Abstract
Peptides/small proteins, encoded by noncanonical open reading frames (ORF) of previously claimed non-coding RNAs, have recently been recognized possessing important biological functions, but largely uncharacterized. 1p36 is an important tumor suppressor gene (TSG) locus frequently deleted in multiple cancers, with critical TSGs like TP73, PRDM16, and CHD5 already validated. Our CpG methylome analysis identified a silenced 1p36.3 gene KIAA0495, previously thought coding long non-coding RNA. We found that the open reading frame 2 of KIAA0495 is actually protein-coding and translating, encoding a small protein SP0495. KIAA0495 transcript is broadly expressed in multiple normal tissues, but frequently silenced by promoter CpG methylation in multiple tumor cell lines and primary tumors including colorectal, esophageal and breast cancers. Its downregulation/methylation is associated with poor survival of cancer patients. SP0495 induces tumor cell apoptosis, cell cycle arrest, senescence and autophagy, and inhibits tumor cell growth in vitro and in vivo. Mechanistically, SP0495 binds to phosphoinositides (PtdIns(3)P, PtdIns(3,5)P2) as a lipid-binding protein, inhibits AKT phosphorylation and its downstream signaling, and further represses oncogenic AKT/mTOR, NF-κB, and Wnt/β-catenin signaling. SP0495 also regulates the stability of autophagy regulators BECN1 and SQSTM1/p62 through modulating phosphoinositides turnover and autophagic/proteasomal degradation. Thus, we discovered and validated a 1p36.3 small protein SP0495, functioning as a novel tumor suppressor regulating AKT signaling activation and autophagy as a phosphoinositide-binding protein, being frequently inactivated by promoter methylation in multiple tumors as a potential biomarker.
Similar content being viewed by others
Introduction
Chromosome 1p36 deletion has been well defined as a common event for multiple malignancies originating from neurons, epithelium, and hematopoietic cells [1]. Through classic genetic studies of this locus, a potent tumor-suppressive region corresponding to 1p36 was defined [2]. Particularly, 1p36.3, which is located at the distal end of 1p, is a tumor suppressor gene (TSG) hotspot or “cancer-gene island” [1,2,3]. Several bona fide TSGs have been identified at this small region, including TP73 [4], CHD5 [5], PRDM16 [6], and AJAP1 [7]. These TSGs are involved in tumor initiation and progression and frequently inactivated by either genetic mutations and/or promoter methylation in various tumors.
Long non-coding RNA (lncRNA) has previously been considered to be non-coding as there was a lack of long/obvious open-reading frames (ORFs). Recently with the advancement in deep sequencing, mass spectrometry and bioinformatics techniques [8,9,10], a subset of lncRNAs with non-canonical small ORFs and high evolutionary conservation has been shown to encode functionally important peptides or small proteins [11,12,13], although the vast majority of them still remain uncharacterized. Some peptides/proteins encoded by non-canonical ORFs of lncRNAs have been reported to possess important oncogenic or tumor-suppressive functions [11, 14], regulating cell proliferation and invasion/metastasis of tumor cells. For examples, the peptides/small proteins encoded by lncRNAs HOXB-AS3 [15], LOC90024 [16], LINC00266-1 [17], suppress colorectal tumor cell growth and metastasis; the circular form of LINC-PINT encodes an 87-aa peptide that inhibits cell proliferation, stemness and enhances DNA damage response in glioblastoma [18]; an ERα-regulated polypeptide ASRPS encoded by LINC00908 is downregulated in triple-negative breast cancer (TNBC) and associated with its poor survival [19]. ASRPS inhibits the angiogenesis of breast tumor cells through the STAT3-VEGF pathway, also as a potential biomarker for TNBC [19]. These studies demonstrate the novel functions and importance of lncRNA-encoded peptides/proteins in human disease pathogenesis.
Autophagy is a highly conserved, regular cellular degradation pathway that targets multiple proteins and removes damaged organelles by lysosomal degradation, and thus plays a crucial role in the maintenance of normal cellular homeostasis and survival [20, 21]. Autophagy is also involved in disease pathogenesis and progression of multiple malignancies, due to the genetic/epigenetic disruption of autophagy regulators [22,23,24]. Loss of Beclin 1/BECN1, a critical regulator of autophagy, through LOH and promoter CpG methylation, has been detected in multiple cancers [25]. On the other hand, SQSTM1/p62, another autophagy adapter and marker, is overexpressed in multiple cancers and significantly associated with aggressive features (distant metastasis, poor disease-free survival) and advanced stages [26, 27]. In certain contexts, induction of autophagy has been shown as a mechanism of tumor suppression, in addition to apoptosis, senescence, and cell cycle regulation [28]. Therefore, autophagy dysregulation frequently contributes to cancer pathogenesis and progression.
In this study, through integrative cancer epigenomics, we identified a 1p36.3 lncRNA KIAA0495 (previously also named as PDAM or TP73-AS1), as a methylated target in multiple tumors. KIAA0495 was frequently silenced by promoter CpG methylation in multiple tumors which could serve as a potential biomarker. We further discovered that the ORF2 of KIAA0495 encoded a small protein (thus named as SP0495), that could be detected both in vivo and in vitro. SP0495 is involved in regulating tumor cell proliferation, apoptosis, autophagy and senescence. We also found that several key oncogenic signaling pathways, including PI3K/AKT/mTOR, NF-κB, and Wnt/β-catenin signaling, were regulated by SP0495. Moreover, we demonstrate that SP0495, as a phosphoinositide-binding protein, represses AKT phosphorylation and downstream signaling activation, and further promotes autophagy via enhancing BECN1 stability to suppress tumorigenesis.
Results
Identification of KIAA0495 as a 1p36.3 transcript silenced in tumors through integrative epigenomics
We performed CpG methylome analysis of multiple tumor cell lines of colorectal, gastric, head and neck and breast, as well as immortalized normal epithelial cells by methylated DNA immunoprecipitation (MeDIP). We detected enriched methylated signals in the promoter region of a previously claimed lncRNA KIAA0495 (NR_033711) in tumor cells of colorectal (HCT116), gastric (SNU719), head and neck (C666-1), and breast (MB231 and MCF7), but not in immortalized mammary epithelial cells (HMEpC) and HCT116 cells with genetic double knockout of DNA methyltransferase DNMT1 and DNMT3B (DKO) (Fig. 1A). Meanwhile independently, using 1-Mb array comparative genomic hybridization (aCGH), we refined a 1p36.3 hemizygous deletion containing KIAA0495 in head and neck (HONE1), and esophageal (HKESC1) carcinoma cell lines (Suppl. Fig. S1A). Gene copy number analysis using the Oncomine database also revealed that KIAA0495 copy number was reduced in multiple tumors compared to normal tissues (Table S1).
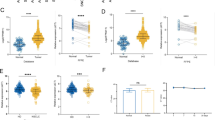
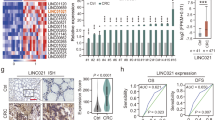
A CpG methylome analysis by MeDIP-chip detected signal enrichment in the KIAA0495 promoter CGI in carcinoma cell lines of colorectal (CRC), gastric (GsCa), head and neck (HNC), breast (BrCa) and immortalized normal mammary cell line HMEpC. Positive signal peaks (red) and the exon 1 of KIAA0495 are marked. B Schematic diagram of KIAA0495 transcript variants. Blank boxes represent exons gapped with lines (introns), blue boxes represent the predicted largest ORF2 of KIAA0495. Positions of RT-PCR primers (F/R) are denoted by arrows. C Broad expression of KIAA0495 in human normal adult tissues and fetal tissues by semi-quantitative RT-PCR, with GAPDH as a control. Sk. muscle, skeletal muscle. D Average β values of all CpG sites located at the KIAA0495 promoter were plotted for CRC, ESCA, and BRCA primary tumors and adjacent normal samples from the TCGA database. Statistical analysis is performed by one-way ANOVA. E KIAA0495 expression levels in different tumor types from TCGA database, as determined by TIMER2.0 (*, p < 0.05; ***, p < 0.001). F Analyses of TCGA datasets reveal an inverse correlation between KIAA0495 mRNA expression and its promoter methylation in CRC, ESCA, and BRCA. Each green circle represents a single clinical sample. Pearson correlation coefficient analysis is used. (G) Kaplan-Meier curve analyses show the overall survival of patients with colorectal, esophageal, lung and breast cancers in GSE and TCGA datasets by log-rank test.
KIAA0495 contains 5 transcripts, with transcript 1 (NR_033711.1) as the longest. First submitted to NCBI database by Kazusa DNA Research Institute (Fig. 1B), transcript 1 shares the same exon 1 and 2 with transcripts 2, 3, 4, thus from the same promoter. Although as a novel transcript in the immediate vicinity of TP73, KIAA0495 has only 218-bp complementary sequence homologous to the TP73 transcript in its C terminal, which was previously renamed as TP73-AS1 (Suppl Fig. S1B). We further examined KIAA0495 expression in normal human tissues using primers targeting all transcript isoforms by semi-quantitative RT-PCR, and detected its ubiquitous but variable expression in human normal adult and fetal tissues (Fig. 1C). We then retrieved KIAA0495 methylation data of colorectal, esophageal, and breast samples from the Cancer Genome Atlas (TCGA) database, and found significantly higher methylation levels of KIAA0495 in these tumor tissues compared to normal tissues (Fig. 1D). KIAA0495 expression was significantly downregulated in TCGA cohorts of multiple cancers including colorectal, gastric, esophageal, lung and breast (Fig. 1E). Further analyses of Gene Expression (SAGE), GENT2, and Oncomine databases also showed KIAA0495 downregulation in a panel of tumor tissues including digestive, head and neck, and breast cancers (Table S2, Suppl. Fig. S2). Moreover, KIAA0495 expression was negatively correlated with its promoter methylation in colorectal, esophageal and breast cancer tissues in TCGA Methylation450K datasets (Fig. 1F).
We further found that decreased expression of KIAA0495 was associated with higher-grade tumors in colorectal, breast, and bladder cancer patients (Table S3, Suppl. Fig. S2C), and poor survival of patients with colorectal, esophageal, lung, breast, bladder, and ovarian cancers (Fig. 1G, Suppl. Fig. S3). These results suggest that silencing of the 1p36.3 transcript KIAA0495, through epigenetic abnormalities, is common during tumorigenesis and associated with poor survival of cancer patients of multiple tissue origins.
