complex. Although the functional pleiotropy of DNA-PK is emerging [66, 67], it is mostly known for its role in NHEJ activation and initiation [68]. We here showed that STING overexpression enhanced the association of DNA-PK complex proteins with the chromatin/nuclear matrix fraction without affecting their level of expression. Although concomitant increase of DNA-PK activity on the chromatin was not experimentally assessed, these observations call for investigating further whether STING could regulate the DDR by favoring the formation of NHEJ-initiation complex at DNA damage sites. Dedicated structure-function studies will be also needed to fully elucidate how these proteins cooperate in the nucleus and at which level(s) STING functionally impacts the DDR (detection, signaling, and/or repair).
This study also shows that a part of the cellular STING pool intrinsically resides in the nucleus of breast cancer cells. In their seminal work, Schirmer and colleagues identified STING as a Nuclear Envelope Transmembrane protein (NET23) in rat liver and tentatively localized it to the outer nuclear membrane (ONM) using high resolution fluorescence imaging [69, 70]. This data does not oppose ours as protein addressing to the INM has been proposed to involve initial protein insertion into ER membranes followed by diffusion to the contiguous ONM and INM (reviewed in ref. [71]). Furthermore, these authors showed that STING failed to localize to the nuclear envelope in lamin A/C-deficient cells [42], which is typically observed for acknowledged INM-resident proteins such as Emerin [72]. Interestingly, there is emerging evidence that the nuclear envelope is involved in DNA repair [73]. In yeast [74] and drosophila [75], persistent or hard-to-repair DSBs relocate to the nuclear periphery where they are anchored to nuclear pore or INM proteins to be repaired. In yeast and mammals, DSBs in chromatin domains associated with the nuclear envelope and nuclear pores are preferentially repaired by error-prone pathways, such as NHEJ, alt-NHEJ, and Break-Induced Replication [76, 77]. In our system, we detected an impact of STING on the overall formation of 53BP1 foci, but not on their repartition within the nucleus. Additional studies are required to fully elucidate whether STING function is confined to the nuclear periphery or extends to the entire nucleus, and by which mechanism.
According to its role as a promoter of antitumor immunity, high intra-tumoral levels of STING have been correlated with good prognosis in various cancers [58, 59]. This was here confirmed for breast cancer (Fig. 8c). In contrast, when we focused on breast (and ovarian) cancer patients that received adjuvant chemotherapy after surgery, high STING expression was correlated with increased recurrence. In samples of chemotherapy-resistant breast tumors, STING levels (and co-localization with lamina) were particularly elevated in proliferating cells that drive tumor regrowth (Fig. 8b). This is consistent with long-term survival in vitro assays showing that increased STING expression promoted survival and regrowth of breast cancer cells exposed to genotoxic stress (Fig. 1c). These observations suggest that, in the context of high chromosomal instability (e.g. chemotherapy), the pro-tumoral DDR-related effects of STING may prevail over its protective role through immunosurveillance. Exploring the balance between these opposite actions of STING in cancer is crucial and requires the development of appropriate animal models.
In conclusion, we uncovered a new subcellular localization of STING at the INM of breast cancer cells and provided evidence supporting its involvement in the DDR. This newly identified function highlights a cell-autonomous pathway by which STING promotes cancer cell survival and resistance to DNA-damaging agents. Importantly, the effects of STING on the DDR, cell survival and drug resistance were independent of cGAS and of IFN responses. Our study pioneers a new field of investigation to elucidate the precise role of STING in the DDR, with potential impact in the clinical setting.
Materials and methods
Antibodies, plasmids, and chemicals
All reagents are described in Supplementary Methods.
Cell lines
MCF7 (Sigma Aldrich, Saint-Louis, Missouri, USA), BT20 (ATCC, Manassas, Virginie, USA), HCC1937 (ATCC), HEK293 STING-WT and KO (Invivogen, San Diego, California, USA) were authenticated by STR profiling. Cells were maintained in DMEM/F12 supplemented with 10% heat-inactivated FBS and 1% penicillin-streptomycin. Cell transfection and transduction procedures are described in Supplementary Methods. Human breast cancer PDX-derived cell lines (HBCx-3 and HBCx-19, ER+; HBCx-2 and HBCx-39, TN) were generated as previously described [18].
