Abstract
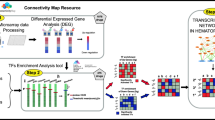
Acute myeloid leukaemia (AML) is a rapidly fatal blood cancer that is characterised by the accumulation of immature myeloid cells in the blood and bone marrow as a result of blocked differentiation. Methods which identify master transcriptional regulators of AML subtype-specific leukaemia cell states and their combinations could be critical for discovering novel differentiation-inducing therapies. In this proof-of-concept study, we demonstrate a novel utility of the Mogrify® algorithm in identifying combinations of transcription factors (TFs) and drugs, which recapitulate granulocytic differentiation of the NB4 acute promyelocytic leukaemia (APL) cell line, using two different approaches. In the first approach, Connectivity Map (CMAP) analysis of these TFs and their target networks outperformed standard approaches, retrieving ATRA as the top hit. We identify dimaprit and mebendazole as a drug combination which induces myeloid differentiation. In the second approach, we show that genetic manipulation of specific Mogrify®-identified TFs (MYC and IRF1) leads to co-operative induction of APL differentiation, as does pharmacological targeting of these TFs using currently available compounds. We also show that loss of IRF1 blunts ATRA-mediated differentiation, and that MYC represses IRF1 expression through recruitment of PML-RARα, the driver fusion oncoprotein in APL, to the IRF1 promoter. Finally, we demonstrate that these drug combinations can also induce differentiation of primary patient-derived APL cells, and highlight the potential of targeting MYC and IRF1 in high-risk APL. Thus, these results suggest that Mogrify® could be used for drug discovery or repositioning in leukaemia differentiation therapy for other subtypes of leukaemia or cancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The raw and processed data were submitted to NCBI GEO. The accession number will be available upon publication.
Code availability
The source code of our pipeline will be available upon publication. Source code for the Mogrify® algorithm is not available.
References
Khwaja A, Bjorkholm M, Gale RE, Levine RL, Jordan CT, Ehninger G, et al. Acute myeloid leukaemia. Nat Rev Dis Prim. 2016;2:16010.
Gonda TJ, Ramsay RG. Directly targeting transcriptional dysregulation in cancer. Nat Rev Cancer. 2015;15:686–94.
Bradner JE, Hnisz D, Young RA. Transcriptional addiction in cancer. Cell. 2017;168:629–43.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663–76.
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131:861–72.
Chao MP, Gentles AJ, Chatterjee S, Lan F, Reinisch A, Corces MR, et al. Human AML-iPSCs reacquire leukemic properties after differentiation and model clonal variation of disease. Cell Stem Cell. 2017;20:329–344.e7.
Kotini AG, Chang C-J, Chow A, Yuan H, Ho T-C, Wang T, et al. Stage-specific human induced pluripotent stem cells map the progression of myeloid transformation to transplantable leukemia. Cell Stem Cell. 2017;20:315–328.e7.
McClellan JS, Dove C, Gentles AJ, Ryan CE, Majeti R. Reprogramming of primary human Philadelphia chromosome-positive B cell acute lymphoblastic leukemia cells into nonleukemic macrophages. Proc Natl Acad Sci USA. 2015;112:4074–9.
Park S-M, Cho H, Thornton AM, Barlowe TS, Chou T, Chhangawala S, et al. IKZF2 drives leukemia stem cell self-renewal and inhibits myeloid differentiation. Cell Stem Cell. 2019;24:153–165.e7.
Wang E, Zhou H, Nadorp B, Cayanan G, Chen X, Yeaton AH, et al. Surface antigen-guided CRISPR screens identify regulators of myeloid leukemia differentiation. Cell Stem Cell. 2021;28:718–731.e6.
Cao Z, Budinich KA, Huang H, Ren D, Lu B, Zhang Z, et al. ZMYND8-regulated IRF8 transcription axis is an acute myeloid leukemia dependency. Mol Cell. 2021;81:3604–3622.e10.
Assi SA, Imperato MR, Coleman DJL, Pickin A, Potluri S, Ptasinska A, et al. Subtype-specific regulatory network rewiring in acute myeloid leukemia. Nat Genet. 2019;51:151–62.
Yun H, Narayan N, Vohra S, Giotopoulos G, Mupo A, Madrigal P, et al. Mutational synergy during leukemia induction remodels chromatin accessibility, histone modifications and three-dimensional DNA topology to alter gene expression. Nat Genet. 2021;53:1443–55.
