Abstract
The role of mitosis in the progression of precancerous skin remains poorly understood. To address this question, we deleted the mitotic Kinase Aurora-A (Aur-A) in hyperplastic mutant p53 mouse skin as an experimental tool to study the G2/M transition in precancerous keratinocytes and AUR-A’s role in this process. Epidermal Aur-A deletion (Aur-AepiΔ) led to marked keratinocyte enlargement, pleomorphism, multinucleation, and attenuated induction of cell death. This phenotype was characteristic of slippage after a stalled mitosis. We also observed altered or impaired epidermal differentiation, indicative of a partial skin barrier defect. The upregulation of mTOR/PI3K signaling was implicated as a mechanism by which keratinocytes may evade cell death after AUR-A deficiency. This was evidenced by the ectopic expression of the pathway readout, p-S6, in the basal layer of Aur-AepiΔ skin and its mitotic upregulation in isolated keratinocytes. We further tested whether our findings were extended to skin carcinoma cells. The chemical inhibition of AUR-A led to a similar mitotic delay, polyploidy/multinucleation, and attenuated cell death in skin cancer cell lines. Moreover, inhibition of mTOR/PI3K signaling ameliorated the effects caused by the deficiency of AUR-A activity but was also associated with the persistence of mitotic p-S6 detection in surviving cancer cells. These results show the induction of multinucleation/polyploidy may be a compensatory state in keratinocytes that allows for cellular survival and maintenance of partial barrier function in face of aberrant cell division or differentiation. Moreover, mTOR/PI3K signaling is active in the mitosis of hyperplastic keratinocytes expressing mutant p53 and is further enhanced by stalled mitosis, indicating a potential resistance mechanism to the use of anti-mitotic drugs in the treatment of skin cancers.
Similar content being viewed by others
Introduction
Cutaneous squamous cell carcinomas (SCCs) are the second most common skin neoplasia but account for the majority of metastasis and fatalities in non-melanoma skin cancers [1]. DNA sequencing of skin SCCs has revealed common drivers that implicate NOTCH1/2, p53, and RAS signaling in its carcinogenesis [2,3,4]. With respect to p53, mutations are found in 58–95% of skin SCCs, which mostly fall within the DNA-binding domain of the protein and lead to the loss-of-function (LOF) of its tumor suppressive activities [3, 5,6,7]. There are also gain-of-function (GOF) mutations that confer oncogenic properties to mutant p53 [8, 9]. LOF and GOF p53 mutations are found in precancerous lesions and tumors arising in sun-exposed or non-sun-exposed skin (see p53.iarc.fr) [10,11,12]. On the other hand, the frequency of RAS mutations in SCCs ranges from 5% to 46% [3, 7, 13]. Moreover, RAS mutations have been reported in precancerous lesions [14] and tumor-related overexpression of RAS or elevated levels of its active GTP-bound form in the absence of activating mutations also contribute to SCC carcinogenesis [15,16,17].
We have focused on two oncogenes associated with aggressive skin cancers, namely, AURORA-A (AUR-A) kinase and the GOF p53R175H mutant. AUR-A has diverse roles in regulating cell division that include promoting the entry into mitosis and bipolar spindle assembly [18]. AUR-A is a cancer-susceptibility gene that is frequently amplified or overexpressed in epithelial tumors, including skin cancers [19,20,21,22]. AUR-A transforms immortalized cells and promotes the malignant conversion of chemically induced skin cancers, resulting in metastasis-prone tumors that are characterized by centrosome amplification and genomic instability [20, 23]. On the other hand, the GOF p53R175H (R172H in mice) mutant is pro-oncogenic in humans and mice; its expression in tumors is associated with genomic instability and metastasis [8, 24,25,26]. With respect to skin, experimental mice that co-express the KRASG12D and p53R172H mutants develop metastasis-prone skin SCCs that display dysregulation of AUR-A [25]. This is in contrast to LOF p53 skin, which is less likely to develop aggressive or metastatic SCCs [25].
The mTOR/PI3K pathway has also been implicated in skin carcinogenesis [27]. The mTOR/PI3K pathway integrates extracellular signals from the microenvironment that can regulate cell size, proliferation, survival, and differentiation. Under non-pathological conditions, the mTOR/PI3K pathway contributes to epidermal skin development and homeostasis [28, 29]. Effectors of the pathway like AKT and S6 are expressed in the upper layers of skin and may protect terminally differentiated keratinocytes (KCs) from apoptosis [30].
In the current study, we aim to dissect the role of AUR-A in the mitotic regulation of preneoplastic KCs by inducing the deletion of Aur-A in hyperplastic LOF/GOF p53 skin. Our results implicate the mTOR/PI3K pathway in mediating multinucleation, cell enlargement, and survival of mutant p53 KCs deficient in AUR-A.
Results
Genetic deletion of Aur-A results in multinucleation and cell survival in hyperplastic mutant p53 skin
In the LOF/GOF p53 skin cancer model, the expression of the KRASG12D mutant is combined with the ablation of p53 or activation of the GOF p53R172H allele [26]. We included the deletion of Aur-A as an experimental tool to understand the regulation of the G2/M phases of precancerous KCs in the context of mutant p53. We analyzed tissues 14 days after the recombination of CreER-inducible alleles by Tamoxifen (TAM) and 7 days after the co-treatment with the tumor promoter TPA (Fig. 1a). TPA allows the normally low cycling basal KCs in the adult epidermis to be hyperplastic [31]. We chose this TPA treatment regime because it led to a significant increase in the percentage of mutant p53 KCs found in S-phase (Supplementary Figure 1). Epidermal deletion of Aur-A (Aur-AepiΔ) skin showed diffused hyperplasia, cellular pleomorphism with marked cell karyomegaly, prominent nucleoli, irregular cell size, and the accumulation of multinucleated cells (Fig. 1b). These effects also seemed more pronounced in Aur-AepiΔ; GOF p53 skin. Nevertheless, epidermal differentiation also appeared impaired and coincided with increased epidermal thickness.
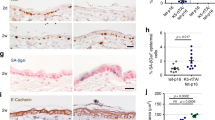
The ablation of Aur-A in hyperplastic mouse skin-induced multinucleation and keratinocyte (KC) survival. a Schema of in vivo studies. Mutant Kras, loss-of-function (LOF) or gain-of-function (GOF) p53 alleles were combined with floxed Aur-A alleles. Mice were fed Tamoxifen containing diets and topically treated with TPA. b Images show the histological appearance of TPA-treated mutant p53 back skin following ablation of Aur-A. Note the hyperplastic appearance of the LOF and GOF p53 epidermis and the enlarged and multinucleated KCs in Aur-A ablated groups. c Co-detection of E-CADHERIN and LMNA (LAMIN-A/C) by immunofluorescence (left panels). E-CADHERIN + cells with more than one LMNA + nuclear structure were quantified (right graph). d Proliferation of skin was assessed by the immunodetection of KI-67 (all cell cycle phases) and by the mitotic marker phospho (p)-HISTONE H3 (p-HH3). Dashed lines indicate the epidermis/dermis boundary. The graph on the right shows the enumeration of p-HH3+ cells. e Left panels show the IHC detection of the apoptotic marker, active CASPASE-3 by IHC. The graph on the right shows the quantification of the staining. f Apoptotic bodies were enumerated at 1000× magnification on H&E stained skin sections. g Images show immunoblotting for the anti-apoptotic protein BCL-XL. Blots were then stripped and probed for β-ACTIN. Quantification of BCL-XL normalized to β-ACTIN revealed no significant changes between genotypes. Scale bar, 10 μm for a, e and 25 μm for c, d. Counting for c, d, e was performed on at least five 200× fields per sample. Counting for f was performed on at least five 1000× fields per sample. Graph values represent mean ± s.d. Sample sizes in c, d, e: n = 5 for NC, n = 5 for LOF p53, n = 5 for GOF p53, n = 6 for Aur-AepiΔ; LOF p53, and n = 4 for Aur-AepiΔ; GOF p53 genotypes. Sample size in f represent n = 5 for NC, n = 6 for LOF p53, n = 6 for GOF p53, n = 7 for Aur-AepiΔ; LOF p53, and n = 8 for Aur-AepiΔ; GOF p53 genotypes. See Material and methods for p-value designated by the stars (“*”) on the graphs (applicable to all figures)
We co-stained with the epithelial membrane marker E-CADHERIN and the nuclear membrane marker LNMA (LAMIN-A/C) (Fig. 1c) to examine more closely the observed multinucleation. We quantified LNMA+ nuclear structures in KCs attached to the basal or suprabasal layers; this included cells with multiple nuclei or smaller micronuclei. Aur-AepiΔ skin had substantially elevated levels of LNMA+ nuclear structures compard to mutant p53 skin (Fig. 1c). Notably, multinucleation was also detected in NC control skin at low levels. Aur-AepiΔ skin was also proliferative, but showed a marked increase in the mitotic index as demonstrated by p-HH3 staining (Supplementary Figure 2 and Fig. 1d). Furthermore, samples with elevated p-HH3+ counts tended to have elevated number of LNMA+ structures (Pearson correlation, r = 0.7, p < 0.0001, n = 24). These results indicate that Aur-AepiΔ KCs stalled in mitosis but underwent mitotic slippage, allowing them to exit without completion of cytokinesis. Moreover, these data also show that multinucleation may be inherent to skin KCs, which can be enhanced by mitotic dysfunction.