KIAA0495 encodes an endogenously expressed small protein SP0495
Although KIAA0495 was previously annotated as a lncRNA, using ORFfinder, we discovered a 606-nucleotide non-canonical ORF2, which is also the largest ORF with coding potential present in transcripts 1–4, encoding a 201-aa small protein (Fig. 2A, B). Exploring ribosome profiling data using GWIPS-viz shows that KIAA0495-ORF2 is a transcriptionally activated region in human cell lines and tissues studied (Fig. 2B). We thus termed this small protein encoded by KIAA0495-ORF2 as SP0495 (small protein of KIAA0495) hereafter.
A Predicted ORFs within the transcript 1 of KIAA0495 (NR_033711.1). The largest ORF - ORF2 with protein-coding potential is marked with a green arrow. As shown in the bottom panel, there is a stop codon (TGA) located right upstream of the start codon (ATG) of ORF2. B Genomic location of the predicted common ORF2 at the four KIAA0495 transcripts. Boxes represent exons gapped with lines (introns), and filled green boxes represent ORF2. RNA-seq (green) and Ribo-seq (red) coverage plots at the KIAA0495 locus from aggregated human data (GWIPS-viz) are shown. Locations of elongating ribosomes (red) generated are consistent with the coding regions of KIAA0495-ORF2. The isolated peaks of ribosome profiling density represent ribosome stalling sites. C Diagram of the IRES fusion constructs of KIAA0495-ORF2. TGA-KIAA0495-ORF2 represents a construct with an additional 5′-UTR fragment added to the upstream of KIAA0495-ORF2 (upper panel). In vitro protein translation of KIAA0495-ORF2 yields a small protein SP0495 at the predicted size (~21 kDa). Asterisks indicate bands of the SP0495 protein at the correct size (lower panel). D Western blot detects ectopic expression of KIAA0495-ORF2/SP0495 protein in HCT116 and KYSE150 cells. α-Tubulin was used as a loading control. Representative data are shown. E Endogenous SP0495 protein is detected in multiple human normal tissues by Western blot using an anti-SP0495 antibody. Representative data are shown. F Immunofluorescence showing the subcellular localization of SP0495 in HONE1 (endogenous) and KYSE150 (exogenous) cells, using anti-SP0495 antibody (Ab) or anti-Flag antibody (green). Nuclei counterstained with DAPI (blue). Original magnification, 400×. Scale bar 10 µM. HONE1 photos were taken by confocal miscroscopy. Representative images are shown. G Co-localization of SP0495 with ER/Golgi compartments. Diagram of the SP0495 protein domain analyzed by bioinformatics is shown on the top panel. Blue box represents the location of a transmembrane & signal peptide domain (1-18-aa). KYSE150 cells with SP0495 expression were co-transfected with either ER- or Golgi-specific plasmid (pDsRed-ER or pEYFP-Golgi, respectively). Nuclei counterstained with DAPI (blue). Original magnification, 400×. Scale bar 10 µM. Representative images are shown. H Representative IHC images of SP0495 expression in gastric and colorectal tumors and adjacent normal tissues. SP0495 expression level is lower in gastric and colorectal tumor tissues compared to their adjacent normal tissues, with statistical significance by Student’s two-tailed unpaired t test analysis (p < 0.001).
We then generated Flag-fused constructs containing ORF2, with or without a 5′-UTR stop codon (TGA) to confirm the coding potential of the predicted ORF2 (Fig. 2B, C), as a 5′ TGA stop codon is located immediately in front of the ATG codon of ORF2. Results of in vitro protein expression system showed that KIAA0495-ORF2/SP0495 could be translated to protein products at the predicted size ~21 kDa, using both anti-Flag and anti-KIAA0495 antibodies (Origene, clone ID: OTI1F8) (Fig. 2C), indicating that the presence of a 5’-TGA stop codon immediately in front of the ATG codon of ORF2 does not affect its translation efficiency. Furthermore, in ectopically expressing cell line systems, abundant expression of KIAA0495-ORF2/SP0495 protein was consistently detected in two different cell lines (Fig. 2D). We also examined the endogenous expression of SP0495 in normal human tissues, and detected the natural, endogenous presence of SP0495 at the same size with ectopically expressed SP0495 in a panel of human normal tissues using an anti-KIAA0495 antibody, except for brain tissue (Fig. 2E). Bioinformatics alignment showed that human KIAA0495-ORF2/SP0495 protein shares high similarity with that of chimpanzee and monkey, with no homolog identified in other species (Suppl. Fig. S4A), indicating that it is likely a primate-specific protein.
We further evaluated the subcellular localization of SP0495. Indirect immunostaining revealed that SP0495 is located predominantly in cell cytoplasm in SP0495-endogenously or -exogenously expressed cells (Fig. 2F). Bioinformatics analysis showed that SP0495 contains a transmembrane and signal peptide domain at its N-terminal (1-18-aa) (Fig. 2G). We thus used ER- or Golgi-specific fluorescent plasmid to evaluate colocalization of SP0495 with ER or Golgi in cells, and found that SP0495 was partially localized with ER but completely colocalized with Golgi (Fig. 2G).
Endogenous SP0495 protein expression was further examined in gastric and colorectal tumor tissue samples and adjacent normal tissue samples by immunohistochemistry (IHC) (Fig. 2H). SP0495 expression levels were significantly higher in adjacent non-tumor tissues than gastric and colorectal tumor tissues (Fig. 2H), and upregulation of SP0495 protein levels was correlated with decreased tumor size in colorectal tumor tissues (Table S4). Thus, the above results demonstrate that KIAA0495-ORF2 indeed encodes a small protein SP0495, which is endogenously expressed but downregulated in multiple tumor tissues.
Promoter methylation of KIAA0495 leads to its silencing in tumor cell lines
We next investigated the expression levels and regulatory mechanisms of KIAA0495 in tumor cells. KIAA0495 promoter contains a typical CpG island spanning its transcription start site to exon 1 (Fig. 3A), suggesting that it should be regulated by promoter CpG methylation. We examined KIAA0495 expression and promoter methylation in multiple tumor cell lines. Semi-quantitative RT-PCR showed that, in contrast to its abundant expression in normal tissues (Fig. 1C) and normal immortalized cell lines (Fig. 3B), KIAA0495 was frequently silenced or downregulated in tumor cell lines of colorectal (11/11), gastric (7/16), head and neck (3/4), esophageal (11/17), breast (4/9), bladder (2/3), cervical (2/4) cancers, as well as non-Hodgkin and Hodgkin lymphomas (12/23) (Fig. 3B, Suppl. Fig. S4B). Similar results were confirmed by real-time RT-PCR at the RNA level and Western blot at the protein level (Suppl. Fig. S4C, Fig. 3C). KIAA0495 silencing/downregulation was uncommon in other cell lines of lung, liver, renal, ovarian, prostate and brain tumors (Suppl. Fig. S4B).
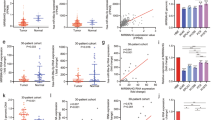
A A typical CpG island (CGI) spans the promoter to exon 1 of KIAA0495. Each vertical bar represents a single CpG site. The transcription start site is indicated by a curved arrow. MSP sites and BGS region are shown. B Silencing of KIAA0495 in representative tumor cell lines correlated with its promoter methylation, while no methylation was detected in normal immortalized cell lines (in green underlined). CRC: colorectal cancer; HNC, head and neck cancer; ESCC: esophageal squamous cell carcinoma; Ca, carcinoma. M, methylated; U, unmethylated. C Endogenous expression of SP0495 in representative tumor cell lines by Western blot, with GAPDH as a loading control. Representative data are shown. Sample names in green underlined are normal tissues and normal immortalized cell lines. D Part of the KIAA0495 promoter CGI (-228 to +260) with 50 CpG sites was analyzed by BGS. Each row represents a single allele of the promoter analyzed, and one circle indicates one CpG site. Filled circle, methylated; Open circle, unmethylated. MSP primers (m1/m2) were indicated. DKO, DNMT1 and DNMT3B double knock-out HCT116 cell line. NPx, nasopharynx. E Pharmacological and genetic demethylation to restore KIAA0495 expression in silenced cell lines of various types was examined by RT-PCR and Western blot. Representative data are shown. A + T, combined treatment using Aza and TSA. 1KO, DNMT1 knock-out, 3BKO, DNMT3B knock-out, DKO, DNMT1 and DNMT3B double knock-out. F KIAA0495 promoter is frequently methylated in primary tumor samples. Colorectal cancer (CRC) tissues (T) and paired surgical marginal non-tumor tissues (N), gastric cancer tissues, nasopharyngeal carcinoma (NPC) and normal nasopharynx (NPx) tissues are examined by MSP. Representative data are shown. Ca, carcinoma. M, methylated; U, unmethylated. G KIAA0495 promoter methylation in representative tumor/normal pairs of CRC and ESCC tissues by BGS. Filled circle, methylated; Open circle, unmethylated. H Real-time PCR and Western blot show the downregulation of KIAA0495 in multiple primary tumors compared to matched normal tissues. Gene expression was normalized to internal control GAPDH. Data are presented as mean ± SEM of three independent experiments via Student’s t test. *, p < 0.05; **, p < 0.01. I Association of KIAA0495 promoter methylation at specific CpG sites with patient overall survival is analyzed in rectum, esophageal, and breast metastatic tumor samples from TCGA datasets by Kaplan Meier analysis. The methylation patient groups are dichotomized by higher (β > cut-off) and lower (β < cut-off), according to a best cut-off point in MethSurv. Ca, cancer.
We next examined the methylation status of KIAA0495 by methylation-specific PCR (MSP). We detected methylated promoters in cell lines with decreased or silenced KIAA0495 expression, and no methylation was found in normal or immortalized cell lines (Fig. 3B, Table 1). KIAA0495 methylation was less frequently observed in lung, liver, renal, ovarian, prostate and brain cancer cells, consistent with its expression status in these tumors (Suppl. Fig. S4B). We validated the MSP results in several cell lines using high-resolution bisulfite genomic sequencing (BGS) analysis of 50 individual CpG sites within the KIAA0495 CGI. BGS results were consistent with those of MSP, in which densely methylated CpG sites were only detected in methylated cell lines, but not in unmethylated normal cell line NP69 (Fig. 3D, Suppl. Fig. S5A). Thus, the KIAA0495 promoter methylation status is well correlated with its expression levels in normal and tumor cell lines.