Rackham OJL, Firas J, Fang H, Oates ME, Holmes ML, Knaupp AS, et al. A predictive computational framework for direct reprogramming between human cell types. Nat Genet. 2016;48:331–5.
Liu TX, Zhang JW, Tao J, Zhang RB, Zhang QH, Zhao CJ, et al. Gene expression networks underlying retinoic acid-induced differentiation of acute promyelocytic leukemia cells. Blood. 2000;96:1496–504.
Yang L, Zhao H, Li S-W, Ahrens K, Collins C, Eckenrode S, et al. Gene expression profiling during all-trans retinoic acid-induced cell differentiation of acute promyelocytic leukemia cells. J Mol Diagn. 2003;5:212–21.
Zheng P-Z, Wang K-K, Zhang Q-Y, Huang Q-H, Du Y-Z, Zhang Q-H, et al. Systems analysis of transcriptome and proteome in retinoic acid/arsenic trioxide-induced cell differentiation/apoptosis of promyelocytic leukemia. Proc Natl Acad Sci USA. 2005;102:7653–8.
de Thé H, Pandolfi PP, Chen Z. Acute promyelocytic leukemia: a paradigm for oncoprotein-targeted cure. Cancer Cell. 2017;32:552–60.
de Thé H. Differentiation therapy revisited. Nat Rev Cancer. 2018;18:117–27.
Lamb J. The connectivity map: using gene-expression signatures to connect small molecules, genes, and disease. Science. 2006;313:1929–35.
Paul F, Arkin Y, Giladi A, Jaitin DA, Kenigsberg E, Keren-Shaul H. et al. Transcriptional heterogeneity and lineage commitment in myeloid progenitors. Cell. 2016;164:325.
Zhang S-D, Gant TW. A simple and robust method for connecting small-molecule drugs using gene-expression signatures. BMC Bioinform. 2008;9:258.
Kuhn M, Szklarczyk D, Franceschini A, von Mering C, Jensen LJ, Bork P. STITCH 3: zooming in on protein-chemical interactions. Nucleic Acids Res. 2012;40:D876–D880.
Kalinyak KA, Sawutz DG, Lampkin BC, Johnson CL, Whitsett JA. Effects of dimaprit on growth and differentiation of human promyelocytic cell line, HL-60. Life Sci. 1985;36:1909–16.
Pantziarka P, Bouche G, Meheus L, Sukhatme V, Sukhatme VP. Repurposing drugs in oncology (ReDO)-mebendazole as an anti-cancer agent. Ecancermedicalscience. 2014;8:443.
McNamara S, Wang H, Hanna N, Miller WH Jr. Topoisomerase IIbeta negatively modulates retinoic acid receptor alpha function: a novel mechanism of retinoic acid resistance. Mol Cell Biol. 2008;28:2066–77.
Nichol JN, Galbraith MD, Kleinman CL, Espinosa JM, Miller WH Jr. NPM and BRG1 mediate transcriptional resistance to retinoic acid in acute promyelocytic leukemia. Cell Rep. 2016;14:2938–49.
Rosenauer A, Raelson JV, Nervi C, Eydoux P, DeBlasio A, Miller WH Jr. Alterations in expression, binding to ligand and DNA, and transcriptional activity of rearranged and wild-type retinoid receptors in retinoid-resistant acute promyelocytic leukemia cell lines. Blood. 1996;88:2671–82.
Shao W, Benedetti L, Lamph WW, Nervi C, Miller WH Jr. A retinoid-resistant acute promyelocytic leukemia subclone expresses a dominant negative PML-RAR alpha mutation. Blood. 1997;89:4282–9.
Duprez E, Benoit G, Flexor M, Lillehaug JR, Lanotte M. A mutated PML/RARA found in the retinoid maturation resistant NB4 subclone, NB4-R2, blocks RARA and wild-type PML/RARA transcriptional activities. Leukemia. 2000;14:255–61.
Testa U, Stellacci E, Pelosi E, Sestili P, Venditti M, Orsatti R, et al. Impaired myelopoiesis in mice devoid of interferon regulatory factor 1. Leukemia. 2004;18:1864–71.
Fiedler K, Brunner C. The role of transcription factors in the guidance of granulopoiesis. Am J Blood Res. 2012;2:57–65.
