Abstract
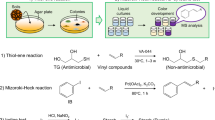
The discovery of novel therapeutic antimicrobials has become an urgent issue in response to the global crisis of the spread of multi-drug-resistant bacteria. In this report, we propose an efficient screening method for antimicrobial agents with therapeutic potential from soil bacteria. With this method, colonies of the soil bacteria were formed first on agar plates containing only an extract of soil, followed by an overlay of soft agar containing the pathogens, an antibiotic target. Then, we selected the colonies that formed the inhibitory zones on soft agar and evaluated the therapeutic efficacy of their culture supernatants using a silkworm bacterial infection model. Using Staphylococcus aureus as an indicator strain to obtain bacteria that produce therapeutically effective antimicrobials, we succeeded in reducing the screening size by 20-fold compared to the conventional method. An analysis of 86 antibiotics producers identified in this study indicated that the majority belonged to Streptomyces sp. and Lysobacter sp., well-known producers of secondary metabolites. Besides, the presence of eight genera and 37 species among the identified species indicated the diversity of antibiotic producers. Based on the finding of our study, we propose this method as an efficient way to discover novel antimicrobial agents that are therapeutically effective.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Tsuzuki S, Matsunaga N, Yahara K, Gu Y, Hayakawa K, Hirabayashi A, et al. National trend of blood-stream infection attributable deaths caused by Staphylococcus aureus and Escherichia coli in Japan. J Infect Chemother. 2020;26:367–71.
Kadri SS. Key takeaways from the U.S. CDC’s 2019 antibiotic resistance threats report for frontline providers. Crit Care Med. 2020;48:939–45.
Butler MS, Paterson DL. Antibiotics in the clinical pipeline in October 2019. J Antibiot (Tokyo). 2020;73:329–64.
Ling LL, Schneider T, Peoples AJ, Spoering AL, Engels I, Conlon BP, et al. A new antibiotic kills pathogens without detectable resistance. Nature. 2015;517:455–9.
Hamamoto H, Urai M, Ishii K, Yasukawa J, Paudel A, Murai M, et al. Lysocin E is a new antibiotic that targets menaquinone in the bacterial membrane. Nat Chem Biol. 2015;11:127–33.
Hayakawa M, Nonomura H. Humic acid-vitamin agar, a new medium for the selective isolation of soil actinomycetes. J Ferment Technol. 1987;65:501–9.
Nett M, Ikeda H, Moore BS. Genomic basis for natural product biosynthetic diversity in the actinomycetes. Nat Prod Rep. 2009;26:1362–84.
Panthee S, Hamamoto H, Suzuki Y, Sekimizu K. In silico identification of lysocin biosynthetic gene cluster from Lysobacter sp. RH2180-5. J Antibiot (Tokyo). 2017;70:204–7.
Panthee S, Hamamoto H, Paudel A, Sekimizu K. Lysobacter species: a potential source of novel antibiotics. Arch Microbiol. 2016;198:839–45.
Hamamoto H, Kurokawa K, Kaito C, Kamura K, Manitra Razanajatovo I, Kusuhara H, et al. Quantitative evaluation of the therapeutic effects of antibiotics using silkworms infected with human pathogenic microorganisms. Antimicrob Agents Chemother. 2004;48:774–9.
Shoji J, Hinoo H, Matsumoto K, Hattori T, Yoshida T, Matsuura S, et al. Isolation and characterization of katanosins A and B. J Antibiot (Tokyo). 1988;41:713–8.
Itoh H, Tokumoto K, Kaji T, Paudel A, Panthee S, Hamamoto H, et al. Total synthesis and biological mode of action of WAP-8294A2: a menaquinone-targeting antibiotic. J Org Chem. 2017;83:6924–35.
Lewis K. Platforms for antibiotic discovery. Nat Rev Drug Disco. 2013;12:371–87.
James N. Soil extract in soil microbiology. Can J Microbiol. 1958;4:363–70.
Taylor CB. Nature of the factor in soil-extract responsible for bacterial growth-stimulation. Nature. 1951;168:115–6.
Nguyen TM, Seo C, Ji M, Paik MJ, Myung SW, Kim J. Effective soil extraction method for cultivating previously uncultured soil bacteria. Appl Environ Microbiol. 2018;84:e01145-18.
Hamaki T, Suzuki M, Fudou R, Jojima Y, Kajiura T, Tabuchi A, et al. Isolation of novel bacteria and actinomycetes using soil-extract agar medium. J Biosci Bioeng. 2005;99:485–92.
Hayakawa M, Yoshida Y, Iimura Y. Selective isolation of bioactive soil actinomycetes belonging to the Streptomyces violaceusniger phenotypic cluster. J Appl Microbiol. 2004;96:973–81.
Craig JW, Chang FY, Kim JH, Obiajulu SC, Brady SF. Expanding small-molecule functional metagenomics through parallel screening of broad-host-range cosmid environmental DNA libraries in diverse proteobacteria. Appl Environ Microbiol. 2010;76:1633–41.
Nkanga EJ, Hagedorn C. Detection of antibiotic-producing Streptomyces inhabiting forest soils. Antimicrob Agents Chemother. 1978;14:51–9.
Panthee S, Paudel A, Hamamoto H, Sekimizu K. Advantages of the silkworm as an animal model for developing novel antimicrobial agents. Front Microbiol. 2017;8:373.
Paudel A, Hamamoto H, Panthee S, Kaneko K, Matsunaga S, Kanai M, et al. A novel spiro-heterocyclic compound identified by the silkworm infection model inhibits transcription in Staphylococcus aureus. Front Microbiol. 2017;8:712.
Yi J, Qinyuan L, Xiu C, Chenglin J. Isolation and cultivation methods of actinobacteria. In: Dhanasekaran D, Jiang Y (eds). Basics and Biotechnological Application. IntechOpen, 2016, pp 39–57.
Akimitsu N, Hamamoto H, Inoue R, Shoji M, Akamine A, Takemori K, et al. Increase in resistance of methicillin-resistant Staphylococcus aureus to beta-lactams caused by mutations conferring resistance to benzalkonium chloride, a disinfectant widely used in hospitals. Antimicrob Agents Chemother. 1999;43:3042–3.
Hamamoto H, Kamura K, Razanajatovo IM, Murakami K, Santa T, Sekimizu K. Effects of molecular mass and hydrophobicity on transport rates through non-specific pathways of the silkworm larva midgut. Int J Antimicrob Agents. 2005;26:38–2.
Acknowledgements
This work was supported by Genome Pharmaceutical Institute, Co., Ltd. and in part by JSPS KAKENHI Grant Numbers 19K07140JP to HH, and in part by JSPS KAKENHI Grant Numbers 15H05783 and 21H02733 to KS and HH, and TBRF, IFO fellowships to SP and KS. We thank Mr. Takashi Yamashita, Ms. Mari Maeda and Ms. Kiyomi Kyogoku for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
KS is a consultant for Genome Pharmaceutical Institute Co, Ltd. The authors declare competing financial interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Hamamoto, H., Panthee, S., Hashimoto, K. et al. An efficient method to screen for the soil bacteria producing therapeutically effective antibiotics. J Antibiot 74, 850–855 (2021). https://doi.org/10.1038/s41429-021-00476-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-021-00476-5