We further investigated whether promoter methylation directly contributes to KIAA0495 silencing, using DNMT1 and DNMT3B double knock-out HCT116 cells (HCT116/DKO). Compared to wild-type HCT116 cells with completely silenced KIAA0495, expression of KIAA0495 was significantly reactivated in HCT116/DKO cells, along with concomitant full demethylation of the promoter (Fig. 3E). After demethylation treatment with DNA methyltransferase (DNMT) inhibitor Aza, alone or together with histone deacetylase inhibitor Trichostatin A, KIAA0495 expression was dramatically restored in several silenced tumor cell lines, along with the significant increase in unmethylated promoter alleles (Fig. 3E). Further high-resolution BGS analysis confirmed the demethylation of the KIAA0495 promoter after pharmacologic or genetic demethylation (Fig. 3D, Suppl. Fig. S5A). These results indicate that KIAA0495 is regulated by promoter CpG methylation in normal and tumor cells, by DNMT1 and DNMT3B together, like other bona fide TSGs which we and others previously characterized [29, 30].
To assess whether genetic alteration, such as mutation, also inactivates SP0495 in tumors, we sequenced all KIAA0495-ORF2 coding exons in a panel of tumor cell lines but found no mutation in any of the 19 cell lines examined (Table S5). These results suggest that genetic point mutation of SP0495 is likely very rare and that epigenetic alteration is the predominant mechanism of its disruption in tumors.
KIAA0495 promoter methylation is common in multiple primary tumors
To evaluate whether KIAA0495 promoter methylation in tumors is of clinical significance for developing as a biomarker for cancer detection and prognosis prediction, we examined KIAA0495 methylation in a series of primary tumors. KIAA0495 methylation was frequently detected in primary tumors of colorectal (14/23, 61%), gastric (15/51, 30%) and nasopharyngeal (28/48, 58%) cancers, but less in esophageal (7/46, 15%) and breast (3/40, 7.5%) cancers (Fig. 3F, Suppl. Fig. S5B). In contrast, no normal nasopharyngeal (0/6), esophageal (0/7), and gastric (0/4) tissues showed KIAA0495 methylation (Fig. 3F, Table 1). Among adjacent normal tissues, no esophageal normal tissue (0/46) and only two normal colorectal tissues (2/12, 17%) showed methylation. BGS analysis confirmed the dense methylation in tumors, but not in paired normal tissues (Fig. 3G and Supp. Fig. S5C). These results demonstrate that KIAA0495 methylation is a frequent and tumor-specific event in multiple solid tumors. In agreement, qRT-PCR results also showed that KIAA0495 expression was reduced dramatically in tumor tissues compared with adjacent normal tissues, and further confirmed by Western blot (Fig. 3H). Furthermore, we detected KIAA0495 methylation in 7/16 (44%) nose swab samples from NPC patients (Suppl. Fig. S5D), suggesting that it could be developed as an epigenetic biomarker for non-invasive cancer diagnosis.
Clinically, through analyzing TCGA cancer dataset, higher KIAA0495 promoter methylation level is significantly associated with poor outcomes of patients with rectum, esophageal, and breast cancers (Fig. 3I). These findings demonstrate the translational value of tumor-specific KIAA0495 promoter methylation as a cancer biomarker, and also the important role of KIAA0495 inactivation during tumor pathogenesis.
Small protein SP0495 functions as a tumor suppressor
Frequent KIAA0495 methylation in multiple carcinomas implies tumor-suppressive functions of its encoded small protein SP0495. To test this, we firstly examined its growth inhibitory effect on tumor cells by colony formation assays. Compared to controls, significant reduction of colony numbers and sizes were observed in cells stably expressing SP0495, in both monolayer and soft-agar culture colony formation assays (Fig. 4A–D, Supp. Fig. S6A), indicating that SP0495 inhibits the anchorage-dependent and -independent growth of tumor cells.
Monolayer-culture (A) and soft-agar (B) colony formation assays of tumor cells. Representative data are shown with three independent experiments. C Stable expression of SP0495 in tumor cells detected by Western blot. D Quantitative analysis of colony numbers in A and B, shown as ±SEM of three independent experiments using Student’s t test. *, p < 0.05. E Representative results of TUNEL assay of tumor cells. Red fluorescent signals indicate TUNEL+ (apoptotic) cells. Representative images are shown. Scale bar, 40 µm. F Flow cytometry analysis of apoptosis by Annexin V-FITC/PI staining of HCT116 and KYSE150 cells. Both early and late apoptotic cells (Annexin V-positive) were counted. Mean ± SD of three independent experiments is shown. **, p < 0.01, ***, p < 0.001. G Increased cleaved PARP in SP0495-expressing HCT116 and KYSE150 cells by Western blot. H Cell cycle analysis using PI staining by flow cytometry. Representative cell cycle histograms of HCT116 and KYSE150 cells with SP0495 expression show significant increase in G1/S populations as well as significant decrease in S phase population, compared to controls. Bar diagram compares variations in cell distribution percentage in each phase of cell cycle of HCT116 and KYSE150 cells. Statistical analysis is performed by one-way ANOVA. Data represented as mean ± SD of three independent experiments in triplicates. *, p < 0.05; **, p < 0.01. I Expression of cell cycle-related proteins in SP0495-expressing HCT116 and KYSE150 cells. Western blot analysis of phosphor-Rb/Rb, CDK4/6, phosphor-p53/p53, and p21. J Time-dependent changes in Cyclin B1 protein level following nocodazole treatment. HCT116 and KYSE150 cells with or without SP0495 expression were treated with nocodazole (1 µM) for the length of time (hours) as indicated. Western blots were performed using indicated antibodies and GAPDH as a loading control. Summary of percentage of cells in each phase of the cell cycle with nocodazole treatment (1 µM, 24 h). Significant differences are indicated as followings: *, p < 0.05; ** p < 0.01 and *** p < 0.001. Bars are mean ± SEM from three independent experiments after Student’s t test analysis. K The activity of senescence-associated β-galactosidase in SP0495-transfected NIH3T3 cells. Representative images of NIH3T3 monolayer cultures stained for SA-β-gal activity are shown. Red arrows indicate positive staining. The percentage of SA-β-gal-positive (blue) cells is calculated (with counting >50 cells per condition) and presented as mean ± SEM of three independent experiments via Student’s t-test. Scale bar 10 µm. ***, p < 0.001. L SP0495 suppresses colorectal tumor formation in vivo. HCT116 cells were transduced with CMV-Firefly luciferase lentivirus encoding SP0495 or empty vector, then injected subcutaneously into BALB/c nude mice. Tumor size and tumor weight were monitored. Tumor fluorescence intensity was quantified. **, p < 0.01. p values determined by unbiased Student’s t test. Representative animals are shown.
We further explored the underlying mechanisms of tumor suppression mediated by SP0495. We found that ectopic SP0495 expression induced apoptosis of HCT116 and KYSE150 tumor cells, as demonstrated by TUNEL assay (Fig. 4E). Additionally, flow cytometry analysis by Annexin V assay showed that the proportion of apoptotic cells was significantly higher in SP0495-expressing HCT116 and KYSE150 cells than in control cells (Fig. 4F). Consistently, 4,6-diamidino-2-phenylindole (DAPI) staining revealed chromatin condensation and nuclear rupture in SP0495-expressing tumor cells—a hallmark of apoptotic cell death (Suppl. Fig. S6B). We also observed increased expression of the apoptotic indicator - cleaved PARP, in SP0495-expressing cells as compared to vector controls (Fig. 4G). We then determined whether SP0495 has an impact on cell cycle progression. We found that the tumor cell cycle pattern was altered by SP0495, with significant increase of cell proportion in G1/S phases and decrease of cell population in S phase (Fig. 4H). We further examined the effect of SP0495 on cell cycle-related protein expression, and found that G1-S associated proteins (phosphorylated Rb, CDK4, and CDK6) were downregulated after SP0495 expression, while the expression levels of p53 and p21 proteins as well as phosphorylated p53 (at Ser15) were increased after SP0495 expression (Fig. 4I). To further explore how SP0495 restoration modifies cell cycle, we treated cells with nocodazole to capture cells in G2/M. Western blot analysis of cyclin B1, a key cell cycle regulator of G2/M transition, showed significant increase of protein expression in control cells after nocodazole treatment, but with only minor changes in SP0495-expressing cells (Fig. 4J). Consistently, we observed that control cells after nocodazole treatment mainly accumulated in G2/M, while SP0495 overexpressing cells were still delayed mainly in G1 phase (Fig. 4J). These data indicated the arrest of G1/S phase progress by SP0495 in tumor cells.
Moreover, as p53 and p21 are both linked to cell senescence, we also detected the impact of SP0495 expression on cell senescence. We detected induction of cell senescence in SP0495-expressing immortalized normal cells by staining for senescence-associated β-galactosidase (SA-β-gal). Elevated β-galactosidase staining was observed in SP0495-expressing cells (Fig. 4K), indicating that cell senescence induction is another growth-suppressive mechanism mediated by SP0495. These findings suggested that SP0495 could function as a tumor suppressor through inducing apoptosis, G1/S cell cycle arrest and cell senescence in tumor cells.
An nude mice animal model was used to investigate whether SP0495 could suppress tumor formation in vivo. HCT116 and MB231 tumor cells with stably expressed SP0495 or control vector were injected into nude mice, with tumor formation efficiency monitored across different time points. SP0495 overexpression significantly decreased tumor growth and average tumor weight of HCT116 and MB231 xenografts in nude mice (Fig. 4L, Suppl. Fig. S6C). Compared with the control group, the fluorescence intensity in SP0495-expressing HCT116 cell group decreased significantly (Fig. 4L), together with slower tumor growth and reduced average tumor weight (Fig. 4L, Suppl. Fig. S6C. We further tested the tumor-suppressive functions of SP0495 under siRNA knock-down. We found that increased tumor cell growth through knockdown of SP0495 by siRNAs in BT549 and T47D cells, accompanied by reduced proportion of G0/G1 cells and elevated proportion of S and G2/M cells, suggesting that SP0495 knockdown could revert the proliferation suppression and G1/S arrest induced by SP0495 (Fig. 5A, B). Thus, SP0495 indeed functions as a tumor suppressor in tumorigenesis.