Next, we determine if cell death was a feature of Aur-AepiΔ skin. The detection of active CASPASE-3 was variable in mutant p53 skin. Unexpectedly, active CASPASE-3 detection was not enhanced by AUR-A deficiency (Fig. 1e). Similarly, modest changes were evident in the histological detection of apoptotic bodies (Fig. 1f). However, no changes were observed in the expression of the apoptotic-associated protein BCL-XL (Fig. 1g). We conclude that apoptosis was attenuated by AUR-A deficiency in hyperplastic mutant p53 skin.
RNA sequencing of AUR-A-deficient skin revealed altered differentiation and enrichment of mitotic gene signatures
We profiled the transcriptome to better understand the cell fate observed in Aur-AepiΔ skin. Few robust changes were observed between GOF vs. LOF p53 skin (cutoff of q < 0.05, Supplementary Tables 1–2), whereas Aur-A deletion had a greater effect on the transcriptome (Fig. 2a and Supplementary Tables 3–4). We found the enrichment of epidermis and ectoderm development genes in the Aur-AepiΔ skin datasets (Fig. 2b and Supplementary Table 5). These datasets included epidermal skin differentiation genes like Krt1, Krt10, and Sprr1b, and “stress” genes like Krt16 that are associated with inflammatory conditions like psoriasis and Harlequin ichthyosis [32,33,34]. We confirmed the downregulation of Krt1 and Krt10 and upregulation of Krt14, Krt16, and Sprr1b in Aur-AepiΔ skin RNA (Supplementary Figure 3a). We also stained for the terminal skin differentiation marker, FILAGGRIN [35]. Its pattern of detection in the stratum corneum of Aur-AepiΔ skin was disorganized and its expression levels reduced (Supplementary Figure 3b). These results indicated that Aur-AepiΔ skin presents with reduced/altered skin differentiation and may have a partial barrier defect.
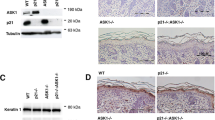
Deletion of Aur-A alters the transcriptome of hyperplastic mutant p53 KCs. a RNA sequencing was performed on total RNA extracted from whole skin. Principle component analysis was performed on transcript profiles from the LOF and GOF p53 groups, with and without Aur-A deletion. Note the proximity of LOF and GOF p53 samples and the separation after Aur-A deletion. b Genelists were compared to generate non-overlapping lists as shown. Category enrichment was determined by comparing gene ontology genesets with the genes that overlapped between Aur-A deleted vs. wildtype (WT) groups. The significance of enrichment was calculated by a Fisher exact test. c QPCR validation was performed on candidate genes identified by RNAseq. Expression was normalized to 18S abundance. Mean ± s.d. is shown. d Panels show detection of AUR-A at KC centrosomes using confocal microscopy. Images show maximal projections that allows for the visualization of both centrosome poles. Arrows depict the location of AUR-A or γ-tubulin. Note the location of γ-tubulin staining and the absence of AUR-A staining in mitotic cells (p-HH3+ cells) in Aur-AepiΔ; LOF p53 skin. This staining pattern was also observed in Aur-A ablated GOF p53 skin. Scale bar, 5 μm. The sample number (n = 19) is shown by the symbols in a and was used in c, d
G2/M and mitotic gene signatures were also over-represented in Aur-AepiΔ skin and included AUR-A interacting partners such as Tpx2, Ajuba, and Bora (Fig. 2b, c and Supplementary Table 5) and mitotic genes like Ccnb1 (Cyclin B1) and Aurora Kinase-B (Aur-B), indicating the enrichment of G2/M cells. CCNB1/CDK1 complexes promote entry into mitosis, but CCNB1 destruction is necessary for mitotic exit or for mitotic slippage [36]. However, Ccnb1 levels were elevated in Aur-AepiΔ skin (Fig. 2c). AUR-B was also upregulated in Aur-AepiΔ skin (Fig. 2c and Supplementary Figure 4a). AUR-B is part of the chromosome passenger complex and is a key regulator of the Spindle Assembly Checkpoint [37]. The activation of mitotic checkpoints likely contributed to stalled Aur-AepiΔ KCs due to the presence of abnormal spindles and microtubules (see below, Fig. 2d and Supplementary Figure 4a-b). Lastly, we also observed elevated levels of Ccnd1 (Cyclin D1) (Fig. 2c). CCND1 accumulates in G1 and is necessary for progression into S-phase, indicating that Aur-AepiΔ KCs remained in cycle.
Examination of p-HH3+ Aur-AepiΔ cells more closely by confocal microscopy revealed other abnormalities in mitosis. These included cells with monopolar spindles as shown by the localization of γ-tubulin to a single mitotic pole (Fig. 2d). In contrast, AUR-A was co-localized with γ-tubulin at both centrosomes in WT cells (Fig. 2d). On the other hand, TPX2 staining revealed abnormally formed microtubules. TPX2 normally forms a complex with AUR-A that enables its activation and has important roles in microtubule dynamics [38]. TPX2 localized with γ-tubulin and centrosomes during WT KC mitosis but was mislocalized in Aur-AepiΔ cells (Supplementary Figure 4b).
MTOR/PI3K signaling is enhanced in AUR-A-deficient mutant p53 skin
Aur-AepiΔ KCs appeared larger than WT cells (see Fig. 1b, c). This may be due to mitotic slippage or involvement of signaling pathways like mTOR/PI3K that can regulate cell size and survival. Applying the Ingenuity pathway analyses software algorithms (qiagenbioinformatics.com/IPA) to our RNAseq datasets predicted the activation of the mTOR/PI3K pathway in Aur-AepiΔ skin (Supplementary Figure 5). By immunoblotting of whole skin samples, we found the levels of p-S6 and p-AKT (S473 and T308) to be elevated in GOF vs. LOF p53 skin, and remained elevated by AUR-A deficiency (Fig. 3a); phosphorylation at both S473 and T308 indicated maximal activation of AKT [39]. We then explored upstream signaling nodes. Similar to AKT, higher levels of p-RICTOR and p-RAPTOR were observed GOF vs. LOF p53 skin (Fig. 3b), which remained elevated in Aur-AepiΔ skin. RAPTOR and RICTOR are obligate members of the mTOR C1 and C2 complexes, respectively [28]. The C1 complex phosphorylates AKT, whereas S6K1, which is an effector of mTORC1 [40], showed reduced levels of phosphorylation following Aur-A deletion. On the other hand, the p-epidermal growth factor receptor (EGFR) was elevated similar to RAPTOR and RICTOR (Fig. 3b). We also observed enhanced p-ERK levels that can also synergize with downstream mTOR/PI3K signaling by the activation of p90RSK [41]. Taken together, the activation of AKT and S6 in Aur-AepiΔ skin would appear to be driven by EGFR and mTORC2 complex activities.
