Abstract
Background
Despite significant advances in multiple myeloma (MM) therapy, disease relapse and treatment resistance remain major obstacles in clinical management. Herein, we have studied the clinical utility of miRNAs in improving patients’ risk-stratification and prognosis.
Methods
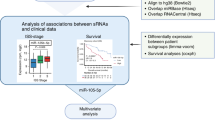
miRNA-seq was performed in CD138+ plasma cells of MM, smoldering multiple myeloma (sMM) and monoclonal gammopathy of undetermined significance (MGUS) patients. The screening MM cohort consisted of 138 patients. miRNA levels of CD138+ plasma cells were quantified by RT-qPCR following 3′-end RNA polyadenylation. Disease progression and patients’ death were used as clinical end-point events. Internal validation was conducted by bootstrap analysis. Clinical net benefit on disease prognosis was assessed by decision curve analysis. Kruykov et al. 2016 served as validation cohort (n = 151).
Results
miRNA-seq highlighted miR-181a to be upregulated in MM vs. sMM/MGUS, and R-ISS III vs. I patients. Screening and validation cohorts confirmed the significantly higher risk for short-term progression and worse survival of the patients overexpressing miR-181a. Multivariate models integrating miR-181a with disease established markers led to superior risk-stratification and clinical benefit for MM prognosis.
Conclusions
CD138+ overexpression of miR-181a was strongly correlated with inferior disease outcome and contributed to superior prediction of MM patients early progression, supporting personalised prognosis and treatment decisions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All the data are available from the corresponding authors on reasonable request.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Rajkumar SV. Multiple myeloma: 2020 update on diagnosis, risk-stratification and management. Am J Hematol. 2020;95:548–67.
Boyle EM, Deshpande S, Tytarenko R, Ashby C, Wang Y, Bauer MA, et al. The molecular make up of smoldering myeloma highlights the evolutionary pathways leading to multiple myeloma. Nat Commun. 2021;12:293.
Dimopoulos MA, Moreau P, Terpos E, Mateos MV, Zweegman S, Cook G, et al. Multiple myeloma: EHA-ESMO Clinical Practice Guidelines for Diagnosis, Treatment and Follow-up. Hemasphere. 2021;5:e528.
Papanota AM, Karousi P, Kontos CK, Ntanasis-Stathopoulos I, Scorilas A, Terpos E. Multiple myeloma bone disease: implication of MicroRNAs in its molecular background. Int J Mol Sci. 2021;22:2375.
Kumar SK, Rajkumar V, Kyle RA, van Duin M, Sonneveld P, Mateos MV, et al. Multiple myeloma. Nat Rev Dis Prim. 2017;3:17046.
Gonzalez-Calle V, Slack A, Keane N, Luft S, Pearce KE, Ketterling RP, et al. Evaluation of revised international staging system (R-ISS) for transplant-eligible multiple myeloma patients. Ann Hematol. 2018;97:1453–62.
Gebert LFR, MacRae IJ. Regulation of microRNA function in animals. Nat Rev Mol Cell Biol. 2019;20:21–37.
Croce CM, Calin GA. miRNAs, cancer, and stem cell division. Cell. 2005;122:6–7.
Diamantopoulos MA, Tsiakanikas P, Scorilas A. Non-coding RNAs: the riddle of the transcriptome and their perspectives in cancer. Ann Transl Med. 2018;6:241.
Di Leva G, Garofalo M, Croce CM. MicroRNAs in cancer. Annu Rev Pathol. 2014;9:287–314.
Chen D, Yang X, Liu M, Zhang Z, Xing E. Roles of miRNA dysregulation in the pathogenesis of multiple myeloma. Cancer Gene Ther. 2021. https://www.nature.com/articles/s41417-020-00291-4.
Rajkumar SV, Dimopoulos MA, Palumbo A, Blade J, Merlini G, Mateos MV, et al. International Myeloma Working Group updated criteria for the diagnosis of multiple myeloma. Lancet Oncol. 2014;15:e538–48.