PDXs and patient tumor samples
Formalin-fixed, paraffin-embedded (FFPE) breast cancer PDX samples were retrieved from the archives of Xentech (Evry, France). PDX establishments, and care and use of animals, were performed as previously described [78, 79] after approval of the Ethics Committees of the Institut Curie and CEEA-Ile de France Paris (official registration number 59). FFPE breast tumor samples were retrieved from the archives of the Department of Pathology of Centre Jean Perrin (Clermont-Ferrand, France). Only samples of breast tumors resistant to neoadjuvant chemotherapy (Fluorouracil-Epirubicin, Cyclophosphamide (FEC)-Taxane regimen), larger than 2 cm in diameter, were used with informed patient consent. This study was approved by the Ethics Committee (CECIC) of the Rhone-Alpes-Auvergne region (Grenoble, France).
Molecular studies
RT-qPCR, co-immunoprecipitations, and immunoblotting were performed following routine procedures. Cell fractionation [80] and chromatin/nuclear matrix enrichment [53] were performed as previously described, with minor modifications. All details are provided in Supplementary Methods.
NanoLC-MS/MS proteomic analyses
Detailed methodology for protein identification, quantification, and data processing is provided in Supplementary Methods.
Immunofluorescence
Immunofluorescence assays [18] and pre-extraction steps [40, 41] were performed as previously described (See Supplementary Methods for details). For co-localization experiments, merged images were treated by Image J software to emphasize (in white) the results of the Manders coefficient with threshold as explained earlier [81]. 53BP1 Foci were quantified using FIJI [82]. For foci distribution within the nucleus, a script was written to segment nuclei in 5 layers from the nuclear periphery to the nuclear center (i.e., 4 concentric layers of 5 pixels and the remaining center), by using the deep learning plugin STARDIST [83] and the Region Of Interest (ROI) erode function of FIJI.
Electron microscopy
MCF7 cells stably expressing Flag-STING-HA were collected in 1.5 mL Eppendorf tube, then fixed, permeabilized, blocked and incubated with anti-Flag antibody as for immunofluorescence analyses. Cells were then incubated 1 h at RT with nanogold-coupled secondary antibody and finally fixed with 2.5% glutaraldehyde EM grade (Sigma, #16210) for 1 h. Further sample processing and data acquisition are provided in Supplementary Methods.
Comet assay
Naïve HEK293 cells were analyzed 3 days post-transfection with mock or STING plasmids. MCF7 cells were analyzed 6 days post-mafosfamide treatment (10 µM) which corresponds to maximal γH2AX accumulation [18]. When indicated, cells were transfected with expression plasmids 8 h prior treatment. Alkaline comet assays were performed according to the manufacturer’s instructions (Trevigen, Gräfelfing, Germany). DNA damage was measured in terms of tail moments using OpenComet plugin in ImageJ software.
Cell viability assay
Cells were treated with vehicle, mafosfamide (10 µM) or etoposide (5 µM or dose-response) 1-day post-seeding. In the siRNA/genotoxic combination setting, cells were transfected 1-day post-seeding and treated with mafosfamide 3 days later. Cell viability was measured between day 3 and day 10 (as indicated) using CellTiter-Glo Luminescent cell viability assay reagent (Promega, Madison, Wisconsin, USA).
Long-term survival assay
This assay was processed as previously described [18]. Briefly, MCF7 cells were treated with 10 µM mafosfamide 1-day post-seeding. BT20 cells were transfected with siRNAs 1-day post-seeding and treated with 10 µM mafosfamide 3 days later. When indicated, cells were transfected with expression plasmids the day of treatment. After 20 (overexpression) or 40 (KD) days, colonies were stained and quantified using the plugin ColonyArea in ImageJ software [84].
Clonogenic assay
Cells were transfected with siRNAs 1-day post-seeding and treated or not with three doses of mafosfamide 2 days later. Two days post-treatment, cells were trypsinized, re-plated in 6-well plates at a density of 1000 cell/well and allowed to grow and form colonies for 2 additional weeks [27]. The number and size of colonies were quantified using imageJ “Analyze particles” plugin.