Analysis of mTOR/PI3K signaling in Aur-A deleted skin. a The phosphorylation status of AKT and S6 was determined in lysates from untreated and TPA treated skin. Lysates from untreated skin were immunoblotted for p-AKT (T308/S473) and p-S6 (S235/236). Blots were then stripped and probed for total protein (i.e., AKT and S6). Blots were stripped once more and re-probed for the loading control β-ACTIN. TPA treated skin showed high levels of p-AKT (T308 and S473) and p-S6 in GOF vs. LOF p53 skin, which remained increased by AUR-A deficiency. The ratios of phospho to total kinase levels were calculated by measuring band intensities (bottom graphs). b The phosphorylation status of RICTOR, RAPTOR, p70-S6K1, ERK, and EGFR was determined in lysates from TPA-treated skin. Blots were then stripped and probed for its total protein. Blots were stripped once more and re-probed for the loading control β-ACTIN. The ratios of phospho to total protein were determined by measuring band intensities. The graphs show the mean of the ratios ± s.d. and the sample size are represented by the lanes shown in the blots
Next, we focused on p-S6 expression in the epidermis as a readout of the mTOR/PI3K pathway activity in Aur-AepiΔ skin. Detection of p-S6 was evident in the upper layers of untreated skin, overlapping with the suprabasal marker KRT1 (Fig. 4a, b) and consistent with the established role of mTOR/PI3K signaling in skin homeostasis [28]. In hyperplastic WT p53 skin, p-S6 and KRT1 detection was co-localized to the suprabasal layers. In contrast, p-S6 staining became more intense and extended down into the hair follicles of mutant p53 skin. This was unlike the pattern observed with KRT1, which remained associated with the upper layers of the skin. AUR-A deficiency extended the detection of p-S6 to the basal layer (Fig. 4a). Its detection appeared higher in the basal layer while reduced in the suprabasal compartment (Fig. 4c). In contrast, S6 appeared to be distributed across the epidermis (Fig. 4a, c). The ratio of p-S6/S6 was higher in the basal layer of the epidermis but remained unchanged in the suprabasal layers. Notably, ectopic expression of p-S6 in the basal layer was undetectable in p53wt/KRASwt embryonic epidermis deficient in AUR-A (Supplementary Figure 6) [42]. Thus, the effects on p-S6 levels and its tissue localization in adult Aur-AepiΔ skin appeared to depend on the p53 and RAS status. Lastly, the detection of KRT1 was reduced in Aur-AepiΔ skin and its pattern of expression disorganized (Fig. 4b), similar to the pattern of FILAGGRIN detection (Supplementary Figure 3b).
Ectopic expression of p-S6 in the basal layer of mutant p53 epidermis deficient in AUR-A. a Images show the immunofluorescent detection of pS6 and S6 in untreated skin and TPA-treated NC and mutant p53 skin. Note that the detection of pS6 in basal cells extends to the differentiated layers in Aur-A deleted epidermis, whereas the pattern of S6 detection is evenly distributed across the epidermis. b Images show the detection of the basal layer marker KRT14 and the suprabasal and early differentiation marker KRT1. Note KRT1’s disorganized pattern of expression in Aur-A deleted epidermis. Representative images are shown for four mice per genotype. Scale bar, 50 μm. c Graphs show the quantification of p-S6 to total protein by pixel intensity per surface area (SA) of the region of interest of KCs proximal to the basement membrane of the epidermis (basal area) and superior to this region (suprabasal). At least three areas in the epidermis were analyzed per slide. Graphs represent mean ± s.d. with n = 6 for each group. Equal number (n = 3) of LOF and GOF p53 samples define the Aur-AWT group, whereas equal number of Aur-AepiΔ; LOF p53 and Aur-AepiΔ; GOF p53 samples define the Aur-AepiΔ group
AUR-A deficiency induces cell cycle-specific changes in AKT and S6 phosphorylation
To define how the ectopic activation of the mTOR/PI3K pathway could protect basal Aur-AepiΔ KCs during stalled mitoses, we performed mass cytometric experiments to correlate p-protein levels with cell cycle states [43]. In these experiments, we use single KCs isolated from 4HOTam/TPA-treated mice ears due to the difficulty of isolating KCs from adult back skin. We also treated for a shorter duration (3 days) with a higher dose of TPA (10 nmol) to avoid analyzing cells that were too different in cell size. We examined the different cell cycle phases (i.e., S, G2, M, G1, Go) by using bivariate plots of incorporated IdU (5′-Iodo-2′-deoxyuridine) vs. CCNB1 and p-HH3Ser28, or p-RBSer807/811 (Fig. 5a) [43]. Control KCs were predominately in G1 but this was reduced at the expense of S-phase and G2-phase in mutant p53 KCs (Fig. 5b and Table 1). Both the S-phase and M-phase fractions were further increased by AUR-A deficiency (Fig. 5b and Table 1). Furthermore, AUR-A deficiency also enhanced the number of p-HH3+ cells and its mean channel intensity (i.e., expression levels) (Fig. 5c). This was correlated with elevated levels of p-S6, p-AKT, and p-EGFR detection in the same cell (Fig. 5c, right panels).
Phospho-S6 and -AKT levels are increased during mitosis of Aur-A-deficient KCs. Suspension of epidermal cells were stained with a panel of metal-tagged antibodies and analyzed by mass cytometry. a Panels show gating strategy to discriminate between the different cell cycle phases. Single cells were identified by 191/193Ir (Iridium) incorporation. Live cells were determined by the exclusion of 195Pt (Platinum) uptake. Immune cells were excluded by gating on 89Y (Yttrium) CD45 negative cells. Detection of 127I (Iodine) labeled DeoxyUridine allowed the determination of DNA synthesis in S-phase. Staining with 164Dy (Dysprosium)-labeled anti-CCNB1 (CYCLIN B1) allowed the detection of G2/M cells. b Pie charts show cell cycle profiles based on p53 and Aur-A status. Cell cycle phases were determined described in a. The average of four independent animal treatments is shown per genotype. See Table 1 for tabulation. c G2/M-gated KCs were analyzed for detection of anti-152Sm (Samarium) p-AKTS473, anti-172Yb (Ytterbium) p-S6S235/S236, anti-146Nd (Neodymium) p-EGFRY1068 (not shown), and anti-175Lu (Lutetium) p-HH3 (Left plots). Note the increase in AKT intensity in p-S6+/p-HH3+ KCs. Right panels show the quantification of channel intensity
Next, we used the Citrus software (cytobank.org) to organize unsupervised clusters around cell cycle markers to determine the differences in p-proteins using statistical correlation models in the same cell [44]. This approach also controls for the increased phosphorylation that normally is observed during mitosis [45]. We compared WT vs. Aur-AepiΔ KCs, regardless of their p53 status. Branches of the clustering tree were organized phenotypically and hierarchically by the expression of cell cycle markers (Supplementary Figure 7a). Statistical modeling indicated that Aur-AepiΔ KCs with elevated p-EGFR and p-Retinoblastoma (p-Rb) levels were distributed in the majority of nodes in actively cycling cells (q < 0.01, Supplementary Figure 7b), whereas KCs with higher levels of p-S6 correlated with p-HH3 levels (Supplementary Figure 7b-d). These results are consistent with the p-S6 analysis from our immunostaining and immunoblotting experiments (see Figs. 3 and 4).