A Monolayer colony formation assay of BT549 and T47D cells treated with two different KIAA0495 siRNAs. Each bar represents mean ± SEM for three independent experiments via One-Way ANOVA with post hoc analysis, with colonies (>50 cells) in empty vector-transfected cells set as 100. B Flow cytometry was performed for cell cycle analysis using PI staining. Representative cell cycle histograms of BT549 cells with SP0495 knockdown by siRNAs are shown. Bar diagram compares variations in cell distribution percentages in each phase of cell cycle of BT549 cells. Statistical analysis is performed by One-Way ANOVA with post hoc analysis. Data represented as mean ± SD of three independent experiments in triplicates. C Effects of SP0495 on six key signaling pathways are assessed by luciferase reporter assays. Results are expressed as fold reduction of activity and shown as mean ± SEM of three independent experiments performed in triplicate analyzed by Student’s t test. *, p < 0.05; **, p < 0.01. D Changes of key molecules in oncogenic signaling pathways (AKT/mTOR, Wnt-β-catenin, NF-kB signaling) shown by Western blot in D SP0495-expressing HCT116 and KYSE150 cells and E BT549 cells with KIAA0495 knockdown by siRNAs. F Real-time PCR results demonstrate that SP0495 represses multiple AKT/mTOR upstream activators (PDK1, ID1, and EEF1A2) and a panel of downstream target genes for AKT/mTOR, Wnt/β-catenin and NF-κB signaling (c-Myc, CCND1, MMP7, MITF, and TWIST1). Data are presented as mean ± SEM of three independent experiments via Student’s t test. *, p < 0.05; **, p < 0.01. G Immunoblot analysis of c-Myc, CCND1, and MMP7 expression levels, as repressed by SP0495 in HCT116 and KYSE150 cells. Representative data are shown. H Analysis of colorectal adenocarcinoma from TCGA database indicates that the mRNA expression status (Z-score) of downstream targets in AKT/mTOR, Wnt/β-catenin and NF-kB signaling pathways analyzed using cBioPortal. Right bottom: expression profiles of indicated genes in Wnt/β-catenin pathway in 32 normal colon tissues and 32 colorectal adenomas are extracted from Oncomine.
SP0495 suppresses AKT and other oncogenic signaling pathways
As SP0495 is a novel tumor suppressor located in the cytoplasm, we thus hypothesize that it might regulate cell signaling to exert its tumor suppression. We utilized several luciferase reporters of critical signaling pathways related to tumorigenesis, including p53-binding sites (bs) (p53), p21 promoter (p21), STATs-bs/GRR5 (STATs), NF-κB-bs (NF-κB), AP1-bs (JNK), SRE (Ras/ERK), and TOPFlash (Wnt) pathways. It was found that the activities of p53 signaling reporters were significantly upregulated, but NF-kB and Wnt signaling reporters were significantly repressed by SP0495, in both HCT116 and KYSE150 cells (Fig. 5C), in addition to the repression of JNK and STAT signaling reporters in KYSE150 only. Moreover, SP0495 suppressed TOPFlash reporter activity in a dose-dependent manner, while with no inhibitory effect on control FOPFlash reporter which harbors mutant TCF/LEF binding sites (Suppl. Fig. S7A).
Furthermore, we examined the regulation of SP0495 on oncogenic signaling pathway regulators. We observed that phosphor-AKT at Ser473 (active form), phosphor-mTOR (Ser2448), phosphor-GSK3β at Ser9 (inactive form), and active β-catenin were downregulated by SP0495 expression (Fig. 5D). We also found that the active forms of NF-κB signaling molecules, including p-IKKα/β, p-IκBα, and p-p65, were suppressed in SP0495-expressing cells (Fig. 5D). Knockdown of SP0495 in turn increased the activities of these signaling pathways (Fig. 5E). Real-time PCR results showed that the expression of AKT-mTOR and NF-κB singling activators, PDK1, ID1, and EEF1A2, was downregulated by SP0495. Meanwhile, multiple downstream effectors of AKT/mTOR, Wnt, and NF-κB signaling, such as c-Myc, CCND1, MMP7, MITF, and TWIST1, were suppressed in SP0495-expressing cells compared to controls, although the mRNA level of β-catenin (CTNNB1) remained unchanged (Fig. 5F). We also confirmed the downregulation of Wnt signaling downstream effectors (c-Myc, CCND1, MMP7) at the protein level by SP0495 expression (Fig. 5G).
We then analyzed the relationship between co-expression of KIAA0495 RNA and signaling molecules in CRC using TCGA colorectal adenocarcinoma database. We found that KIAA0495 overexpression is associated with reduced expression of AKT/mTOR (EIF4EBP1 and RPS6KB1), NF-κB (BCL2, BCL2L1, BIRC5/Survivin, and SQSTM1/p62) and Wnt (ID1, CCND1, MYC and MMP7) signaling molecules (Fig. 5H), with ID1 expression mutual exclusive with KIAA0495 expression (p = 0.020). Co-expression analysis of Oncomine database showed that KIAA0495 was highly expressed in normal colon tissues, but with much lower expression in colorectal adenomas, while levels of Wnt signaling molecules negatively correlate with KIAA0495 levels across these samples (Fig. 5H).These results suggest that KIAA0495 (SP0495) contributes to the suppression of oncogenic signaling axis during tumor formation and progression.
SP0495 induces autophagy in carcinoma cells
To further investigate the molecular mechanisms underlying SP0495 tumor suppression, we analyzed changes in gene expression profile mediated by SP0495 in tumor cells and immortalized normal cells through RNA-sequencing and microarray expression analysis. GO enrichment analysis showed that regulation of apoptosis and multiple oncogenic signaling pathways were the mainly enriched biological processes in HCT116 and KYSE150 cells (Fig. 6A). Microarray expression analysis showed that the enriched signaling pathways affected by doxycycline (DOX)-induced SP0495 expression in 293 cells included focal adhesion, senescence, and autophagy (Suppl. Fig. S7B, C). Moreover, GSEA showed that SP0495 significantly induced apoptotic signaling response to ER stress and positively regulated the signal transduction by p53 mediators (Fig. 6B). These results are consistent with the suppression of autophagy-related oncogenic signaling (AKT/mTOR, NF-κB, and Wnt) and activation of p53 signaling by SP0495 which we observed above in tumor cells.
A GO enrichment analysis of differentially expressed genes identified through comparisons of SP0495-expressing tumor cells vs control cells. Q value < = 0.05 is regarded as significant enrichment. B Gene set enrichment analysis (GESA) revealed significantly enriched pathways in SP0495-expressing group vs control group. NES, normalized enrichment score. C Electron microscopy images of autolysosome formation in SP0495-expressing HCT116 cells. Data are representative images by transmission electron microscopy (TEM) from three independent assays. Autophagosomes with double-membrane (yellow arrows) and autolysosomes with cargo in different stages of digestion (red arrows) are indicated. Scale bar, 5 µm. D Representative images detecting autophagy flux regulated by SP0495 in carcinoma cells infected with mRFP-GFP-LC3 adenovirus. Quantification of mRFP-GFP-LC3 shows that SP0495 significantly increases autophagy flux in HCT116 and KYSE150 cells. ~50 cells are randomly selected for each statistical analysis. Data are presented as mean ± SEM of three independent experiments via Student’s t-test. **, p < 0.01; ***, p < 0.001. Scale bar, 20 µm. Original magnification, 400×. E, F Western blot analysis of the expression of apoptosis and autophagy proteins in HCT116 and KYSE150 cells with SP0495 expression or empty vector, with or without BECN1 siRNA treatment. Results are representative of three independent experiments. G SP0495 facilitates cell response to autophagy by Western blot analysis of the expression of autophagy regulator proteins. HCT116 and KYSE150 cells were exposed to Torin1 (250 nM, 500 nM) for 4 h. Results are representative of three independent experiments. H SP0495-expressing HCT116 and KYSE150 cells are treated with 10 μM BafA1 for 24 h, with LC3A/B and p62 levels measured by Western blot. Representative data are shown. I BafA1 suppresses colony formation in SP0495-expressing HCT116 and KYSE150 cells. Representative images are shown. Quantitative analyses of colony numbers are shown as values of mean ± SEM of three independent experiments using Student’s t test, with colonies (>50 cells) of vector cells (DMSO treatment) set as 100. *, p < 0.05; **, p < 0.01; ***, p < 0.001.
We thus further examined the effects of SP0495 on autophagy in tumor cells. Transmission electron microscopy showed increase in the formation of autophagic vesicles in SP0495-expressing HCT116 cells (Fig. 6C). Further autophagic flux using an mRFP-GFP-LC3 reporter construct showed that more ectopically expressed mRFP-GFP-LC3 was detected as red and yellow speckles in SP0495-expressing tumor cells than control cells, indicating that SP0495 induces accumulation of autophagosomes in tumor cells (Fig. 6D).
As SQSTM1/p62 and BECN1 levels are critical indicators of autophagic flux, we further detected autophagy regulators to confirm the regulation of SP0495 on autophagy. It was found that SP0495 upregulated the levels of cleaved-PARP, BECN1 and ATG5, while downregulated the levels of BCL2 and SQSTM1/p62, thus mediating autophagosome form LC3-II conversion in tumor cells (Fig. 6E). To demonstrate the role of BECN1 upregulation in SP0495-mediated autophagy, knockdown of BECN1 using siRNA was used in SP0495-expressing HCT116 and KYSE150 cells. Results showed that cleaved-PARP and L3CII protein levels were decreased after BECN1 knockdown but p62 levels were increased, suggesting that BECN1 upregulation by SP0495 is responsible for its tumor-suppressive functions (Fig. 6F). We next examined the effects of SP0495 on Torin1-induced autophagy and found that SP0495 upregulated LC3II protein levels after Torin1 treatment, suggesting that SP0495-expressing tumor cells are more susceptible to cell death stimuli (Fig. 6G). Moreover, Bafilomycin A1 (BafA1), a lysosomal inhibitor, abrogated the autophagy induction by SP0495 in tumor cells (Fig. 6H). We further investigated whether the blockage of autophagic response in tumor cells preceded cell growth inhibition by SP0495. We found that inhibition of autophagic response with the lysosomal inhibitor BafA1 significantly enhanced SP0495-mediated inhibition of cell clonogenic ability, in both HCT116 and KYSE150 cells (Fig. 6I). These data suggest that SP0495 induces autophagic flux through regulating autophagy proteins in tumor cells.