Kumar S, Paiva B, Anderson KC, Durie B, Landgren O, Moreau P, et al. International Myeloma Working Group consensus criteria for response and minimal residual disease assessment in multiple myeloma. Lancet Oncol. 2016;17:e328–e46.
Kryukov F, Nemec P, Radova L, Kryukova E, Okubote S, Minarik J, et al. Centrosome associated genes pattern for risk sub-stratification in multiple myeloma. J Transl Med. 2016;14:150.
Friedlander MR, Mackowiak SD, Li N, Chen W, Rajewsky N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 2012;40:37–52.
Wong N, Wang X. miRDB: an online resource for microRNA target prediction and functional annotations. Nucleic Acids Res. 2015;43:D146–52.
Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 2016;44:W90–7.
Supek F, Bosnjak M, Skunca N, Smuc T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE. 2011;6:e21800.
Vickers AJ, Elkin EB. Decision curve analysis: a novel method for evaluating prediction models. Med Decis Mak. 2006;26:565–74.
Chen CZ, Li L, Lodish HF, Bartel DP. MicroRNAs modulate hematopoietic lineage differentiation. Science. 2004;303:83–6.
Schmidt TM, Barwick BG, Joseph N, Heffner LT, Hofmeister CC, Bernal L, et al. Gain of Chromosome 1q is associated with early progression in multiple myeloma patients treated with lenalidomide, bortezomib, and dexamethasone. Blood Cancer J. 2019;9:94.
Schmidt TM, Fonseca R, Usmani SZ. Chromosome 1q21 abnormalities in multiple myeloma. Blood Cancer J. 2021;11:83.
Landgren O, Rajkumar SV. New developments in diagnosis, prognosis, and assessment of response in multiple myeloma. Clin Cancer Res. 2016;22:5428–33.
Dimopoulos MA, Jakubowiak AJ, McCarthy PL, Orlowski RZ, Attal M, Blade J, et al. Developments in continuous therapy and maintenance treatment approaches for patients with newly diagnosed multiple myeloma. Blood Cancer J. 2020;10:17.
Rasche L, Chavan SS, Stephens OW, Patel PH, Tytarenko R, Ashby C, et al. Spatial genomic heterogeneity in multiple myeloma revealed by multi-region sequencing. Nat Commun. 2017;8:268.
Rupaimoole R, Slack FJ. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov. 2017;16:203–22.
Avgeris M, Panoutsopoulou K, Papadimitriou MA, Scorilas A. Circulating exosomal miRNAs: clinical significance in human cancers. Expert Rev Mol Diagn. 2019;19:979–95.
Morelli E, Leone E, Cantafio ME, Di Martino MT, Amodio N, Biamonte L, et al. Selective targeting of IRF4 by synthetic microRNA-125b-5p mimics induces anti-multiple myeloma activity in vitro and in vivo. Leukemia. 2015;29:2173–83.
Yin Y, Yang W, Zhang L, Liu K, Luo Z. Long non-coding RNA ANRIL and its target microRNAs (microRNA-34a, microRNA-125a and microRNA-186) relate to risk stratification and prognosis in multiple myeloma. Hematology. 2021;26:160–9.
Misso G, Zarone MR, Lombardi A, Grimaldi A, Cossu AM, Ferri C, et al. miR-125b upregulates miR-34a and sequentially activates stress adaption and cell death mechanisms in multiple myeloma. Mol Ther Nucleic Acids. 2019;16:391–406.
Di Martino MT, Leone E, Amodio N, Foresta U, Lionetti M, Pitari MR, et al. Synthetic miR-34a mimics as a novel therapeutic agent for multiple myeloma: in vitro and in vivo evidence. Clin Cancer Res. 2012;18:6260–70.
Leone E, Morelli E, Di Martino MT, Amodio N, Foresta U, Gulla A, et al. Targeting miR-21 inhibits in vitro and in vivo multiple myeloma cell growth. Clin Cancer Res. 2013;19:2096–106.