Inhibition of AUR-A resulted in multinucleation/polyploidy and M-phase activation of p-S6 in skin SCC cell lines
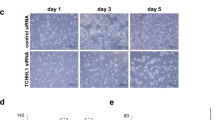
We tested the AUR-A small molecule inhibitors, VX680 and TC-A2317, in human SCC cell lines to determine whether our findings in precancerous mouse skin were preserved in squamous tumor cells. The survival of Colo16 [46] and SRB-P9 [47], and SRB-M7 [48] cells was modestly reduced by the inhibitors (Fig. 6a and Supplementary Figure 8a). We also could not detect the induction of Annexin V-associated apoptosis in drug-treated cells (Fig. 6a). Although VX680 inhibits the AURORA Kinase family and TC-A2317 is more selective for AUR-A [49], both inhibitors produced enlarged cells, indicative of mitotic arrest and slippage (Fig. 6b and Supplementary Figure 8a-b). This was also shown by the presence of drug-treated cells with greater than 4N DNA content (Fig. 6c, Supplementary Figures 8a and 9a). We confirmed the inhibitory effects of TC-A2317 in Colo16 and SRB-P9 cells by probing for the phosphorylation of AUR-A at T288, a surrogate for its activation [50]. TC-A2317 reduced the phosphorylation at T288 (Fig. 6d and Supplementary Figure 9b) and that of the AUR-A target protein TACC3 in Colo16 cells (Fig. 6d) [51].
The chemical inhibition of AUR-A attenuated cell death and induced polyploidy/multinucleation in skin carcinoma cells. a Cell viability was measured by absorbance of formazan by-products from MTS at 48 h after drug treatment, whereas apoptosis was measured by the fluorescence detection of Annexin V exposure on the cell surface at time points between 1 and 48 h. The Annexin V graphs shown are at 24 h and are representative of the results obtained over the 48 h time course. Each bars represents at least n = 8 samples per condition. b Morphological changes on Colo16 and SRB-P9 skin cancer cells after 48 h exposure to the AUR-A selective inhibitor TC-A2317. Similar effects were observed on M7 cells (Supplementary Figure 8b). Scale bar, 100 μm. c Cell cycle analysis was performed on drug-treated Colo16 cells by BrdU/PI flow cytometry. Cells were characterized 48 h after treatment. Note the appearance of greater than 4N cell populations. d Phosphorylation of AUR-A and its target protein, TACC3, was determined 8 h following drug treatment by immunoblotting. Blots were then stripped and probed for the loading control β-ACTIN. e Western blotting for p-S6 and p-AKT after 48 h of drug treatment. Note the lower levels of AKT and p-AKT detection in whole cell lysates. The graph shows the quantification of p-S6/S6 ratios from two independent experiments (n = 6 for each condition). f Colo16 cells (n = 5 independent plates per condition) were treated with TC-A2317 or vehicle for 48 h. Cell were then processed, stained, analyzed for mass cytometry. Cell cycle distribution was determined as in Fig. 5a. Pie charts values are an average of five independent plates per condition. g p-S6 was analyzed in “live” and M phase gates. Note the increase in p-S6+/p-HH3+ cells with TC-A2317 (top graphs). Bottom graph shows quantitation of p-S6 mean channel intensity (n = 5, mean ± s.d.). Data is representative of at least three independent experiments for a–d, and two independent experiments for e–g
Inhibition of mTOR/PI3K signaling reduced cell viability and enlargement of TC-A2317-treated carcinoma cells
In subsequent experiments we focused on the effects of TC-A2317 on mTOR/PI3K signaling of Colo16 and SRB-P9 cells using low doses of the inhibitor. Western blotting for p-S6 in unsynchronized cells did not reveal large changes between vehicle and TC-A2317-treated cells (Fig. 6e and Supplementary Figure 9c). Notably, the levels of p-AKT/AKT were much lower than S6 but followed the same pattern as the detection of p-S6. We then characterized the cell cycle distribution of p-S6 using mass cytometry. Similar to our previous observations, we observed an increase in G2/M cells in AUR-A-inhibited cells (Fig. 6f and Supplementary Figure 9d). Drug treated cells also showed an increased fraction of p-S6+/p-HH3+ cells and an increase in p-S6 channel intensity (Fig. 6f, g and Supplementary Figure 9e). Next, we explored the effects of the dual mTOR/PI3K inhibitor BEZ235 on TC-A2317-treated cells [52, 53]. Alone, BEZ235 suppressed cell viability (Fig. 7a, b) by reducing the fraction of S-phase cells. Colo16 cells appeared less sensitive to the effects of BEZ235 than SRB-P9 cells. BEZ235 also did not appear to induce Annexin V-associated apoptosis (Fig. 7a). In combination, TC-A2317 had a modest synergy on BEZ235-mediated inhibition of cell viability (Fig. 7a). Nevertheless, BEZ235 appeared to decrease the enlargement and multinucleation of TC-A2317-treated carcinoma cells (Fig. 7b). BEZ235 also reduced the fraction of S-phase cells induced by TC-A2317 in synchronized or unsynchronized cells (Fig. 7c and Supplementary Figure 9d, respectively). Gating on “live” cells or G2/M showed that the overall levels of p-S6+ cells were reduced by BEZ235 (Fig. 7d and Supplementary Figure 9e). This also was seen in the p-S6 mean channel intensity (Fig. 7e and Supplementary Figure 9e). However, a persistent population of M-phase cells retained the expression of p-S6 after BEZ235 treatment, which was further increased in dual inhibitor-treated cells (Fig. 7d, e and Supplementary Figure 9e). This observation suggests that there may be additional pathways that activate S6 signaling in mitosis when mTOR/PI3K signaling is impaired. Lastly, we probed for p-S6 and S6 by immunoblotting in BEZ235/TC-A2317-treated cells. The ratio of p-S6/S6 appeared elevated in dual inhibitor treated Colo16 cells, but unchanged in SRB-P9 cells (Fig. 7f and Supplementary Figure 9f). These results indicate that the flux of mTOR/PI3K signaling may depend on cellular context, but the changes observed in p-S6 expression during M-phase cannot be simply ascribed to changes in overall S6 levels.
BEZ235 attenuated the effects on cell size and polyploidy/multinucleation by AUR-A inhibition in skin carcinoma cells. a Cell viability as measured by the absorbance of Luminescent formazan by-products from metabolic active cells at 24 h. Apoptosis was measured by the fluorescence detection of Annexin V exposure on the cell surface at time points between 1 and 48 h. The Annexin V graphs shown are at 24 h and are representative of the results obtained over the 48 h time course. Each bars represents n = 8 samples per condition. b Morphological changes on synchronized Colo16 and SRB-P9 skin carcinoma cells after 48 h exposure to BEZ245, or the combination of TC-A2317 and BEZ235. Scale bar, 100 μm. c Synchronized Colo16 cells (n = 5 independent plates per condition) were treated with TC-A2317 or with the combination TCA2317/BEZ235 for 48 h. Cells were then processed, stained, and analyzed for mass cytometry. Cell cycle distribution was determined as in Fig. 5a. Pie chart values are an average of five independent plates per condition. d The pattern of p-S6 detection was analyzed in the G2/M gate of TCA2317/BEZ235-treated Colo16 cells. The fraction of p-S6+ cells using a “live” and G2/M gate decreased with TC-A2317/BEZ235. Note the persistence of p-S6+ population in M-phase (i.e., p-HH3+ cells). e The graphs show the quantification of p-S6 channel intensity in different phase of the cell cycle (n = 5, mean ± s.d.). Note the pattern of p-S6 detection was similar to that observed with SRB-P9 cells (see Supplementary Figure 9). f Western blotting for p-S6 and p-AKT after a 48 h treatment with inhibitors is shown. Note the lower levels of p-AKT and AKT detection in whole cell lysates. TC-A2317 also appears to elevate p-S6 levels in BEZ235-treated Colo16 cells. The graph shows the quantification of p-S6/S6 levels from two independent experiments (n = 6 for each condition). Data is representative of at least three independent experiments for a and b and two independent experiments for c–f
Discussion
The predominant response to Aur-A deletion in mutant p53 hyperplastic skin was delayed mitosis, mitotic survival, slippage, and multinucleation. The cell fate decision between death vs. slippage after mitotic delay remains poorly understood. Current models suggest that the interplay between the levels of CCNB1/CDK1 complex and the strength of pro-apoptotic signaling determines whether cells can slip out of a stalled mitosis or undergo catastrophic cell death. Our results would indicate that pro-apoptotic signaling was attenuated in adult Aur-AepiΔ hyperplastic skin. However, this conclusion may depend on the developmental stage. Aur-A deletion at 1–2 cell stage or during mid-gestation led to embryonic lethality [54, 55]. Similar findings were observed when Aur-A was deleted in the developing epidermis at the onset of stratification (E.12.5–E15.5) [42]. Detection of cell death declined in later embryonic stages and in adult skin [42, 56]. Lastly, we now showed that cell death due to the inhibition of AUR-A is attenuated in skin carcinoma cells.