Kaplan–Meier survival analysis
The RFS curves associated to STING (TMEM173) expression status (high versus low using the auto-select best cutoff) were generated according to molecular subtypes of breast cancer and/or chemotherapy as documented in the Kaplan–Meier Plotter integrated database (http://kmplot.com/) [85].
Statistical analysis
Data were analyzed using GraphPad Prism 6 software (GraphPad, San Diego, California, USA) and are presented as mean ± s.d. unless otherwise indicated. Comparisons between two groups or multiple groups were performed by Student’s t-test or one-way ANOVA, respectively. To examine the influence of two independent parameters on multiple groups, two-way ANOVA was used. Statistically significant differences are indicated as follows: ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001; ∗∗∗∗p < 0.0001. Statistical details of experiments can be found in the figure legends.
References
Ishikawa H, Ma Z, Barber GN. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature. 2009;461:788–92.
Ishikawa H, Barber GN. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature. 2008;455:674–8.
Liu S, Cai X, Wu J, Cong Q, Chen X, Li T, et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science. 2015;347:aaa2630.
Ablasser A, Goldeck M, Cavlar T, Deimling T, Witte G, Rohl I, et al. cGAS produces a 2’-5’-linked cyclic dinucleotide second messenger that activates STING. Nature. 2013;498:380–4.
Abe T, Barber GN. Cytosolic-DNA-mediated, STING-dependent proinflammatory gene induction necessitates canonical NF-kappaB activation through TBK1. J Virol. 2014;88:5328–41.
Ranoa DRE, Widau RC, Mallon S, Parekh AD, Nicolae CM, Huang X, et al. STING promotes homeostasis via regulation of cell proliferation and chromosomal stability. Cancer Res. 2019;79:1465–79.
Bakhoum SF, Ngo B, Laughney AM, Cavallo JA, Murphy CJ, Ly P, et al. Chromosomal instability drives metastasis through a cytosolic DNA response. Nature. 2018;553:467–72.
Dunphy G, Flannery SM, Almine JF, Connolly DJ, Paulus C, Jonsson KL, et al. Non-canonical activation of the DNA sensing adaptor STING by ATM and IFI16 mediates NF-kappaB signaling after nuclear DNA damage. Mol Cell. 2018;71:745–60 e5.
Barber GN. STING-dependent cytosolic DNA sensing pathways. Trends Immunol. 2014;35:88–93.
Deng L, Liang H, Xu M, Yang X, Burnette B, Arina A, et al. STING-dependent cytosolic DNA sensing promotes radiation-induced type I interferon-dependent antitumor immunity in immunogenic tumors. Immunity. 2014;41:843–52.
Vanpouille-Box C, Alard A, Aryankalayil MJ, Sarfraz Y, Diamond JM, Schneider RJ, et al. DNA exonuclease Trex1 regulates radiotherapy-induced tumour immunogenicity. Nat Commun. 2017;8:15618.
Wang H, Hu S, Chen X, Shi H, Chen C, Sun L, et al. cGAS is essential for the antitumor effect of immune checkpoint blockade. Proc Natl Acad Sci USA. 2017;114:1637–42.
Harding SM, Benci JL, Irianto J, Discher DE, Minn AJ, Greenberg RA. Mitotic progression following DNA damage enables pattern recognition within micronuclei. Nature. 2017;548:466–70.
Brown JS, Sundar R, Lopez J. Combining DNA damaging therapeutics with immunotherapy: more haste, less speed. Br J Cancer. 2018;118:312–24.
Ahn J, Xia T, Konno H, Konno K, Ruiz P, Barber GN. Inflammation-driven carcinogenesis is mediated through STING. Nat Commun. 2014;5:5166.
Wu Z, Oeck S, West AP, Mangalhara KC, Sainz AG, Newman LE, et al. Mitochondrial DNA stress signalling protects the nuclear genome. Nat Metab. 2019;1:1209–18.
Chen Q, Boire A, Jin X, Valiente M, Er EE, Lopez-Soto A, et al. Carcinoma-astrocyte gap junctions promote brain metastasis by cGAMP transfer. Nature. 2016;533:493–8.