SP0495 promotes autophagy through enhancing BECN1 stability
Autophagy is mainly regulated by BECN1 and SQSTM1/p62, with p62 as an autophagic degradation marker, we thus detected effects of SP0495 on BECN1 and p62 expression levels. Results showed that SP0495 had no significant effect on the mRNA expression levels of BECN1 and p62 (Fig. 7A), however endogenous SP0495 expression level is correlated with upregulated BECN1 level in a panel of cell lines (Fig. 7B), suggesting that SP0495 might be involved in regulating BECN1 expression at the protein level.
A Expression levels of BECN1 and p62 at the mRNA level in SP0495-expressing tumor cells. Bars show relative fold changes by quantitative real-time PCR (qRT-PCR) analysis. All data were normalized with GAPDH expression and given as relative to control. p values determined by Student unpaired t test. n.s. no significant change. B Western blot examined the expression of BECN1 and p62 at the protein level in a panel of tumor cell lines with or without endogenous SP0495 expression. C Cycloheximide (CHX)-chase assay for the half-life of BECN1 and p62 in KYSE150 cells. Left panel, KYSE150 cells with SP0495 or vector expression are treated with CHX (20 μg/mL) for the indicated time points, and Western blotting with indicated antibodies. Right panel, the levels of remaining BECN1 and p62 at different time points are normalized to GAPDH from three independent experiments. D–G SP0495 decreases BECN1 ubiquitination and increases the ubiquitination of p62. T-REx-293 with inducible SP0495 expression and SP0495-stable expressing KYSE150 cells were transfected with or without His-Ub plasmid and/or treatment of 10 μM MG132 for 6 h. Cell lysates were immunoprecipitated with anti-BECN1 (D, E) or anti-p62 (F, G) antibody, the precipitated materials are subjected to immunoblot for endogenous Ub (D, F) or exogenous His-Ub (E, G) to detect the ubiquitination levels of BECN1 and p62. Results are representative of three independent experiments. H BECN1 and p62 ubiquitination are performed on Ni-NTA pull-down ubiquitination analysis. HEK293T and KYSE150 cells were transfected with indicated plasmids and then treated with MG132 (10 μM) for 6 h. BECN1 and p62 ubiquitination were monitored by immunoblots performed on Ni-NTA purified proteins. Results are representative of three independent experiments.
As autophagy is a cellular degradation process through stabilizing BECN1 or degrading p62 via protein modifications such as ubiquitination, we next detected the half-life of endogenous BECN1 and p62 affected by SP0495 in tumor cells. CHX assay showed that SP0495 extended the half-life of BECN1 over 8 h, but shortened the half-life of p62 from 6 h to ~2 h, suggesting that SP0495 modulates the stability of both BECN1 and p62 proteins (Fig. 7C).
We next evaluated whether modulation of protein ubiquitination by SP0495 led to the regulation of BECN1 and p62 protein stabilities. To assess the effects of SP0495 on endogenous or exogenous ubiquitin chain of BECN1 and p62, we performed coimmunoprecipitation (co-IP) assays in SP0495-inducible expressed 293 cells and stably-expressed KYSE150 cells. We found that SP0495 decreased endogenous or exogenous ubiquitin linked with BECN1, in the presence and absence of proteasome inhibitor MG132 (Fig. 7D, E), while increased endogenous or exogenous ubiquitin linked with p62, in the presence and absence of MG132 (Fig. 7F, G). Furthermore, Ni-NTA (nickel nitrilotriacetic acid) ubiquitination assay showed decreased BECN1 ubiquitination but increased p62 ubiquitination by SP0495 in HEK293T and KYSE150 cells co-transfected with His-Ub and Flag-SP0495, after MG132 treatment (Fig. 7H). These results suggest that SP0495 regulates both BECN1 and p62 stability to induce autophagy in tumor cells.
SP0495 acts as a lipid-binding-protein to regulate AKT signaling and autophagy
To elucidate the underlying molecular mechanism of SP0495 in regulating the stabilities of BECN1 and p62 proteins, we examined the interaction of SP0495 with BECN1 or p62 by co-IP assay. However, no direct binding was observed (data not shown).
Autophagy is a membrane-driven catabolic pathway through the interaction of membrane lipids with autophagy machinery proteins. Phosphoinositides and phosphoinositide-binding proteins play essential roles in the regulation of lipid membrane trafficking/signaling, autophagy and cell signaling events, especially AKT activation and signaling [31]. Thus, we sought to investigate the possible interaction of SP0495 protein with phosphoinositides. First, we performed 3D structure model analysis of SP0495 protein by Phyre2 (Fig. 8A, left panel). The secondary structure of SP0495 was described as disordered of 53%, alpha-helix of 19%, and beta-strand of 17%. We also validated the predicted 3D structure of SP0495 by Swiss-Model (Fig. 8A, right panel). The MolProbity Score was 2.59 and Ramachandran favored 77.78% with QMEAN, Cβ, all-atom, solvation, and torsion values of −1.28, −0.83, −0.10, −0.89, and −1.01, respectively. The domain analysis by Phyre2 showed a 3D structure of lipid-binding-protein template (c1lshB) out of SP0495 (Fig. 8B). Further protein motif analysis by DisoLipPred revealed two lipid-binding motifs within the SP0495 protein (Fig. 8B). These results suggest possible binding of SP0495 with phosphoinositides for the regulation of autophagy and cell signaling.
A Structure of the SP0495 protein predicted by Phyre2 and Swiss Model. B The tertiary structure of SP0495 is predicted by Phyre2, based on the template of c1lshB as a lipid-binding protein (left panel). Lipid binding domain predicted by DisoLipPred shows two lipid-protein binding motifs within SP0495 (right panel). C Schematic diagram of SP0495 and its deletion mutants. SP0495-mut1 lacks the lipid-binding domain1 and SP0495-mut2 lacks the lipid-binding domain2. The in vitro expressed recombinant KIAA0495-ORF2/SP0495 protein and its two mutants are shown. D Membrane and cytoplasmic fractionations of HCT116 and KYSE150 cells with or without SP0495 expression were performed. Proteins in each fraction were detected using antibodies against SP0495, INSR (membrane) and GAPDH (cytoplasm) by Western blot. E Colony formation assay shows the difference in colony formation suppression abilities of SP0495 full-length and mutants. Quantitative analyses of colony numbers are shown as values of mean ± SEM of three independent experiments using One-Way ANOVA with post hoc analysis. Colonies (>50 cells) in empty vector-transfected cells set as 100. F Effects of SP0495 lipid binding domain deletions on cell apoptosis by flow cytometry analysis via Annexin V-FITC/PI staining. Both early and late apoptotic cells (Annexin V-positive) were counted. Mean ± SD of three independent experiments. Quantitative analyses of apoptotic cells using One-Way ANOVA with post hoc analysis. G Western blot analysis of oncogenic signaling and autophagy protein level changes in HCT116 and KYSE150 cells expressing SP0495 and mutants. Results are representative of three independent experiments. H Recombinant SP0495 and its mutant proteins were used to probe PIP StripsTM and PIP ArraysTM. Bound proteins were detected using anti-SP0495 antibody. Representative data are shown of three independent experiments. LPA, lysophosphatidic acid; LPC, lysophosphatidylcholine; PI, phosphatidylinositol; PE, phosphatidylethanolamine; PC, phosphatidylcholine; S1P, sphingosine-1-phosphate; PA, phosphatidic acid; PS, phosphatidylserine. I Immunostaining of KYSE150 cells expressing SP0495 and its mutants with antibody against PI(3,4,5)P3 (green). Nuclei counterstained with DAPI (blue). Original magnification, 400×. Scale bar 40 µM. J Schematic diagram illustrates the role of SP0495, as a plasma membrane phosphoinositide-binding protein, functioning as a tumor suppressor through regulating AKT phosphorylation/activation and autophagy during multiple tumor pathogenesis.
To assess that SP0495 functions as a lipid-binding-protein, we constructed two lipid-binding motif deletion mutants, SP0495-mutant 1 with lipid-binding domain 1 deleted and SP0495-mutant 2 with lipid-binding domain 2 deleted (Fig. 8C), with recombinant proteins of these SP0495 constructs successfully detected (Fig. 8C). We observed that the full-length SP0495 was mainly located on cell membrane but its mutants mainly located in the cytoplasm (Fig. 8D). Colony formation assay showed that mutant 1 lost the ability of cell growth inhibition and apoptosis induction, while the ability of cell growth inhibition and apoptosis induction of mutant 2 was only slightly decreased or not affected at all (Fig. 8E, F). We further examined the effects of SP0495 mutants on cell signaling and autophagy marker expression. Results showed that SP0495-mutant 1 was seriously impaired in the ability to activate p53 signaling, suppress AKT/mTOR signaling and Rb phosphorylation, and autophagic signaling induction (Fig. 8G). These results suggested that the lipid-binding domain 1 is a major and critical functional domain for SP0495-mediated tumor-suppressive functions.