The cell fate response to AUR-A deficiency may be inherent to adult skin. First, multinucleation/polyploidy is a feature of suprabasal skin differentiation [57, 58]. In primary human KC cultures, loss of WT p53 triggers differentiation and mitotic slippage [57]. Second, PI3K/mTOR signaling is important for skin development and tissue homeostasis [28, 29]. In particular, activation of the PI3K/AKT pathway provides survival signals and regulates CASPASE activity in differentiated KCs [30]. Third, activation of this pathway allows suprabasal KCs to survive in conditions of defective skin barrier [59]. Fourth, PI3K/AKT signaling has been shown in cultured cells to regulate the G2/M transition and may provide survival signals during normal mitosis [60, 61]. Increased AKT signaling has been shown to rescue cultured cells arrested in G2/M by DNA damage [62]. Lastly, mTOR may also regulate the G2/M transition [63]. In our study, the inhibition of mTOR/PI3K signaling by BEZ235 reduced the cell enlargement and polyploidy/multinucleation observed in AUR-A-inhibited carcinoma cells. Intriguingly, S6 remained phosphorylated in mitotic cells surviving BEZ235 exposure. This would suggest that other pathways (e.g., p90RSK) could compensate for impaired mTOR/PI3K signaling. Although the role of p-S6 in mitosis remains elusive [63,64,65], its function may be related to cell survival. Our data suggests that hyperplastic KCs expressing mutant p53 and RAS, or skin carcinoma cells can also tap into these endogenous survival pathways when faced with impaired mitosis.
To date anti-mitotic agents have given mixed results in cancer clinical trials [66]. This is likely due to the fact that new drugs are tested in xenograft models that may not faithfully represent de novo tumor formation. The GOF/LOF p53 model may be an ideal in vivo platform to test the preclinical effects of anti-mitotic drugs in combination with other inhibitors. We show that the AUR-A inhibition led to the survival of mutant p53 and KRAS KCs in polyploidy/multinucleated states; this may be an impediment to using anti-mitotic agents in skin cancer treatment. However, the use of anti-mitotic and mTOR/PI3K inhibitors may be an effective combination as recently shown for Large B-cell Lymphomas [67]. This combination may also be relevant for organ transplant patients who are at high risk of developing metastatic or recurring SCCs and whose immune systems are suppressed with the use of mTOR inhibitors [68].
In summary, we show that the induction of multinucleation/polyploidy may be a compensatory state in KCs allowing for cellular survival and maintenance of partial barrier function in face of aberrant cell division and differentiation in adult skin, which may depend on mTOR/PI3K signaling.
Material and methods
Animal models and KC isolation
The University of Colorado Institutional Animal Care and Use Committee approved all animal studies described in this manuscript. Mice with the p53f, p53lslR172H, and KrasLSLG12D knock-in alleles were crossed with the K14CreER(T) transgene to generate LOF p53 mice (p53f/f; KraslslG12D; K14CreER) or GOF p53 mice (p53f/lslR172H; KraslslG12D; K14CreER) [9, 26, 42]. A floxed Aur-A allele [42] was introduced to GOF or LOF p53 mice to generate Aur-Af/f; GOF or Aur-Af/f; LOF p53 mice. Recombination of floxed alleles in back skin was induced by feeding 8–10-week-old shaved mice with TAM containing chow (250 mg/kg weight) (Harland Laboratories) for 14 days. TPA (12-O-tetradecanoylphorbol-13-acetate, LC Laboratories, Woburn, MA) (1 nmol) was applied topically to the back skin for 7 days, in order to promote the proliferation of skin epidermal KC stem cells [20]. In experiments for mass cytometry or propidium iodide (PI) DNA content by flow cytometry, the ears of mice were treated topically with 4-hydroxyTAM (4HOT) (25 μg) for 7 days. TPA then was co-applied at 1 nmol for 7 days for PI flow cytometry or at 10 nmol for 3 days for mass cytometric experiments.
Mice studies utilized pooled cohorts from different litters and treatments on different days due to the challenge of obtaining mice with a complete set of experimental alleles (e.g., p53f/lslR172H; KraslslG12D; K14CreER). Thus, for practical reasons the sample size was determined from available genotypes and investigators were not blind to the genotypes of each mouse. Tissues were then harvested for RNA, protein, or histological analysis. Isolation of KCs for flow cytometry was performed as previously described [20]. Briefly, ear skin was floated O/N on 0.25% Trypsin (ThermoFisher, Waltham, MA) to separate epidermis from dermis. KCs were tapped out in media containing serum. For mass cytometry experiments, ear skin was floated on Dispase (Roche, Basel, Switzerland) O/N to separate epidermis from dermis. Peeled epidermis was then floated on Trypsin. Next, media with serum was added to neutralize Trypsin and dislodge KCs from the epidermis.
Cell lines and tissue culture
Colo16 cells [46] were obtained from Dr. Numsen Hail (University of Colorado Anschutz Medical Campus, Aurora, CO) and SRB-P9 and SRB-M7 cells [47, 48] were obtained from Dr. John Clifford (Louisiana State University, Shreveport, Louisiana). Cells were initially expanded and frozen down. Mycoplasma contamination was routinely tested using the Lonza MycoAlert™ PLUS Mycoplasma Detection Kit (Lonza, Basel, Switzerland). Cell line authentication was performed at the University of Colorado Cancer Center Protein Production/Mab/Tissue Culture Core using STR detection (Promega Madison, WI). Mutations in p53 in the SRB-M7 cell line was determined by Sanger Sequencing of amplicons covering hot spot exons (5–9) at the UCCC Molecular Pathology Shared Resource Core (University of Colorado Anschutz Medical campus, Aurora, CO). Mutations in the SRB-P9 and Colo16 cell lines were determined by next generation sequencing using the Archer VariantPlex Solid Tumor kit (ArcherDx, Boulder, CO) with 50 ng of tumor cell DNA. Libraries were sequenced via Illumina NextSeq (Illumina, San Diego, CA). Raw sequence data was processed for mutational calling by using the Archer Analysis software package (version 5.1.2.2; ArcherDx). Mutations were manually inspected in the Integrative Genomics Viewer (Broad Institute, Cambridge, MA) by visualizing BAM files. Using these approaches, we determined a 319Gln[stop] mutation in SRB-P9 cells and an Y205F mutation in SRB-M7 cells for p53. Colo16 cells contained WT p53 sequences but harbored an HRAS mutation (Gly13Arg).
Tissue culture experiments were performed within the last 3 years with Mycoplasm negative cells up to passage 30 from the original expansion. Colo16 cells were grown in RPMI-containing 2% FBS, whereas SRB-P9/M7 cells were grown in 10% FBS DMEM/F12 media (Thermo Fisher). Promega’s CellTiter Aqueous MTS colorimetric or RealTime-Glo MT Cell Viability assays were used to determine viability as per the manufacturer’s instructions (Promega). TC-A2317 (Tocris, Minneapolis, MN) was dissolved in ethanol at a 1 mM concentration. BEZ235 (Selleckchem, Houston, TX) and VX680 (Selleckchem.com) were dissolved in DMSO at a 1 mM concentrations. Cell synchronization was performed as previously published using medium containing Thymidine [69].