Gaston J, Cheradame L, Yvonnet V, Deas O, Poupon MF, Judde JG, et al. Intracellular STING inactivation sensitizes breast cancer cells to genotoxic agents. Oncotarget. 2016;7:77205–24.
Li T, Chen ZJ. The cGAS-cGAMP-STING pathway connects DNA damage to inflammation, senescence, and cancer. J Exp Med. 2018;215:1287–99.
Erdal E, Haider S, Rehwinkel J, Harris AL, McHugh PJ. A prosurvival DNA damage-induced cytoplasmic interferon response is mediated by end resection factors and is limited by Trex1. Genes Dev. 2017;31:353–69.
Parkes EE, Walker SM, Taggart LE, McCabe N, Knight LA, Wilkinson R, et al. Activation of STING-dependent innate immune signaling by S-phase-specific DNA damage in breast cancer. J Natl Cancer Inst. 2017;109:djw199.
Mackenzie KJ, Carroll P, Martin CA, Murina O, Fluteau A, Simpson DJ, et al. cGAS surveillance of micronuclei links genome instability to innate immunity. Nature 2017;548:461–5.
Lan YY, Londono D, Bouley R, Rooney MS, Hacohen N. Dnase2a deficiency uncovers lysosomal clearance of damaged nuclear DNA via autophagy. Cell Rep. 2014;9:180–92.
Hartlova A, Erttmann SF, Raffi FA, Schmalz AM, Resch U, Anugula S, et al. DNA damage primes the type I interferon system via the cytosolic DNA sensor STING to promote anti-microbial innate immunity. Immunity. 2015;42:332–43.
Mazur L, Opydo-Chanek M, Stojak M, Wojcieszek K. Mafosfamide as a new anticancer agent: preclinical investigations and clinical trials. Anticancer Res. 2012;32:2783–9.
Basit A, Cho MG, Kim EY, Kwon D, Kang SJ, Lee JH. The cGAS/STING/TBK1/IRF3 innate immunity pathway maintains chromosomal stability through regulation of p21 levels. Exp Mol Med. 2020;52:643–57.
Franken NA, Rodermond HM, Stap J, Haveman J, van Bree C. Clonogenic assay of cells in vitro. Nat Protoc. 2006;1:2315–9.
Gaston J, Cheradame L, Yvonnet V, Deas O, Poupon MF, Judde JG, et al. Correction: intracellular STING inactivation sensitizes breast cancer cells to genotoxic agents. Oncotarget. 2019;10:4249–51.
Schultz LB, Chehab NH, Malikzay A, Halazonetis TD. p53 binding protein 1 (53BP1) is an early participant in the cellular response to DNA double-strand breaks. J Cell Biol. 2000;151:1381–90.
Lukas C, Savic V, Bekker-Jensen S, Doil C, Neumann B, Pedersen RS, et al. 53BP1 nuclear bodies form around DNA lesions generated by mitotic transmission of chromosomes under replication stress. Nat Cell Biol. 2011;13:243–53.
Chen H, Pei R, Zhu W, Zeng R, Wang Y, Wang Y, et al. An alternative splicing isoform of MITA antagonizes MITA-mediated induction of type I IFNs. J Immunol. 2014;192:1162–70.
Liu H, Zhang H, Wu X, Ma D, Wu J, Wang L, et al. Nuclear cGAS suppresses DNA repair and promotes tumorigenesis. Nature. 2018;563:131–6.
Burleigh K, Maltbaek JH, Cambier S, Green R, Gale M, Jr, James RC, et al. Human DNA-PK activates a STING-independent DNA sensing pathway. Sci Immunol. 2020;5:eaba4219.
Ferguson BJ, Mansur DS, Peters NE, Ren H, Smith GL. DNA-PK is a DNA sensor for IRF-3-dependent innate immunity. Elife 2012;1:e00047.
Kondo T, Kobayashi J, Saitoh T, Maruyama K, Ishii KJ, Barber GN, et al. DNA damage sensor MRE11 recognizes cytosolic double-stranded DNA and induces type I interferon by regulating STING trafficking. Proc Natl Acad Sci USA. 2013;110:2969–74.