We further investigated the possible interaction of phosphoinositide with SP0495. Lipid overlay experiments using PIP strips were performed using recombinant human SP0495 protein and its two mutants (Fig. 8H). We found that human SP0495 protein mainly interacted with phosphatidic acid (PA), PtdIns(3)P, PtdIns(5)P, and PtdIns(3, 5)P2, with a preference for PA, weakly with PtdIns(3,4,5)P3, PtdIns(4, 5)P2, and PtdIns(4)P (Fig. 8H), while mutant1 primarily interacted with PI(3, 4, 5)P3 and PI (4, 5)P2, but mutant 2 remained similar binding feature as the wild-type SP0495 protein. Protein-lipid binding assay using PIP arrays also revealed that SP0495 strongly interacted with PI(3)P and PI(3,5)P2, while mutant1 displayed almost no PI3P interaction (Fig. 8H). We further performed immunofluorescence staining to examine the expression levels of PI(3,4,5)P3 in tumor cells expressing SP0495 and its mutants. Results showed that anti-PI(3,4,5)P3 staining was diminished or greatly reduced in tumor cells expressing SP0495 or mutant 2, while expression of mutant 1 did not affect PI(3,4,5)P3 staining much (Fig. 8I). These results suggest that SP0495, indeed as a lipid-binding protein, regulates autophagy and cell signaling through interacting with phosphoinositides in cells.
Discussion
1p36.3 is an important TSG locus implicated in the early events of tumorigenesis in multiple cancers and is thus believed to harbor critical TSGs [1, 2]. Although several TSGs residing in this locus, including TP73 [4], CHD5 [5], PRDM16 [6], and AJAP1 [7], have already been validated, more TSG candidates are likely still waiting to be characterized as 1p36.3 is a gene-rich region. In this report, through integrative epigenome study, we identified a novel 1p36.3 gene, KIAA0495, frequently methylated and silenced in broad cancers in a tumor-specific manner, indicating its tumorigenesis-associated functions. We further present direct evidence that although KIAA0495 was previously claimed to be a lncRNA, it actually encodes a small protein SP0495, which induces tumor cell apoptosis, cell cycle arrest, senescence, autophagy, and inhibits tumor cell growth in vivo. SP0495 represses oncogenic AKT/mTOR, NF-κB, and Wnt/β-catenin signaling. As a lipid-binding protein, SP0495 regulates autophagy through disrupting autophagic/proteasomal degradation of p62 and BECN1. We also found that although KIAA0495 transcript/protein is broadly expressed in multiple normal tissues, it is silenced in multiple tumors by promoter CpG methylation, and its downregulation/methylation is associated with high-grade stage and poor survival of multiple cancer patients. Thus, our results validate that KIAA0495/SP0495 is a bona fide TSG/tumor suppressor being frequently inactivated by epigenetic mechanisms in multiple cancers.
Genome-wide association studies have pinpointed 1p36 as a susceptibility locus for multiple cancers [32,33,34] and even certain developmental disorders such as azoospermia [35]. Other 1p36.3 TSGs have previously been reported to contribute to multiple tumorigeneses. For example, TP73 is mapped to a minimal region of 1p36.3 commonly deleted in neuroblastomas, and functions as a p53-like TSG, inducing cell cycle arrest and apoptosis. Epigenetic silencing of TP73 leads to cell cycle deregulation in hematological and oligodendroglial tumors [4, 36]. Through mouse chromosome engineering, Bagchi et al. identified a critical 1p36.31 tumor suppressor—the chromodomain helicase DNA binding domain 5 (CHD5), which controls cell proliferation, apoptosis, and senescence via p14ARF/p53 pathway [5], and is also epigenetically silenced in multiple cancers [37,38,39,40]. Thus, novel 1p36 TSGs may be important for cancers and other diseases associated with genetic loss of this small genome fragment or epigenetic inactivation of its encoded genes.
A long 1p36.3 transcript KIAA0495 was first submitted to the NCBI database by Kazusa DNA Research Institute in 1997 [41]. In the current NCBI database, this gene is names as TP73-AS1, as this gene is in the immediate vicinity of TP73 and the KIAA0495 transcript is partially complementary to the TP73 transcript. However, the complement is only at a small region (~218 bp) of the C terminal of the TP73 gene, thus the previously named term TP73-AS1 is actually somewhat misleading (Suppl Fig. S2). A previous report indicated that full-length KIAA0495 cDNA (~6.4 kb) may not produce protein by in vitro translation analysis [42]. However, the authors did observe a protein product (~24 kDa) when expressing the ORF2 of KIAA0495 (606 nucleotides, 201 aa residues) using in-house made anti-PDAM/KIAA0495 antibody (a peptide TTSDLSAREDATPSC against 66-79 amino acid residues) [42], which is consistent with our findings regarding the coding potential of KIAA0495-ORF2 (SP0495).
Recently with the advancement of biological technology, it has been recognized that some non-canonical ORFs derived from lncRNAs or UTRs do possess peptide/small protein-coding properties [12, 13, 43,44,45]. Some of these peptide/small proteins have been validated as functional oncogenes or TSGs in tumorigenesis [18, 19, 45]. Here, we demonstrate that the KIAA0495-ORF2 codes a small protein SP0495, through in vitro translation and further endogenous protein detection in cell lines and normal tissues, by Western blot and immunostaining using an antibody targeting the SP0495 protein. KIAA0495/SP0495 is readily expressed in multiple normal tissues, although with various expression levels. However, it is frequently downregulated, but rarely mutated, in multiple tumors including esophageal, colorectal, gastric, and breast cancers, and correlated with poor survival of multiple cancer patients, indicating its important roles in cancer pathogenesis.
As SP0495 protein is a previously uncharacterized protein, we investigated its biological functions and underlying mechanisms in-depth. SP0495 contains a transmembrane and signal peptide domain, and is located in the cytoplasm and partly co-localized with ER and Golgi. SP0495 exerts tumor-suppressive functions through inhibiting proliferation, inducing apoptosis and G1/S cell cycle arrest. Moreover, SP0495 suppresses tumor growth in vivo, thus indeed as a bona fide tumor suppressor. Mechanistically, SP0495 negatively regulates AKT/mTOR, NF-κB, and Wnt/β-catenin signaling cascades, through repressing AKT/mTOR and NF-kB upstream effectors including PDK1/PDPK1 [46], ID1 [47], EEF1A2 [48], and downstream targets, although PI3K remains unaffected. Genome-wide dataset analysis revealed that KIAA0495 is unregulated ~1.75 fold in tumor cells which are sensitive to PI3K/AKT inhibitor (GDC-0941), compared to resistant cells [49], suggesting that KIAA0495 inactivation may confer tumor cells resistance to such targeted therapy. Furthermore, SP0495-regulated genes by RNA-seq and microarray analysis are mainly enriched in signaling pathways of apoptotic regulation and p53 signaling, which is consistent with deregulation of AKT/mTOR, Wnt/β-catenin and NF-κB signaling cascades by SP0495.
We further found that SP0495 expression enhances apoptosis, cell cycle G1/S arrest, cell senescence and autophagy, by inducing elevated protein levels of p53, phosphorylated p53 at Ser15 and p21, which confirmed our RNA-seq and microarray data. In cancers, genetic/epigenetic aberrations of autophagy regulators highlight the importance of autophagy dysregulation in cancer pathogenesis and even drug resistance. Multiple tumor suppressors have been identified to induce autophagy, including BECN1, p53, PTEN, DAPK1, and LKB1/STK11 [50]. Autophagy regulators p62 and BECN1 are inversely correlated and play crucial roles in autophagy regulation in cells.
Our study demonstrates that SP0495 as a functional tumor suppressor induces autophagy in tumor cells. SP0495 promotes BECN1 accumulation but inhibits p62 accumulation, through interfering with their ubiquitination-related degradation, then further promoting autophagy. We further found that knockdown of BECN1 in SP0495-expressing tumor cells impaired its induced apoptosis and autophagy, suggesting a key role of BECN1 in SP0495-mediated tumor suppression. However, unlike other regulators, SP0495 does not bind p62 and BECN1 directly. We speculate that SP0495 may regulate autophagy as a transmembrane signal peptide, since autophagy is a membrane-driven process with lipids playing a central role in its regulation. Phosphoinositides and phosphoinositide-binding proteins play essential roles in the regulation of lipid membrane trafficking/signaling, autophagy and cell signaling events, especially AKT activation and signaling [31]. AKT needs to bind to PI(3,4,5)P3 on the plasma membrane inner leaflet via its PH domain, then undergoes conformational changes, phosphorylation/activation and further downstream signaling cascade.
However, how lipids interact with autophagy machinery regulators by SP0495 still remains unclear. Our structure analysis shows that SP0495 has similarities with lipid-binding proteins and contains two lipid-binding domains, indicating that SP0495 possibly regulates autophagy and AKT signaling as a lipid-binding protein. Phosphoinositides (PtdIns; phosphorylated derivatives of PI), consisting of ~1% of phospholipids, play key roles in lipid signaling and membrane trafficking pathways including autophagy. Emerging evidence demonstrates the important role of PI(5)P in positively regulating autophagy, through association with autophagy effectors that bind PI(3)P [51]. PI(5)P and PI(3,5)P2 are mainly localized at lysosome and autophagosome within the intracellular membrane system [52, 53]. Moreover, phosphatidic acid (PA) acts as a positive regulator of autophagy via inhibiting mTORC1 [54]. Our data show that SP0495 interacts with distinct types of lipids by protein-lipid overlay assays, including PA, PI(3)P, PI(5)P, and PI(3,5)P2, which facilitates the biogenesis and maturation of autophagosomes. Moreover, SP0495 predominantly binds PI(3)P and PI(3,5)P2, supporting our hypothesis that SP0495 regulates autophagy and AKT signaling through binding phosphoinositides. Future studies like the liposome flotation assay might verify these hypothesized protein-phosphoinositide interactions and identify the functional domain or residues of SP0495 involved in the interaction.
We also assessed the regulatory mechanisms of KIAA0495 downregulation in cancers. Promoter CpG methylation mediates KIAA0495 silencing/downregulation in multiple tumors, but rarely in normal tissues and immortalized normal cells. Moreover, no mutations were detected in any of the tumor cell lines examined, suggesting a predominant role for epigenetic inactivation of KIAA0495 in multiple cancers. We also found that KIAA0495 promoter methylation is significantly associated with poor survival of rectum, esophageal, and breast cancers, thus could be an attractive epigenetic biomarker for tumor diagnosis in future.
In conclusion, our integrative epigenomic analysis elucidates a new molecular link of a novel 1p36 tumor suppressor to multiple tumorigeneses. The small protein SP0495, encoded by the pre-claimed lncRNA KIAA0495 (TP73-AS1), functions as a bona fide tumor suppressor for multiple cancers through suppressing oncogenic signaling and regulating cell cycle, apoptosis, senescence and autophagy (Fig. 8J). The discovered tumor-specific methylation of KIAA0495 promoter could serve as a potential tumor biomarker in future.