RNA and QPCR experiments
RNA from frozen biopsies was isolated as previously described [42]. For whole skin RNA sequencing (RNAseq), TruSEQ Stranded mRNA Sample Prep (Illumina) was used with 500 ng of total RNA. Barcoded samples were pooled and run across three lanes of 50 bp reads sequencing on the HiSEQ 2500 machine (Illumina). Reads quality was checked using FastQC software. Raw files were deposited in the Gene Expression Omnibus (ncbi.nlm.nih.gov/geo) under accession #GSE106281. The median number of reads per condition was 42 million. Raw reads were aligned and quantitated with Partek Flow (Partek.com) and TopHat_v2/Cufflinks_v2.02.2 [70]. Differential gene expression was determined using Partek Flow or Cuffdiff v2.0.2 [70]. Pathway analysis was performed using IPA ingenuity software (Qiagen). QPCR analysis was performed as previously described [42] using a SYBR assay (KappaBiosystems, Wilmington, MA) or with predesigned assays from IDT (idtdna.com, Skokie, IL). Assay catalog numbers and oligo sequences are listed in Supplementary Table 6.
Immunofluorescence and immunoblotting assays
Skin biopsies were fixed in 4% paraformaldehyde in PBS and processed for paraffin embedding and sectioning. Histopathological analysis was performed on H&E-stained sections. Mitotic indexes and number of apoptotic cells was determined by counting five fields per section under 1000× magnification. Nucleated cell numbers were measured from basal KCs to the start of keratinization. Immunodetection was done as previously described [42] using primary antibodies listed in Supplementary Table 7. Secondary antibodies were purchased from ThermoFisher. Sections were then mounted with DAPI-containing anti-fade solution (Vector Laboratories, Burlingame, CA) and staining visualized on a Nikon 90i conventional microscope or a C2 confocal microscope. To quantitate the p-S6 signal, we determined pixel intensity vs. surface area of the “Region of Interest” associated with the cells attached to the basal layer or cells found directly superior in the suprabasal layers. First, we determined the p-S6/S6 ratios in the basal layer and then in the suprabasal layer. As an alternative approach, we calculated the ratio of basal to suprabasal intensity in each sample to better normalize the detection of p-S6 and total S6. We compared the averaged values from WT AUR-A skin (i.e., GOF and LOF p53 skin) with that of AUR-A deficient skin (i.e., Aur-AepiΔ; GOF p53 and Aur-AepiΔ; LOF p53 skin).
For western blot analysis, cell or tissue lysates were prepared in non-denaturing Lysis Buffer 17 (R&D Systems, Minneapolis, MN) or 1× cell lysis buffer (Cell Signaling, Danvers, MA) containing IGEPAL® CA-360 and phosphatase and protease inhibitors (Roche). Protein concentrations were determined via a Bradford Assay (Biorad, Hercules, CA) and fractionated on Novex® 4–12% Bis-Tris Protein gels (ThermoFisher). Proteins on gels were then transferred onto nitrocellulose support. Membranes were probed with primary antibodies (see Supplementary Table 7) and then incubated with appropriate HRP-conjugated secondary antibodies. Chemiluminescent signal was detected using Supersignal Dura or Femto substrate (ThermoFisher). Membranes were then stripped and reprobed for β-ACTIN (SantaCruz). Images were either captured on film or with an Odyssey Fc imaging system (LI-COR Biosciences, Lincoln, Nebraska). Images on film were digitized and band intensity determined by measuring mean adjusted pixel density using ImageJ software (imagej.nih.gov) [71]. Quantification for westerns captured on the Odyssey system was performed with Image studio software (LI-COR).
Flow and mass cytometry
Freshly isolated KCs were fixed in PBS/ethanol and then re-suspended with PI solution. Fluorescence was evaluated on a Beckman Coulter Gallios cell analyzer for DNA content analysis. BromodeoxyUridine (BrdU) incorporation and detection was performed as previously described [72]. Cell preparation, antibody staining, and fixation for mass cytometry was performed with reagents and protocols from Fluidigm Corp (Fluidigm.com) and from previous reports [43, 44]. Barcoding of samples for tissue culture experiments was performed as per manufacturer instructions (Fluidigm) and from published protocols [73]. Samples were analyzed using a Helios system and raw data either normalized using the Helios software (Fluidigm) or through software available from the Nolan Lab (http://web.stanford.edu/group/nolan/resources.html). Normalized data was visualized and gated using Cytobank (cytobank.org). Citrus clustering analyses were performed on Cytobank.
Data and statistical analyses
Mice studies were repeated independently at least twice. Cell culture studies were repeated independently at least three times unless otherwise stated. Statistical ratios and analyses were performed with Partek Genomic Flow, IPA, Cytobank or GraphPad Prism (v6) software. For simple comparisons of means between two genotypes or treatments (e.g., Vehicle vs Drug treatment), an unpaired double-sided t-test was performed. We assumed equal standard deviation in each of the sample populations. The subsequent “P” value was used to evaluate the likelihood of an actual difference between the comparison groups. P-values are denoted in graphs by asterisks: *P = 0.01–0.05; **P = 0.001–0.01; ***P = 0.0001–0.001; ****P < 0.0001. An adjusted P-value (q value) was calculated using the Storey method [74] in Partek Genomic Flow software to account for false discovery in the analysis of RNAseq data. A cutoff of q < 0.05 was used for gene expression comparisons. Core facilities staff (e.g., Flow Core) processed samples blindly in RNAseq or cytometric experiments.
References
Karia PS, Han J, Schmults CD. Cutaneous squamous cell carcinoma: estimated incidence of disease, nodal metastasis, and deaths from disease in the United States, 2012. J Am Acad Dermatol. 2013;68:957–66.
Nassar D, Latil M, Boeckx B, Lambrechts D, Blanpain C. Genomic landscape of carcinogen-induced and genetically induced mouse skin squamous cell carcinoma. Nat Med. 2015;21:946–54.
South AP, Purdie KJ, Watt SA, Haldenby S, den Breems NY, Dimon M, et al. NOTCH1 mutations occur early during cutaneous squamous cell carcinogenesis. J Invest Dermatol. 2014;134:2630–8.
Yilmaz AS, Ozer HG, Gillespie JL, Allain DC, Bernhardt MN, Furlan KC, et al. Differential mutation frequencies in metastatic cutaneous squamous cell carcinomas versus primary tumors. Cancer. 2016;123:1184–93.
Pierceall WE, Mukhopadhyay T, Goldberg LH, Ananthaswamy HN. Mutations in the p53 tumor suppressor gene in human cutaneous squamous cell carcinomas. Mol Carcinog. 1991;4:445–9.
Brash DE, Rudolph JA, Simon JA, Lin A, McKenna GJ, Baden HP, et al. A role for sunlight in skin cancer: UV-induced p53 mutations in squamous cell carcinoma. Proc Natl Acad Sci USA. 1991;88:10124–8.
Pickering CR, Zhou JH, Lee JJ, Drummond JA, Peng SA, Saade RE, et al. Mutational landscape of aggressive cutaneous squamous cell carcinoma. Clin Cancer Res. 2014;20:6582–92.
Lang GA, Iwakuma T, Suh YA, Liu G, Rao VA, Parant JM, et al. Gain of function of a p53 hot spot mutation in a mouse model of Li-Fraumeni syndrome. Cell. 2004;119:861–72.
Olive KP, Tuveson DA, Ruhe ZC, Yin B, Willis NA, Bronson RT, et al. Mutant p53 gain of function in two mouse models of Li-Fraumeni syndrome. Cell. 2004;119:847–60.
Nelson MA, Einspahr JG, Alberts DS, Balfour CA, Wymer JA, Welch KL, et al. Analysis of the p53 gene in human precancerous actinic keratosis lesions and squamous cell cancers. Cancer Lett. 1994;85:23–9.
Jonason AS, Kunala S, Price GJ, Restifo RJ, Spinelli HM, Persing JA, et al. Frequent clones of p53-mutated keratinocytes in normal human skin. Proc Natl Acad Sci USA. 1996;93:14025–9.