Leon J, Ferrandiz N, Acosta JC, Delgado MD. Inhibition of cell differentiation: a critical mechanism for MYC-mediated carcinogenesis? Cell Cycle. 2009;8:1148–57.
Prunier C, Zhang M-Z, Kumar S, Levy L, Ferrigno O, Tzivion G, et al. Disruption of the PHRF1 Tumor suppressor network by PML-RARα drives acute promyelocytic leukemia pathogenesis. Cell Rep. 2015;10:883–90.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Shi L, Perin JC, Leipzig J, Zhang Z, Sullivan KE. Genome-wide analysis of interferon regulatory factor I binding in primary human monocytes. Gene. 2011;487:21–28.
Zeller KI, Zhao X, Lee CWH, Chiu KP, Yao F, Yustein JT, et al. Global mapping of c-Myc binding sites and target gene networks in human B cells. Proc Natl Acad Sci USA. 2006;103:17834–9.
Fujiwara T, O’Geen H, Keles S, Blahnik K, Linnemann AK, Kang Y-A, et al. Discovering hematopoietic mechanisms through genome-wide analysis of GATA factor chromatin occupancy. Mol Cell. 2009;36:667–81.
Lan X, Witt H, Katsumura K, Ye Z, Wang Q, Bresnick EH, et al. Integration of Hi-C and ChIP-seq data reveals distinct types of chromatin linkages. Nucleic Acids Res. 2012;40:7690–704.
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P. The Molecular Signatures Database Hallmark Gene Set Collection. Cell Syst. 2015;1:417–25.
Pan X-N, Chen J-J, Wang L-X, Xiao R-Z, Liu L-L, Fang Z-G, et al. Inhibition of c-Myc overcomes cytotoxic drug resistance in acute myeloid leukemia cells by promoting differentiation. PLoS ONE. 2014;9:e105381.
Mei S, Qin Q, Wu Q, Sun H, Zheng R, Zang C, et al. Cistrome Data Browser: a data portal for ChIP-Seq and chromatin accessibility data in human and mouse. Nucleic Acids Res. 2017;45:D658–D662.
Tan Y, Wang X, Song H, Zhang Y, Zhang R, Li S, et al. A PML/RARα direct target atlas redefines transcriptional deregulation in acute promyelocytic leukemia. Blood. 2021;137:1503–16.
Gao T, He B, Liu S, Zhu H, Tan K, Qian J. EnhancerAtlas: a resource for enhancer annotation and analysis in 105 human cell/tissue types. Bioinformatics. 2016;32:3543–51.
Wang P, Tang Z, Lee B, Zhu JJ, Cai L, Szalaj P, et al. Chromatin topology reorganization and transcription repression by PML-RARα in acute promyeloid leukemia. Genome Biol. 2020;21:110.
Hu M-H, Wang Y-Q, Yu Z-Y, Hu L-N, Ou T-M, Chen S-B, et al. Discovery of a new four-leaf clover-like ligand as a potent c-MYC transcription inhibitor specifically targeting the promoter G-Quadruplex. J Med Chem. 2018;61:2447–59.
Michalska A, Blaszczyk K, Wesoly J, Bluyssen HAR. A positive feedback amplifier circuit that regulates interferon (IFN)-stimulated gene expression and controls type I and type II IFN responses. Front Immunol. 2018;9:1135.
Sapio L, Gallo M, Illiano M, Chiosi E, Naviglio D, Spina A, et al. The natural cAMP elevating compound forskolin in cancer therapy: is it time? J Cell Physiol. 2017;232:922–7.
Sawutz DG, Kalinyak K, Whitsett JA, Johnson CL. Histamine H2 receptor desensitization in HL-60 human promyelocytic leukemia cells. J Pharmacol Exp Ther. 1984;231:1–7.
Shayo C, Davio C, Brodsky A, Mladovan AG, Legnazzi BL, Rivera E, et al. Histamine modulates the expression of c-fos through cyclic AMP production via the H2 receptor in the human promonocytic cell line U937. Mol Pharm. 1997;51:983–90.
Brodsky A, Davio C, Shayo C, Lemos Legnazzi B, Barbosa M, Lardo M, et al. Forskolin induces U937 cell line differentiation as a result of a sustained cAMP elevation. Eur J Pharmacol. 1998;350:121–7.