Methods
CpG methylomes
CpG methylome analysis of cell lines was performed by methylated DNA immunoprecipitation (MeDIP) coupled with promoter microarray hybridization (MeDIP-chip) [55]. Briefly, genomic DNA of CRC cell lines (HCT116-DKO, HCT116), gastric cell line (SNU719), NPC cell line (C666-1), breast cancer cell lines (MB231, MCF7), and immortalized mammary epithelial cell line (HMEpC) was immunoprecipitated using a monoclonal antibody against 5-methylcytidine (33D3, Diagenode, Seraing, Belgium), purified, labeled and hybridized to NimbleGen™ HG18 Meth (385K CGI plus) promoter arrays (Array Star, Inc., MD). Array data analysis of methylome data was performed using SignalMap by NimbleGen Systems, Inc. as previously described [55].
In vitro translation of KIAA0495-ORF2 protein
In vitro protein expression was performed using the Human In Vitro Protein Expression Kit for DNA Templates (Thermo Fisher Scientific, Rockford, IL) according to the manufacturer’s instructions. Protein products were analyzed in SDS-PAGE gel and immunoblot with antibodies against FLAG tag (F3165, Sigma) or KIAA0495-ORF2/SP0495 (TA503634, Origene).
In vivo Xenograft models
Female BALB/c nude mice aged 4 weeks were used for tumor implantation experiments. HCT116 cells with luciferase-tag (2–5 × 106 cells in PBS) were injected subcutaneously into the flanks of nude mice with randomization (n = 6). No blinding to the group allocation during the experiment was done. Starting on day 10 after the first injection, tumor growth was monitored once every 7–10 days for 40 days according to the actual tumor formation and animal welfare ethics regulations (tumor diameter <20 mm). For luciferase luminescent detection, a dose of 10 µl/g D Luciferin (15 mg/mL) was injected intraperitoneally before anesthesia testing. Tumor images were captured using an IVIS® Lumina LT for whole live-animal imaging (PerkinElmer). Total fluorescence expression in the AVERAGE area was calculated as ([p/s]/[μW/cm2]). Tumor volume was calculated as [π/6 × L (length) × W (width) × H (height)]. All animal work was approved by the Institutional Ethics Committees of the First Affiliated Hospital of Chongqing Medical University.
Autophagic flux and quantification
To measure autophagic flux, we used monomeric LC3 proteins fused to a pH-stable RFP and a pH-sensitive GFP fluorophore. The mRFP-GFP-LC3 adenoviral particles were purchased from HanBio Technology (Shanghai, China). HCT116 and KYSE150 cells were infected with adenoviral particles according to the manufacturer’s instructions. After infection, cells were cultured for another 24 h for immunostaining. RFP punctate indicates both early autophagosome and autophagic lysosomes. Yellow punctate appearing after red and green fluorescence merged indicates early autophagosomes alone, as GFP fluorescence is quenched when autophagosomes fuse with lysosomes. The ratio of early autophagosomes over total autolysosomes was calculated as Flux %: Flux % = (100 – ((Red and Green)/Red) × 100). Images were captured by a fluorescence microscope Olympus BX51 microscope (Olympus Corporation, Tokyo, Japan).
Protein ubiquitination assay
Protein ubiquitination assay was performed as described previously [56,57,58]. Briefly, T-REx-293 cells with inducible SP0495 expression and KYSE150 cells with stable expression of SP0495 were lysed, or transfected with His-Ub plasmids for 48 h. After cell treatment with 10 μM MG132 (Sigma-Aldrich, Saint Louis, MO) for 6 h before harvest, endogenous BECN1 and p62 were immunoprecipitated with BECN1 and p62 antibodies, followed by immunoblot with anti-Ub or anti-His antibody to detect the ubiquitinated BECN1 and p62 proteins.
Ni-NTA pull-down assay was performed as previously described [56]. Cell lysates were affinity purified with Ni-NTA-agarose beads (#30210, Qiagen), and analyzed by immunoblot with specific antibodies targeting BECN1 and p62.
Lipid-protein overlay assay
PIP StripsTM and PIP ArraysTM (Echelon Biosciences) were blocked in TBST containing 3% BSA for 1 h at room temperature (RT) and incubated overnight at 4 °C with 0.5 µg/ml recombinant KIAA0495-ORF2/SP0495 protein (Origene, TP310801) and customized SP0495-mutant 1 and mutant 2 recombinant proteins produced with C-terminal DDK tag from human HEK293 cells (Origene) in TBST + 3% BSA. After three times washes, anti-KIAA0495-ORF2/SP0495 antibody (Origene, TA503634) was added to TBST + 3% BSA solution and incubated for 1 h at RT. Bound proteins were then detected using an HRP-coupled anti-mouse-IgG antibody, followed by visualization using the ECL detection system (GE Healthcare).
Additional materials and methods
Additional information on cell lines, tumor and normal tissue samples, array-comparative genomic hybridization (CGH), semi-quantitative RT-PCR and real-time PCR analyses, bisulfite treatment and promoter methylation analyses, plasmid constructs and generation of cell lines, immunofluorescence, immunohistochemical staining, colony formation assay, in vivo tumor formation assay, apoptosis and cell cycle analyses, Senescence-specific β-galactosidase staining, luciferase reporter assay, transmission electron microscopy, Western blot, protein stability analysis, generation of microarray data and online URLs, and statistical analyses are provided in Supplementary information.
Availability of data and materials
All data generated or analyzed during this study are included in this manuscript.
References
Henrich KO, Schwab M, Westermann F. 1p36 tumor suppression-a matter of dosage? Cancer Res. 2012;72:6079–88.
Bagchi A, Mills AA. The quest for the 1p36 tumor suppressor. Cancer Res. 2008;68:2551–6.
Solimini NL, Xu Q, Mermel CH, Liang AC, Schlabach MR, Luo J, et al. Recurrent hemizygous deletions in cancers may optimize proliferative potential. Science. 2012;337:104–9.
Kawano S, Miller CW, Gombart AF, Bartram CR, Matsuo Y, Asou H, et al. Loss of p73 gene expression in leukemias/lymphomas due to hypermethylation. Blood. 1999;94:1113–20.
Bagchi A, Papazoglu C, Wu Y, Capurso D, Brodt M, Francis D, et al. CHD5 is a tumor suppressor at human 1p36. Cell. 2007;128:459–75.
Xinh PT, Tri NK, Nagao H, Nakazato H, Taketazu F, Fujisawa S, et al. Breakpoints at 1p36.3 in three MDS/AML(M4) patients with t(1;3)(p36;q21) occur in the first intron and in the 5’ region of MEL1. Genes Chromosomes Cancer. 2003;36:313–6.
Lin N, Di C, Bortoff K, Fu J, Truszkowski P, Killela P, et al. Deletion or epigenetic silencing of AJAP1 on 1p36 in glioblastoma. Mol Cancer Res. 2012;10:208–17.
Ingolia NT, Brar GA, Stern-Ginossar N, Harris MS, Talhouarne GJ, Jackson SE, et al. Ribosome profiling reveals pervasive translation outside of annotated protein-coding genes. Cell Rep. 2014;8:1365–79.
Fields AP, Rodriguez EH, Jovanovic M, Stern-Ginossar N, Haas BJ, Mertins P, et al. A regression-based analysis of ribosome-profiling data reveals a conserved complexity to mammalian translation. Mol Cell. 2015;60:816–27.
Calviello L, Mukherjee N, Wyler E, Zauber H, Hirsekorn A, Selbach M, et al. Detecting actively translated open reading frames in ribosome profiling data. Nat Methods. 2016;13:165–70.
Chen J, Brunner AD, Cogan JZ, Nunez JK, Fields AP, Adamson B, et al. Pervasive functional translation of noncanonical human open reading frames. Science. 2020;367:1140–6.
Magny EG, Pueyo JI, Pearl FM, Cespedes MA, Niven JE, Bishop SA, et al. Conserved regulation of cardiac calcium uptake by peptides encoded in small open reading frames. Science. 2013;341:1116–20.
Andrews SJ, Rothnagel JA. Emerging evidence for functional peptides encoded by short open reading frames. Nat Rev Genet. 2014;15:193–204.
Prensner JR, Enache OM, Luria V, Krug K, Clauser KR, Dempster JM, et al. Noncanonical open reading frames encode functional proteins essential for cancer cell survival. Nat Biotechnol. 2021;39:697–704.
Huang JZ, Chen M, Chen, Gao XC, Zhu S, Huang H, et al. A peptide encoded by a putative lncRNA HOXB-AS3 suppresses colon cancer growth. Mol Cell. 2017;68:171–84.e176.
Meng N, Chen M, Chen, Chen XH, Wang JZ, Zhu S, et al. Small protein hidden in lncRNA LOC90024 promotes “cancerous” RNA splicing and tumorigenesis. Adv Sci. 2020;7:1903233.
Zhu S, Wang JZ, Chen D, He YT, Meng N, Chen M, et al. An oncopeptide regulates m(6)A recognition by the m(6)A reader IGF2BP1 and tumorigenesis. Nat Commun. 2020;11:1685.
Zhang M, Zhao K, Xu X, Yang Y, Yan S, Wei P, et al. A peptide encoded by circular form of LINC-PINT suppresses oncogenic transcriptional elongation in glioblastoma. Nat Commun. 2018;9:4475.
Wang Y, Wu S, Zhu X, Zhang L, Deng J, Li F, et al. LncRNA-encoded polypeptide ASRPS inhibits triple-negative breast cancer angiogenesis. J Exp Med. 2020;217:jem.20190950.
Mathew R, Karantza-Wadsworth V, White E. Role of autophagy in cancer. Nat Re Cancer. 2007;7:961–7.
Santana-Codina N, Mancias JD, Kimmelman AC. The role of autophagy in cancer. Annu Rev Cancer Biol. 2017;1:19–39.
Tyutyunyk-Massey L, Gewirtz DA. Roles of autophagy in breast cancer treatment: Target, bystander or benefactor. Semin Cancer Biol. 2020;66:155–62.