Kanellou P, Zaravinos A, Zioga M, Stratigos A, Baritaki S, Soufla G, et al. Genomic instability, mutations and expression analysis of the tumour suppressor genesp14(ARF), p15(INK4b), p16(INK4a) and p53 in actinic keratosis. Cancer Lett. 2008;264:145–61.
Pierceall WE, Goldberg LH, Tainsky MA, Mukhopadhyay T, Ananthaswamy HN. Ras gene mutation and amplification in human nonmelanoma skin cancers. Mol Carcinog. 1991;4:196–202.
Spencer JM, Kahn SM, Jiang W, DeLeo VA, Weinstein IB. Activated ras genes occur in human actinic keratoses, premalignant precursors to squamous cell carcinomas. Arch Dermatol. 1995;131:796–800.
Dajee M, Lazarov M, Zhang JY, Cai T, Green CL, Russell AJ, et al. NF-kappaB blockade and oncogenic Ras trigger invasive human epidermal neoplasia. Nature. 2003;421:639–43.
Miyamoto H, Harada M, Isobe H, Akita HD, Haneda H, Yamaguchi E, et al. Prognostic value of nuclear DNA content and expression of the ras oncogene product in lung cancer. Cancer Res. 1991;51:6346–50.
Lu SL, Herrington H, Reh D, Weber S, Bornstein S, Wang D, et al. Loss of transforming growth factor-beta type II receptor promotes metastatic head-and-neck squamous cell carcinoma. Genes Dev. 2006;20:1331–42.
Nikonova AS, Astsaturov I, Serebriiskii IG, Dunbrack RL Jr., Golemis EA. Aurora A kinase (AURKA) in normal and pathological cell division. Cell Mol Life Sci. 2013;70:661–87.
Ewart-Toland A, Dai Q, Gao YT, Nagase H, Dunlop MG, Farrington SM, et al. Aurora-A/STK15 T + 91A is a general low penetrance cancer susceptibility gene: a meta-analysis of multiple cancer types. Carcinogenesis. 2005;26:1368–73.
Torchia EC, Chen Y, Sheng H, Katayama H, Fitzpatrick J, Brinkley WR, et al. A genetic variant of Aurora kinase A promotes genomic instability leading to highly malignant skin tumors. Cancer Res. 2009;69:7207–15.
Wang X, Zhou YX, Qiao W, Tominaga Y, Ouchi M, Ouchi T, et al. Overexpression of aurora kinase A in mouse mammary epithelium induces genetic instability preceding mammary tumor formation. Oncogene. 2006;25:7148–58.
Mobley A, Zhang S, Bondaruk J, Wang Y, Majewski T, Caraway NP, et al. Aurora kinase A is a biomarker for bladder cancer detection and contributes to its aggressive behavior. Sci Rep. 2017;7:40714.
Zhou H, Kuang J, Zhong L, Kuo WL, Gray JW, Sahin A, et al. Tumour amplified kinase STK15/BTAK induces centrosome amplification, aneuploidy and transformation. Nat Genet. 1998;20:189–93.
Liu G, McDonnell TJ, Montes de Oca Luna R, Kapoor M, Mims B, El-Naggar AK, et al. High metastatic potential in mice inheriting a targeted p53 missense mutation. Proc Natl Acad Sci USA. 2000;97:4174–9.
Torchia EC, Caulin C, Acin S, Terzian T, Kubick BJ, Box NF, et al. Myc, Aurora Kinase A, and mutantp53(R172H) co-operate in a mouse model of metastatic skin carcinoma. Oncogene. 2012;31:2680–90.
Caulin C, Nguyen T, Lang GA, Goepfert TM, Brinkley BR, Cai WW, et al. An inducible mouse model for skin cancer reveals distinct roles for gain- and loss-of-function p53 mutations. J Clin Invest. 2007;117:1893–901.
Hertzler-Schaefer K, Mathew G, Somani AK, Tholpady S, Kadakia MP, Chen Y, et al. Pten loss induces autocrine FGF signaling to promote skin tumorigenesis. Cell Rep. 2014;6:818–26.
Ding X, Bloch W, Iden S, Ruegg MA, Hall MN, Leptin M, et al. mTORC1 and mTORC2 regulate skin morphogenesis and epidermal barrier formation. Nat Commun. 2016;7:13226.
Dainichi T, Hayden MS, Park SG, Oh H, Seeley JJ, Grinberg-Bleyer Y, et al. PDK1 is a regulator of epidermal differentiation that activates and organizes asymmetric cell division. Cell Rep. 2016;15:1615–23.
Janes SM, Ofstad TA, Campbell DH, Eddaoudi A, Warnes G, Davies D, et al. PI3-kinase-dependent activation of apoptotic machinery occurs on commitment of epidermal keratinocytes to terminal differentiation. Cell Res. 2009;19:328–39.
Yuspa SH, Hennings H, Lichti U. Initiator and promoter induced specific changes in epidermal function and biological potential. J Supramol Struct Cell Biochem. 1981;17:245–57.
Zhang L, Ferreyros M, Feng W, Hupe M, Crumrine DA, Chen J, et al. Defects in stratum corneum desquamation are the predominant effect of impaired ABCA12 function in a novel mouse model of harlequin ichthyosis. PLoS ONE. 2016;11:e0161465.
Cottle DL, Ursino GM, Ip SC, Jones LK, Ditommaso T, Hacking DF, et al. Fetal inhibition of inflammation improves disease phenotypes in harlequin ichthyosis. Hum Mol Genet. 2015;24:436–49.
Oestreicher P. 4th Annual pharmacogenomics and medicine lectures. Pharmacogenomics. 2001;2:291–6.
Steven AC, Bisher ME, Roop DR, Steinert PM. Biosynthetic pathways of filaggrin and loricrin–two major proteins expressed by terminally differentiated epidermal keratinocytes. J Struct Biol. 1990;104:150–62.
Brito DA, Rieder CL. Mitotic checkpoint slippage in humans occurs via cyclin B destruction in the presence of an active checkpoint. Curr Biol. 2006;16:1194–1200.
Vader G, Medema RH, Lens SM. The chromosomal passenger complex: guiding Aurora-B through mitosis. J Cell Biol. 2006;173:833–7.
Gruss OJ, Wittmann M, Yokoyama H, Pepperkok R, Kufer T, Sillje H, et al. Chromosome-induced microtubule assembly mediated by TPX2 is required for spindle formation in HeLa cells. Nat Cell Biol. 2002;4:871–9.
Sarbassov DD, Guertin DA, Ali SM, Sabatini DM. Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science. 2005;307:1098–101.
Fingar DC, Salama S, Tsou C, Harlow E, Blenis J. Mammalian cell size is controlled by mTOR and its downstream targets S6K1 and 4EBP1/eIF4E. Genes Dev. 2002;16:1472–87.
Aksamitiene E, Kholodenko BN, Kolch W, Hoek JB, Kiyatkin A. PI3K/Akt-sensitive MEK-independent compensatory circuit of ERK activation in ER-positive PI3K-mutant T47D breast cancer cells. Cell Signal. 2010;22:1369–78.
Torchia EC, Zhang L, Huebner AJ, Sen S, Roop DR. Aurora kinase-A deficiency during skin development impairs cell division and stratification. J Invest Dermatol. 2013;133:78–86.
Behbehani GK, Bendall SC, Clutter MR, Fantl WJ, Nolan GP. Single-cell mass cytometry adapted to measurements of the cell cycle. Cytometry A. 2012;81:552–66.
Bruggner RV, Bodenmiller B, Dill DL, Tibshirani RJ, Nolan GP. Automated identification of stratifying signatures in cellular subpopulations. Proc Natl Acad Sci USA. 2014;111:E2770–7.
Dephoure N, Zhou C, Villen J, Beausoleil SA, Bakalarski CE, Elledge SJ, et al. A quantitative atlas of mitotic phosphorylation. Proc Natl Acad Sci USA. 2008;105:10762–7.