Rossi M, Altomare E, Botta C, Gallo Cantafio ME, Sarvide S, Caracciolo D, et al. miR-21 antagonism abrogates Th17 tumor promoting functions in multiple myeloma. Leukemia. 2021;35:823–34.
Amodio N, Di Martino MT, Foresta U, Leone E, Lionetti M, Leotta M, et al. miR-29b sensitizes multiple myeloma cells to bortezomib-induced apoptosis through the activation of a feedback loop with the transcription factor Sp1. Cell Death Dis. 2012;3:e436.
Pan Y, Zhang Y, Liu W, Huang Y, Shen X, Jing R, et al. LncRNA H19 overexpression induces bortezomib resistance in multiple myeloma by targeting MCL-1 via miR-29b-3p. Cell Death Dis. 2019;10:106.
Xu J, Su Y, Xu A, Fan F, Mu S, Chen L, et al. miR-221/222-mediated inhibition of autophagy promotes dexamethasone resistance in multiple myeloma. Mol Ther. 2019;27:559–70.
Zhao JJ, Chu ZB, Hu Y, Lin J, Wang Z, Jiang M, et al. Targeting the miR-221-222/PUMA/BAK/BAX pathway abrogates dexamethasone resistance in multiple myeloma. Cancer Res. 2015;75:4384–97.
Qin Y, Zhang S, Deng S, An G, Qin X, Li F, et al. Epigenetic silencing of miR-137 induces drug resistance and chromosomal instability by targeting AURKA in multiple myeloma. Leukemia. 2017;31:1123–35.
Yang N, Chen J, Zhang H, Wang X, Yao H, Peng Y, et al. LncRNA OIP5-AS1 loss-induced microRNA-410 accumulation regulates cell proliferation and apoptosis by targeting KLF10 via activating PTEN/PI3K/AKT pathway in multiple myeloma. Cell Death Dis. 2017;8:e2975.
Emming S, Chirichella M, Monticelli S. MicroRNAs as modulators of T cell functions in cancer. Cancer Lett. 2018;430:172–8.
Pichiorri F, Suh SS, Ladetto M, Kuehl M, Palumbo T, Drandi D, et al. MicroRNAs regulate critical genes associated with multiple myeloma pathogenesis. Proc Natl Acad Sci USA. 2008;105:12885–90.
Roccaro AM, Sacco A, Thompson B, Leleu X, Azab AK, Azab F, et al. MicroRNAs 15a and 16 regulate tumor proliferation in multiple myeloma. Blood. 2009;113:6669–80.
Li YJ, Li DP, Yan ZL, Qi KM, Chen LL, Zhang ZY, et al. Potential relationship and clinical significance of miRNAs and Th17 cytokines in patients with multiple myeloma. Leuk Res. 2014;38:1130–5.
Giri S, Huntington SF, Wang R, Zeidan AM, Podoltsev N, Gore SD, et al. Chromosome 1 abnormalities and survival of patients with multiple myeloma in the era of novel agents. Blood Adv. 2020;4:2245–53.
Abdallah N, Greipp P, Kapoor P, Gertz MA, Dispenzieri A, Baughn LB, et al. Clinical characteristics and treatment outcomes of newly diagnosed multiple myeloma with chromosome 1q abnormalities. Blood Adv. 2020;4:3509–19.
Peng J, Thakur A, Zhang S, Dong YF, Wang XQ, Yuan RL, et al. Expressions of miR-181a and miR-20a in RPMI8226 cell line and their potential as biomarkers for multiple myeloma. Tumor Biol. 2015;36:8545–52.
Liu N, Yang J, Yuan R, Peng J, Liu L, Guo X. Effects of miR181a on the biological function of multiple myeloma. Oncol Rep. 2019;42:291–300.
Shen XX, Bai H, Zhu HY, Yan Q, Yang Y, Yu WJ, et al. Long non-coding RNA MEG3 functions as a competing endogenous RNA to regulate HOXA11 expression by sponging miR-181a in multiple myeloma. Cell Physiol Biochem. 2018;49:87–100.