Norsworthy KJ, Altman JK. Optimal treatment strategies for high-risk acute promyelocytic leukemia. Curr Opin Hematol. 2016;23:127–36.
Testa U, Lo-Coco F. Prognostic factors in acute promyelocytic leukemia: strategies to define high-risk patients. Ann Hematol. 2016;95:673–80.
Dos Santos GA, Kats L, Pandolfi PP. Synergy against PML-RARa: targeting transcription, proteolysis, differentiation, and self-renewal in acute promyelocytic leukemia. J Exp Med. 2013;210:2793–802.
Ravandi F, Estey E, Jones D, Faderl S, O’Brien S, Fiorentino J, et al. Effective treatment of acute promyelocytic leukemia with all-trans-retinoic acid, arsenic trioxide, and gemtuzumab ozogamicin. J Clin Oncol. 2009;27:504–10.
Lucena-Araujo AR, Coelho-Silva JL, Pereira-Martins DA, Silveira DR, Koury LC, Melo RAM, et al. Combining gene mutation with gene expression analysis improves outcome prediction in acute promyelocytic leukemia. Blood. 2019;134:951–9.
Lin X, Qiao N, Shen Y, Fang H, Xue Q, Cui B, et al. Integration of genomic and transcriptomic markers improves the prognosis prediction of acute promyelocytic leukemia. Clin Cancer Res. 2021;27:3683–94.
Vicente C, Conchillo A, García-Sánchez MA, Odero MD. The role of the GATA2 transcription factor in normal and malignant hematopoiesis. Crit Rev Oncol Hematol. 2012;82:1–17.
Chen A, Licht JD, Wu Y, Hellinger N, Scher W, Waxman S. Retinoic acid is required for and potentiates differentiation of acute promyelocytic leukemia cells by nonretinoid agents. Blood. 1994;84:2122–9.
Stegmaier K, Ross KN, Colavito SA, O’Malley S, Stockwell BR, Golub TR. Gene expression-based high-throughput screening(GE-HTS) and application to leukemia differentiation. Nat Genet. 2004;36:257–63.
Li Y, Thomas D, Deutzmann A, Majeti R, Felsher DW, Dill DL. Mebendazole for differentiation therapy of acute myeloid leukemia identified by a lineage maturation index. Sci Rep. 2019;9:16775.
Quenech’Du N, Ruchaud S, Khelef N, Guiso N, Lanotte M. A sustained increase in the endogenous level of cAMP reduces the retinoid concentration required for APL cell maturation to near physiological levels. Leukemia. 1998;12:1829–33.
Zhao Q, Tao J, Zhu Q, Jia P-M, Dou A-X, Li X, et al. Rapid induction of cAMP/PKA pathway during retinoic acid-induced acute promyelocytic leukemia cell differentiation. Leukemia. 2004;18:285–92.
Nasr R, Guillemin M-C, Ferhi O, Soilihi H, Peres L, Berthier C, et al. Eradication of acute promyelocytic leukemia-initiating cells through PML-RARA degradation. Nat Med. 2008;14:1333–42.
Padmanabhan A, Li X, Bieberich CJ. Protein kinase A regulates MYC protein through transcriptional and post-translational mechanisms in a catalytic subunit isoform-specific manner. J Biol Chem. 2013;288:14158–69.
Liu Q, Nguyen E, Døskeland S, Ségal-Bendirdjian É. cAMP-dependent protein kinase A (PKA)-mediated c-Myc degradation is dependent on the relative proportion of PKA-I and PKA-II isozymes. Mol Pharm. 2015;88:469–76.
Walf-Vorderwülbecke V, Pearce K, Brooks T, Hubank M, van den Heuvel-Eibrink MM, Zwaan CM, et al. Targeting acute myeloid leukemia by drug-induced c-MYB degradation. Leukemia. 2018;32:882–9.
Coccia EM, Stellacci E, Valtieri M, Masella B, Feccia T, Marziali G, et al. Ectopic expression of interferon regulatory factor-1 potentiates granulocytic differentiation. Biochem J. 2001;360:285–94.
Si J, Yu X, Zhang Y, DeWille JW. Myc interacts with Max and Miz1 to repress C/EBPδ promoter activity and gene expression. Mol Cancer. 2010;9:92.