Moore GE, Merrick SB, Woods LK, Arabasz NM. A human squamous cell carcinoma cell line. Cancer Res. 1975;35:2684–8.
Clifford JL, Yang X, Walch E, Wang M, Lippman SM. Dominant negative signal transducer and activator of transcription 2 (STAT2) protein: stable expression blocks interferon alpha action in skin squamous cell carcinoma cells. Mol Cancer Ther. 2003;2:453–9.
Hail N Jr, Lotan R. Examining the role of mitochondrial respiration in vanilloid-induced apoptosis. J Natl Cancer Inst. 2002;94:1281–92.
Min YH, Kim W, Kim JE. The Aurora kinase A inhibitor TC-A2317 disrupts mitotic progression and inhibits cancer cell proliferation. Oncotarget. 2016;7:84718–35.
Eyers PA, Erikson E, Chen LG, Maller JL. A novel mechanism for activation of the protein kinase Aurora A. Curr Biol. 2003;13:691–7.
Kinoshita K, Noetzel TL, Pelletier L, Mechtler K, Drechsel DN, Schwager A, et al. Aurora A phosphorylation of TACC3/maskin is required for centrosome-dependent microtubule assembly in mitosis. J Cell Biol. 2005;170:1047–55.
Maira SM, Stauffer F, Brueggen J, Furet P, Schnell C, Fritsch C, et al. Identification and characterization of NVP-BEZ235, a new orally available dual phosphatidylinositol 3-kinase/mammalian target of rapamycin inhibitor with potent in vivo antitumor activity. Mol Cancer Ther. 2008;7:1851–63.
Serra V, Markman B, Scaltriti M, Eichhorn PJ, Valero V, Guzman M, et al. NVP-BEZ235, a dual PI3K/mTOR inhibitor, prevents PI3K signaling and inhibits the growth of cancer cells with activating PI3K mutations. Cancer Res. 2008;68:8022–30.
Sasai K, Parant JM, Brandt ME, Carter J, Adams HP, Stass SA, et al. Targeted disruption of Aurora A causes abnormal mitotic spindle assembly, chromosome misalignment and embryonic lethality. Oncogene. 2008;27:4122–7.
Cowley DO, Rivera-Perez JA, Schliekelman M, He YJ, Oliver TG, Lu L, et al. Aurora-A kinase is essential for bipolar spindle formation and early development. Mol Cell Biol. 2009;29:1059–71.
Perez de Castro I, Aguirre-Portoles C, Fernandez-Miranda G, Canamero M, Cowley DO, Van Dyke T, et al. Requirements for Aurora-A in tissue regeneration and tumor development in adult mammals. Cancer Res. 2013;73:6804–15.
Freije A, Molinuevo R, Ceballos L, Cagigas M, Alonso-Lecue P, Rodriguez R, et al. Inactivation of p53 in human keratinocytes leads to squamous differentiation and shedding via replication stress and mitotic slippage. Cell Rep. 2014;9:1349–60.
Zanet J, Freije A, Ruiz M, Coulon V, Sanz JR, Chiesa J, et al. A mitosis block links active cell cycle with human epidermal differentiation and results in endoreplication. PLoS ONE. 2010;5:e15701.
Yanagi T, Akiyama M, Nishihara H, Miyamura Y, Sakai K, Tanaka S, et al. AKT has an anti-apoptotic role in ABCA12-deficient keratinocytes. J Invest Dermatol. 2011;131:1942–5.
Dangi S, Cha H, Shapiro P. Requirement for phosphatidylinositol-3 kinase activity during progression through S-phase and entry into mitosis. Cell Signal. 2003;15:667–75.
Lee SR, Park JH, Park EK, Chung CH, Kang SS, Bang OS. Akt-induced promotion of cell-cycle progression at G2/M phase involves upregulation of NF-Y binding activity in PC12 cells. J Cell Physiol. 2005;205:270–7.
Kandel ES, Skeen J, Majewski N, Di Cristofano A, Pandolfi PP, Feliciano CS, et al. Activation of Akt/protein kinase B overcomes a G(2)/m cell cycle checkpoint induced by DNA damage. Mol Cell Biol. 2002;22:7831–41.
Ramirez-Valle F, Badura ML, Braunstein S, Narasimhan M, Schneider RJ. Mitotic raptor promotes mTORC1 activity, G(2)/M cell cycle progression, and internal ribosome entry site-mediated mRNA translation. Mol Cell Biol. 2010;30:3151–64.
Egervari G, Mark A, Hajdu M, Barna G, Sapi Z, Krenacs T, et al. Mitotic lymphoma cells are characterized by high expression of phosphorylated ribosomal S6 protein. Histochem Cell Biol. 2011;135:409–17.
Schmidt T, Wahl P, Wuthrich RP, Vogetseder A, Picard N, Kaissling B, et al. Immunolocalization of phospho-S6 kinases: a new way to detect mitosis in tissue sections and in cell culture. Histochem Cell Biol. 2007;127:123–9.
Pitts TM, Davis SL, Eckhardt SG, Bradshaw-Pierce EL. Targeting nuclear kinases in cancer: development of cell cycle kinase inhibitors. Pharmacol Ther. 2014;142:258–69.
Islam S, Qi W, Morales C, Cooke L, Spier C, Weterings E, et al. Disruption of aneuploidy and senescence induced by aurora inhibition promotes intrinsic apoptosis in double hit or double expressor diffuse large B-cell lymphomas. Mol Cancer Ther. 2017;16:2083–93.
Ulrich C, Arnold R, Frei U, Hetzer R, Neuhaus P, Stockfleth E. Skin changes following organ transplantation: an interdisciplinary challenge. Dtsch Arztebl Int. 2014;111:188–94.
Bostock CJ, Prescott DM, Kirkpatrick JB. An evaluation of the double thymidine block for synchronizing mammalian cells at the G1-S border. Exp Cell Res. 1971;68:163–8.
Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25:1105–11.
Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9:671–5.
Terzian T, Torchia EC, Dai D, Robinson SE, Murao K, Stiegmann RA, et al. p53 prevents progression of nevi to melanoma predominantly through cell cycle regulation. Pigment Cell Melanoma Res. 2010;23:781–94.
Zunder ER, Finck R, Behbehani GK, Amir el AD, Krishnaswamy S, Gonzalez VD, et al. Palladium-based mass tag cell barcoding with a doublet-filtering scheme and single-cell deconvolution algorithm. Nat Protoc. 2015;10:316–33.
Storey JD. The positive false discovery rate: a Bayesian interpretation and the q-value. Ann Stat. 2003;31:2013–35.
Acknowledgements
A Dermatology Foundation Research Career Development Award and an American Cancer Association Institutional Research grant #57-001-53 to E.C. Torchia supported this work. Additional support came from the Department of Dermatology and the Gates Center for Regenerative Medicine at the University of Colorado. Weston K. Ryan received both a University of Colorado Cancer Center (UCCC) summer student fellowship and a Gates Center Summer Internship award. We also acknowledge the University of Colorado Skin Disease Center Morphology and Phenotyping Cores (NIAMS P30 AR057212), the UCCC Genomic, and the Protein Production/Mab/Tissue Culture Facilities, the Molecular Pathology Shared Resource Cores for technical assistance (P30CA046934). We are especially grateful to Karen Helm, Christine Childs and the UCCC FACS core for their expert advice and technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Edited by: G. Del Sal
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Ryan, W.K., Fernandez, J., Peterson, M.K. et al. Activation of S6 signaling is associated with cell survival and multinucleation in hyperplastic skin after epidermal loss of AURORA-A Kinase. Cell Death Differ 26, 548–564 (2019). https://doi.org/10.1038/s41418-018-0167-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-018-0167-7
This article is cited by
-
Polyploidy and mTOR signaling: a possible molecular link
Cell Communication and Signaling (2024)
-
Divide and conquer: two stem cell populations in squamous epithelia, reserves and the active duty forces
International Journal of Oral Science (2019)