Tang J, Deng R, Luo RZ, Shen GP, Cai MY, Du ZM, et al. Low expression of ULK1 is associated with operable breast cancer progression and is an adverse prognostic marker of survival for patients. Breast Cancer Res Treat. 2012;134:549–60.
Li ZL, Zhang HL, Huang Y, Huang JH, Sun P, Zhou NN, et al. Autophagy deficiency promotes triple-negative breast cancer resistance to T cell-mediated cytotoxicity by blocking tenascin-C degradation. Nat Commun. 2020;11:3806.
Li Z, Chen B, Wu Y, Jin F, Xia Y, Liu X. Genetic and epigenetic silencing of the beclin 1 gene in sporadic breast tumors. BMC Cancer. 2010;10:98.
Rolland P, Madjd Z, Durrant L, Ellis IO, Layfield R, Spendlove I. The ubiquitin-binding protein p62 is expressed in breast cancers showing features of aggressive disease. Endocr Relat Cancer. 2007;14:73–80.
Luo RZ, Yuan ZY, Li M, Xi SY, Fu J, He J. Accumulation of p62 is associated with poor prognosis in patients with triple-negative breast cancer. Oncol Targets Ther. 2013;6:883–8.
Lu Z, Luo RZ, Lu Y, Zhang X, Yu Q, Khare S, et al. The tumor suppressor gene ARHI regulates autophagy and tumor dormancy in human ovarian cancer cells. J Clin Invest. 2008;118:3917–29.
Ying J, Li H, Seng TJ, Langford C, Srivastava G, Tsao SW, et al. Functional epigenetics identifies a protocadherin PCDH10 as a candidate tumor suppressor for nasopharyngeal, esophageal and multiple other carcinomas with frequent methylation. Oncogene. 2006;25:1070–80.
Jin H, Wang X, Ying J, Wong AH, Cui Y, Srivastava G, et al. Epigenetic silencing of a Ca(2+)-regulated Ras GTPase-activating protein RASAL defines a new mechanism of Ras activation in human cancers. Proc Natl Acad Sci USA. 2007;104:12353–8.
Kim S, Heo S, Brzostowski J, Kang D. Endosomal mTORC2 is required for phosphoinositide-dependent AKT activation in platelet-derived growth factor-stimulated glioma cells. Cancers. 2021;13:2405.
Dong J, Hu Z, Wu C, Guo H, Zhou B, Lv J, et al. Association analyses identify multiple new lung cancer susceptibility loci and their interactions with smoking in the Chinese population. Nat Genet. 2012;44:895–9.
Zhang H, Zhai Y, Hu Z, Wu C, Qian J, Jia W, et al. Genome-wide association study identifies 1p36.22 as a new susceptibility locus for hepatocellular carcinoma in chronic hepatitis B virus carriers. Nat Genet. 2010;42:755–8.
Stacey SN, Gudbjartsson DF, Sulem P, Bergthorsson JT, Kumar R, Thorleifsson G, et al. Common variants on 1p36 and 1q42 are associated with cutaneous basal cell carcinoma but not with melanoma or pigmentation traits. Nat Genet. 2008;40:1313–8.
Hu Z, Xia Y, Guo X, Dai J, Li H, Hu H, et al. A genome-wide association study in Chinese men identifies three risk loci for non-obstructive azoospermia. Nat Genet. 2011;44:183–6.
Corn PG, Kuerbitz SJ, van Noesel MM, Esteller M, Compitello N, Baylin SB, et al. Transcriptional silencing of the p73 gene in acute lymphoblastic leukemia and Burkitt’s lymphoma is associated with 5’ CpG island methylation. Cancer Res. 1999;59:3352–6.
Fujita T, Igarashi J, Okawa ER, Gotoh T, Manne J, Kolla V, et al. CHD5, a tumor suppressor gene deleted from 1p36.31 in neuroblastomas. J Natl Cancer Inst. 2008;100:940–9.
Gorringe KL, Choong DY, Williams LH, Ramakrishna M, Sridhar A, Qiu W, et al. Mutation and methylation analysis of the chromodomain-helicase-DNA binding 5 gene in ovarian cancer. Neoplasia. 2008;10:1253–8.
Mulero-Navarro S, Esteller M. Chromatin remodeling factor CHD5 is silenced by promoter CpG island hypermethylation in human cancer. Epigenetics. 2008;3:210–5.
Du Z, Li L, Huang X, Jin J, Huang S, Zhang Q, et al. The epigenetic modifier CHD5 functions as a novel tumor suppressor for renal cell carcinoma and is predominantly inactivated by promoter CpG methylation. Oncotarget. 2016;7:21618–30.
Seki N, Ohira M, Nagase T, Ishikawa K, Miyajima N, Nakajima D, et al. Characterization of cDNA clones in size-fractionated cDNA libraries from human brain. DNA Res. 1997;4:345–9.
Pang JC, Li KK, Lau KM, Ng YL, Wong J, Chung NY, et al. KIAA0495/PDAM is frequently downregulated in oligodendroglial tumors and its knockdown by siRNA induces cisplatin resistance in glioma cells. Brain Pathol. 2010;20:1021–32.
Wu P, Mo Y, Peng M, Tang T, Zhong Y, Deng X, et al. Emerging role of tumor-related functional peptides encoded by lncRNA and circRNA. Mol Cancer. 2020;19:22.
Hartford CCR, Lal A. When long noncoding becomes protein coding. Mol Cell Biol. 2020;40:e00528–19.
Huang N, Li F, Zhang M, Zhou H, Chen Z, Ma X, et al. An upstream open reading frame in phosphatase and tensin homolog encodes a circuit breaker of lactate metabolism. Cell Metab. 2021;33:128–44.e129.
Alessi DR, James SR, Downes CP, Holmes AB, Gaffney PR, Reese CB, et al. Characterization of a 3-phosphoinositide-dependent protein kinase which phosphorylates and activates protein kinase Balpha. Curr Biol. 1997;7:261–9.
Li B, Cheung PY, Wang X, Tsao SW, Ling MT, Wong YC, et al. Id-1 activation of PI3K/Akt/NFkappaB signaling pathway and its significance in promoting survival of esophageal cancer cells. Carcinogenesis. 2007;28:2313–20.
Amiri A, Noei F, Jeganathan S, Kulkarni G, Pinke DE, Lee JM. eEF1A2 activates Akt and stimulates Akt-dependent actin remodeling, invasion and migration. Oncogene. 2007;26:3027–40.
Garnett MJ, Edelman EJ, Heidorn SJ, Greenman CD, Dastur A, Lau KW, et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature. 2012;483:570–5.
Maiuri MC, Tasdemir E, Criollo A, Morselli E, Vicencio JM, Carnuccio R, et al. Control of autophagy by oncogenes and tumor suppressor genes. Cell Death Differ. 2009;16:87–93.
Vicinanza M, Korolchuk VI, Ashkenazi A, Puri C, Menzies FM, Clarke JH, et al. PI(5)P regulates autophagosome biogenesis. Mol Cell. 2015;57:219–34.
Hasegawa J, Strunk BS, Weisman LS. PI5P and PI(3,5)P2: minor, but essential phosphoinositides. Cell Struct Funct. 2017;42:49–60.
Dall’Armi C, Devereaux KA, Di, Paolo G. The role of lipids in the control of autophagy. Curr Biol. 2013;23:R33–45.
Fang Y, Vilella-Bach M, Bachmann R, Flanigan A, Chen J. Phosphatidic acid-mediated mitogenic activation of mTOR signaling. Science. 2001;294:1942–5.
Li L, Zhang Y, Fan Y, Sun K, Su X, Du Z, et al. Characterization of the nasopharyngeal carcinoma methylome identifies aberrant disruption of key signaling pathways and methylated tumor suppressor genes. Epigenomics. 2015;7:155–73.
Li L, Li W, Xiao L, Xu J, Chen X, Tang M, et al. Viral oncoprotein LMP1 disrupts p53-induced cell cycle arrest and apoptosis through modulating K63-linked ubiquitination of p53. Cell Cycle. 2012;11:2327–36.
Li L, Li Z, Zhou S, Xiao L, Guo L, Tao Y, et al. Ubiquitination of MDM2 modulated by Epstein-Barr virus encoded latent membrane protein 1. Virus Res. 2007;130:275–80.
Li L, Tao Q, Jin H, van Hasselt A, Poon FF, Wang X, et al. The tumor suppressor UCHL1 forms a complex with p53/MDM2/ARF to promote p53 signaling and is frequently silenced in nasopharyngeal carcinoma. Clin Cancer Res. 2010;16:2949–58.
Acknowledgements
We thank Drs B Vogelstein, G Srivastava, GSW Tsao, Sun Young Rha, the late Dolly Huang and KT Yao for cell lines; DSMZ (Germany) for KYSE cell lines (Shimada et al., Cancer 69: 277–284 (1992); Kazusa DNA Research Institute for the full-length clone of KIAA0495; Drs Christof Niehrs and Dong-Yan Jin for TOPFlash and FOPFlash constructs; Dr. Ben CB Ko for autophagy assay, and FanFong Poon for some technical support.
Funding
This study was supported by HK-RGC (#14115019; #14115920; #14102923), NSFC (#81772869), China MOST National Key Research and Development Program (#2017YFE0191700), and Johns Hopkins Singapore/A*STAR.
Author information
Authors and Affiliations
Contributions
QT, LL: conceptual design; LL, XS, HG, JY, LG, JL, TX, LW: performed experiments; BBM, ATC, XZ, RFA: provided materials and reviewed manuscript; LL, XS, HG, QT: drafted the manuscript; LL, QT: finalized the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The animal experiments were performed according to guidelines approved by the Institutional Ethics Committees of the First Affiliated Hospital of Chongqing Medical University and abided by the Declaration of Helsinki.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Consent for publication All authors have agreed to publish this manuscript.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, L., Shu, Xs., Geng, H. et al. A novel tumor suppressor encoded by a 1p36.3 lncRNA functions as a phosphoinositide-binding protein repressing AKT phosphorylation/activation and promoting autophagy. Cell Death Differ 30, 1166–1183 (2023). https://doi.org/10.1038/s41418-023-01129-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-023-01129-w