Sui H, Zhou M, Imamichi H, Jiao X, Sherman BT, Lane HC, et al. STING is an essential mediator of the Ku70-mediated production of IFN-lambda1 in response to exogenous DNA. Sci Signal. 2017;10:eaah5054.
Xia T, Konno H, Ahn J, Barber GN. Deregulation of STING signaling in colorectal carcinoma constrains DNA damage responses and correlates with tumorigenesis. Cell Rep. 2016;14:282–97.
Saitoh T, Fujita N, Hayashi T, Takahara K, Satoh T, Lee H, et al. Atg9a controls dsDNA-driven dynamic translocation of STING and the innate immune response. Proc Natl Acad Sci USA. 2009;106:20842–6.
Dobrzynska A, Gonzalo S, Shanahan C, Askjaer P. The nuclear lamina in health and disease. Nucleus 2016;7:233–48.
Britton S, Coates J, Jackson SP. A new method for high-resolution imaging of Ku foci to decipher mechanisms of DNA double-strand break repair. J Cell Biol. 2013;202:579–95.
Sawasdichai A, Chen HT, Abdul Hamid N, Jayaraman PS, Gaston K In situ subcellular fractionation of adherent and non-adherent mammalian cells. J Vis Exp. 2010;41:1958.
Malik P, Korfali N, Srsen V, Lazou V, Batrakou DG, Zuleger N, et al. Cell-specific and lamin-dependent targeting of novel transmembrane proteins in the nuclear envelope. Cell Mol Life Sci. 2010;67:1353–69.
Shang G, Zhang C, Chen ZJ, Bai XC, Zhang X Cryo-EM structures of STING reveal its mechanism of activation by cyclic GMP-AMP. Nature. 2019;389–93.
Gonugunta VK, Sakai T, Pokatayev V, Yang K, Wu J, Dobbs N, et al. Trafficking-mediated STING degradation requires sorting to acidified endolysosomes and can be targeted to enhance anti-tumor response. Cell Rep. 2017;21:3234–42.
Chen H, Sun H, You F, Sun W, Zhou X, Chen L, et al. Activation of STAT6 by STING is critical for antiviral innate immunity. Cell. 2011;147:436–46.
Smeenk G, Wiegant WW, Marteijn JA, Luijsterburg MS, Sroczynski N, Costelloe T, et al. Poly(ADP-ribosyl)ation links the chromatin remodeler SMARCA5/SNF2H to RNF168-dependent DNA damage signaling. J Cell Sci. 2013;126:889–903. Pt 4
Lans H, Marteijn JA, Vermeulen W. ATP-dependent chromatin remodeling in the DNA-damage response. Epigenetics Chromatin. 2012;5:4.
Oliveira DV, Kato A, Nakamura K, Ikura T, Okada M, Kobayashi J, et al. Histone chaperone FACT regulates homologous recombination by chromatin remodeling through interaction with RNF20. J Cell Sci. 2014;127:763–72. Pt 4
Clarke TL, Sanchez-Bailon MP, Chiang K, Reynolds JJ, Herrero-Ruiz J, Bandeiras TM, et al. PRMT5-dependent methylation of the TIP60 coactivator RUVBL1 is a key regulator of homologous recombination. Mol Cell. 2017;65:900–16 e7.
Potts PR, Porteus MH, Yu H. Human SMC5/6 complex promotes sister chromatid homologous recombination by recruiting the SMC1/3 cohesin complex to double-strand breaks. EMBO J. 2006;25:3377–88.
Shin YJ, Kim MS, Kim MS, Lee J, Kang M, Jeong JH. High-mobility group box 2 (HMGB2) modulates radioresponse and is downregulated by p53 in colorectal cancer cell. Cancer Biol Ther. 2013;14:213–21.
Blackford AN, Jackson SP. ATM, ATR, and DNA-PK: The Trinity at the Heart of the DNA Damage Response. Mol Cell. 2017;66:801–17.
Ochi T, Blackford AN, Coates J, Jhujh S, Mehmood S, Tamura N, et al. DNA repair. PAXX, a paralog of XRCC4 and XLF, interacts with Ku to promote DNA double-strand break repair. Science. 2015;347:185–8.
Xia T, Konno H, Barber GN. Recurrent loss of STING signaling in melanoma correlates with susceptibility to viral oncolysis. Cancer Res. 2016;76:6747–59.