Zhang L, Li J, Xu H, Shao X, Fu L, Hou Y, et al. Myc-Miz1 signaling promotes self-renewal of leukemia stem cells by repressing Cebpα and Cebpδ. Blood. 2020;135:1133–45.
Wang K, Wang P, Shi J, Zhu X, He M, Jia X, et al. PML/RARα targets promoter regions containing PU.1 Consensus and RARE half sites in acute promyelocytic leukemia. Cancer Cell. 2010;17:186–97.
Lourenco C, Resetca D, Redel C, Lin P, MacDonald AS, Ciaccio R, et al. MYC protein interactors in gene transcription and cancer. Nat Rev Cancer. 2021;21:579–91.
Huang M-J, Cheng Y-C, Liu C-R, Lin S, Liu HE. A small-molecule c-Myc inhibitor, 10058-F4, induces cell-cycle arrest, apoptosis, and myeloid differentiation of human acute myeloid leukemia. Exp Hematol. 2006;34:1480–9.
Nason-Burchenal K, Gandini D, Botto M, Allopenna J, Seale JR, Cross NC, et al. Interferon augments PML and PML/RAR alpha expression in normal myeloid and acute promyelocytic cells and cooperates with all-trans retinoic acid to induce maturation of a retinoid-resistant promyelocytic cell line. Blood. 1996;88:3926–36.
Ng KP, Manjeri A, Lee LM, Chan ZE, Tan CY, Tan QD, et al. The arginase inhibitor Nω-hydroxy-nor-arginine (nor-NOHA) induces apoptosis in leukemic cells specifically under hypoxic conditions but CRISPR/Cas9 excludes arginase 2 (ARG2) as the functional target. PLoS ONE. 2018;13:e0205254.
Sanjana NE, Shalem O, Zhang F. Improved vectors and genome-wide libraries for CRISPR screening. Nat Methods. 2014;11:783–4.
Shyamsunder P, Shanmugasundaram M, Mayakonda A, Dakle P, Teoh WW, Han L, et al. Identification of a novel enhancer of CEBPE essential for granulocytic differentiation. Blood. 2019;133:2507–17.
Corces MR, Buenrostro JD, Wu B, Greenside PG, Chan SM, Koenig JL, et al. Lineage-specific and single-cell chromatin accessibility charts human hematopoiesis and leukemia evolution. Nat Genet. 2016;48:1193–203.
Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A, et al. STRING v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 2013;41:D808–D815.
Acknowledgements
The authors would like to acknowledge Dr. Sonia P. Chothani for providing the RNA-Seq data analysis pipeline, the ssCMAP developer, Dr. Shu-Dong Zhang, for helpful discussion on the algorithm’s internal calculations, the Duke-NUS Genome Biology Facility (DGBF) for RNA, ChIP- and ATAC-sequencing services, as well as Dr. Gee Chuan Wong/Bryan Y.W. Teo, SGH Department of Haematology for providing primary APL samples. This work was funded by the National Medical Research Council (NMRC) of Singapore (MOH-000059/MOH-CSASI18may-0002) and NMRC/CIRG/1429/2015 to STO). OJLR is supported by NMRC YIRG (NMRC/OFYIRG/0022/2016) and by a Singapore National Research Foundation grant [NRF-CRP20-2017-0002].
Author information
Authors and Affiliations
Contributions
Wet lab experiments were performed by LML and PS. RNA-Seq was planned and designed by KLL. RNA-Seq data QC, analysis, Gene Set Functional Enrichments, querying of CMAP, drug ranking, and network analysis were performed by EGC. ChIP data were analysed by BJC. KLL provided guidance on TF GSEA analyses. EGC and LML prepared the manuscript and generated the figures with input from co-authors. KLL, OJLR, EP, and STO designed the study, after conceptualisation by OJLR, EP, and STO. ES and TKF provided the NB4/MR2/LR2 cell lines. GCW provided primary human APL samples from the Singapore General Hospital Haematology Repository.
Corresponding authors
Ethics declarations
Competing interests
OJLR is a co-inventor of the patent (WO/2017/106932) and is a co-founder, shareholder and director of Mogrify Ltd, a cell therapy company. All other authors declare no competing interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, L.M., Christodoulou, E.G., Shyamsunder, P. et al. A novel network pharmacology approach for leukaemia differentiation therapy using Mogrify®. Oncogene 41, 5160–5175 (2022). https://doi.org/10.1038/s41388-022-02505-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02505-5