Panoutsopoulou K, Avgeris M, Magkou P, Mavridis K, Dreyer T, Dorn J, et al. miR-181a overexpression predicts the poor treatment response and early-progression of serous ovarian cancer patients. Int J Cancer. 2020;147:3560–73.
Taylor MA, Sossey-Alaoui K, Thompson CL, Danielpour D, Schiemann WP. TGF-beta upregulates miR-181a expression to promote breast cancer metastasis. J Clin Invest. 2013;123:150–63.
Nishimura J, Handa R, Yamamoto H, Tanaka F, Shibata K, Mimori K, et al. microRNA-181a is associated with poor prognosis of colorectal cancer. Oncol Rep. 2012;28:2221–6.
Meijer LL, Garajova I, Caparello C, Le Large TYS, Frampton AE, Vasile E, et al. Plasma miR-181a-5p downregulation predicts response and improved survival after FOLFIRINOX in pancreatic ductal adenocarcinoma. Ann Surg. 2020;271:1137–47.
Mi Y, Zhang D, Jiang W, Weng J, Zhou C, Huang K, et al. miR-181a-5p promotes the progression of gastric cancer via RASSF6-mediated MAPK signalling activation. Cancer Lett. 2017;389:11–22.
Liu X, Liao W, Peng H, Luo X, Luo Z, Jiang H, et al. miR-181a promotes G1/S transition and cell proliferation in pediatric acute myeloid leukemia by targeting ATM. J Cancer Res Clin Oncol. 2016;142:77–87.
Lyu X, Li J, Yun X, Huang R, Deng X, Wang Y, et al. miR-181a-5p, an inducer of Wnt-signaling, facilitates cell proliferation in acute lymphoblastic leukemia. Oncol Rep. 2017;37:1469–76.
Fragoso R, Mao T, Wang S, Schaffert S, Gong X, Yue S, et al. Modulating the strength and threshold of NOTCH oncogenic signals by mir-181a-1/b-1. PLoS Genet. 2012;8:e1002855.
Haque S, Vaiselbuh SR. Silencing of exosomal miR-181a reverses pediatric acute lymphocytic leukemia cell proliferation. Pharmaceuticals (Basel). 2020;13:241.
Acknowledgements
We thank all patients and families for their participation. We thank IEMBITHEK (Greece) for partially funded the study. We thank Bodossaki Foundation (Athens, Greece) for partially supported the study by fellowship to PGA.
Author information
Authors and Affiliations
Contributions
Conception and design: AS, ET, MAD, MA; Development of methodology: MAP, AMP, MA; Acquisition of data: MAP, AMP, PGA, KMP, CIL, PM, NMK, DP, EEP, MG, EK, MA; Analysis and interpretation of data: MAP, AMP, PGA, KMP, MA, MAD, ET, AS; Acquired and managed patients: AMP, MG, EK, MAD, ET; Drafting of the manuscript: MAP, AMP, PGA, MA; Critical revision of the manuscript: MAD, ET, AS; Study supervision: AS, ET, MAD, MA; Approval of the submitted and final version: All authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The study was approved by the Ethics Committee of “Alexandra” Hospital, Athens, Greece, and all patients who participated were informed and signed an informed consent according to 1975 Declaration of Helsinki ethical standards, as revised in 2008.
Consent for publication
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Papadimitriou, MA., Papanota, AM., Adamopoulos, P.G. et al. miRNA-seq and clinical evaluation in multiple myeloma: miR-181a overexpression predicts short-term disease progression and poor post-treatment outcome. Br J Cancer 126, 79–90 (2022). https://doi.org/10.1038/s41416-021-01602-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-021-01602-8
This article is cited by
-
miRNA-seq identification and clinical validation of CD138+ and circulating miR-25 in treatment response of multiple myeloma
Journal of Translational Medicine (2023)
-
Comprehensive small RNA-sequencing of primary myeloma cells identifies miR-105-5p as a predictor of patient survival
British Journal of Cancer (2023)