Konno H, Yamauchi S, Berglund A, Putney RM, Mule JJ, Barber GN. Suppression of STING signaling through epigenetic silencing and missense mutation impedes DNA damage mediated cytokine production. Oncogene. 2018;37:2037–51.
Liang D, Xiao-Feng H, Guan-Jun D, Er-Ling H, Sheng C, Ting-Ting W, et al. Activated STING enhances Tregs infiltration in the HPV-related carcinogenesis of tongue squamous cells via the c-jun/CCL22 signal. Biochim Biophys Acta. 2015;1852:2494–503.
Baird JR, Friedman D, Cottam B, Dubensky TW Jr, Kanne DB, Bambina S, et al. Radiotherapy combined with novel STING-targeting oligonucleotides results in regression of established tumors. Cancer Res. 2016;76:50–61.
Bu Y, Liu F, Jia QA, Yu SN. Decreased expression of TMEM173 predicts poor prognosis in patients with hepatocellular carcinoma. PLoS One. 2016;11:e0165681.
Song S, Peng P, Tang Z, Zhao J, Wu W, Li H, et al. Decreased expression of STING predicts poor prognosis in patients with gastric cancer. Sci Rep. 2017;7:39858.
Bakhoum SF, Cantley LC. The multifaceted role of chromosomal instability in cancer and its microenvironment. Cell. 2018;174:1347–60.
Litvin O, Schwartz S, Wan Z, Schild T, Rocco M, Oh NL, et al. Interferon alpha/beta enhances the cytotoxic response of MEK inhibition in melanoma. Mol Cell. 2015;57:784–96.
Linsley PS, Speake C, Whalen E, Chaussabel D. Copy number loss of the interferon gene cluster in melanomas is linked to reduced T cell infiltrate and poor patient prognosis. PLoS One. 2014;9:e109760.
Chen Y, Wang L, Jin J, Luan Y, Chen C, Li Y, et al. p38 inhibition provides anti-DNA virus immunity by regulation of USP21 phosphorylation and STING activation. J Exp Med. 2017;214:991–1010.
Jiang H, Xue X, Panda S, Kawale A, Hooy RM, Liang F, et al. Chromatin-bound cGAS is an inhibitor of DNA repair and hence accelerates genome destabilization and cell death. EMBO J. 2019;38:e102718.
Chen H, Chen H, Zhang J, Wang Y, Simoneau A, Yang H, et al. cGAS suppresses genomic instability as a decelerator of replication forks. Sci Adv. 2020;6:eabb8941.
Goodwin JF, Knudsen KE. Beyond DNA repair: DNA-PK function in cancer. Cancer Discov. 2014;4:1126–39.
Mohiuddin IS, Kang MH. DNA-PK as an emerging therapeutic target in cancer. Front Oncol. 2019;9:635.
Neal JA, Meek K. Choosing the right path: does DNA-PK help make the decision? Mutat Res. 2011;711:73–86.
Schirmer EC, Florens L, Guan T, Yates JR 3rd, Gerace L. Nuclear membrane proteins with potential disease links found by subtractive proteomics. Science. 2003;301:1380–2.
Malik P, Zuleger N, las Heras JI, Saiz-Ros N, Makarov AA, Lazou V, et al. NET23/STING promotes chromatin compaction from the nuclear envelope. PLoS ONE. 2014;9:e111851.
Katta SS, Smoyer CJ, Jaspersen SL. Destination: inner nuclear membrane. Trends Cell Biol. 2014;24:221–9.
Sullivan T, Escalante-Alcalde D, Bhatt H, Anver M, Bhat N, Nagashima K, et al. Loss of A-type lamin expression compromises nuclear envelope integrity leading to muscular dystrophy. J Cell Biol. 1999;147:913–20.
Marnef A, Legube G. Organizing DNA repair in the nucleus: DSBs hit the road. Curr Opin Cell Biol. 2017;46:1–8.
Oza P, Jaspersen SL, Miele A, Dekker J, Peterson CL. Mechanisms that regulate localization of a DNA double-strand break to the nuclear periphery. Genes Dev. 2009;23:912–27.
Ryu T, Spatola B, Delabaere L, Bowlin K, Hopp H, Kunitake R, et al. Heterochromatic breaks move to the nuclear periphery to continue recombinational repair. Nat Cell Biol. 2015;17:1401–11.
Lemaitre C, Grabarz A, Tsouroula K, Andronov L, Furst A, Pankotai T, et al. Nuclear position dictates DNA repair pathway choice. Genes Dev. 2014;28:2450–63.
Chung DK, Chan JN, Strecker J, Zhang W, Ebrahimi-Ardebili S, Lu T, et al. Perinuclear tethers license telomeric DSBs for a broad kinesin- and NPC-dependent DNA repair process. Nat Commun. 2015;6:7742.
Legrier ME, Bieche I, Gaston J, Beurdeley A, Yvonnet V, Deas O, et al. Activation of IFN/STAT1 signalling predicts response to chemotherapy in oestrogen receptor-negative breast cancer. Br J Cancer. 2016;114:177–87.
Marangoni E, Vincent-Salomon A, Auger N, Degeorges A, Assayag F, de Cremoux P, et al. A new model of patient tumor-derived breast cancer xenografts for preclinical assays. Clin Cancer Res. 2007;13:3989–98.
Baghirova S, Hughes BG, Hendzel MJ, Schulz R. Sequential fractionation and isolation of subcellular proteins from tissue or cultured cells. MethodsX. 2015;2:440–5.
Bolte S, Cordelieres FP. A guided tour into subcellular colocalization analysis in light microscopy. J Microsc. 2006;224:213–32. Pt 3
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Schmidt U, Weigert M, Broaddus C, Myers G, editors. Cell Detection with Star-convex Polygons. In Medical Image Computing and Computer Assisted Intervention – MICCAI 2018 (eds. Frangi, A. F. et al.), Springer International Publishing, 2018;11071:265–73.
Guzman C, Bagga M, Kaur A, Westermarck J, Abankwa D. ColonyArea: an ImageJ plugin to automatically quantify colony formation in clonogenic assays. PLoS One. 2014;9:e92444.
Nagy A, Lanczky A, Menyhart O, Gyorffy B. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci Rep. 2018;8:9227.
Acknowledgements
This study was funded by XenTech, annual funding from Inserm and the University Paris Descartes, and ECLER non-profit association. JG and LC were recipient of a CIFRE fellowship from the Association Nationale de la Recherche et de la Technologie (ANRT). MM is supported by the Ligue Nationale contre le Cancer. We are grateful to Patrice Codogno, Etienne Morel, Ganna Panasyuk, Thierry Dubois and Frédéric Rieux-Laucat for helpful discussions, to Dr Orgunc and Pr S. Fillatreau for providing biological materials, and to Katheryn Meek for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
Conception and design: LC, JG, SC, VG. Development of methodology: LC, ICG, NG, AS, SC, VG. Acquisition of data: LC, ICG, AS, VJ, MP. Analysis and interpretation of data: LC, ICG, NG, AS, NR-R, MM, SC, VG. Writing, review, and/or revision of the manuscript: LC, ICG, AS, MM, J-GJ, SC, VG. Administrative, technical, or material support: LC, ICG, AS, VJ, NR-R, J-GJ, SC, VG. Study supervision: SC, VG.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Cheradame, L., Guerrera, I.C., Gaston, J. et al. STING protects breast cancer cells from intrinsic and genotoxic-induced DNA instability via a non-canonical, cell-autonomous pathway. Oncogene 40, 6627–6640 (2021). https://doi.org/10.1038/s41388-021-02037-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-02037-4
This article is cited by
-
The tumor cell-intrinsic cGAS–STING pathway is associated with the high density of CD8+ T cells after chemotherapy in esophageal squamous cell carcinoma
Esophagus (2024)
-
Ligands stimulating antitumour immunity as the next G-quadruplex challenge
Molecular Cancer (2022)
-
Nuclear soluble cGAS senses double-stranded DNA virus infection
Communications Biology (2022)
-
Topoisomerase I poison-triggered immune gene activation is markedly reduced in human small-cell lung cancers by impairment of the cGAS/STING pathway
British Journal of Cancer (2022)