Abstract
Purpose
The SERPINA1 Z allele is associated with cystic fibrosis (CF)-related liver disease (CFLD), a common manifestation in patients with CF. We estimated CFLD incidence based on the SERPINA1 genotype in 3328 CF patients with CFLD-phenotype information.
Methods
The associations of SERPINA1 Z (rs28929474) and S (rs17580) alleles with age at CFLD onset and the development of CFLD-related complications (severe liver disease with cirrhosis, portal hypertension, esophageal varices) were analyzed.
Results
Overall, 3% of patients carried the SERPINA1 Z allele and 13% carried the S allele. The cumulative incidence of CFLD increased more rapidly in patients carrying the Z allele (hazard ratio [HR] = 1.6; 95% confidence interval [CI] = 1.1–2.4, P = 0.019), reaching 47% by age 25 compared with 30% in noncarriers. Increased risk was similar for patients with severe CFLD (HR = 1.5, 95% CI = 0.7–3.2, P = 0.31) but failed to reach significance due to a limited sample size of Z-allele carriers. No significant effect was found for the S allele.
Conclusion
CF patients carrying the SERPINA1 Z allele had an increased risk of developing CFLD and related complications compared with noncarriers. Routine SERPINA1 Z genotyping upon CF diagnosis is warranted for identifying patients worthy of closer liver disease monitoring.
Similar content being viewed by others
INTRODUCTION
Cystic fibrosis (CF) is the most common, severe, autosomal recessive genetic disease in Caucasians. Pathogenic variants in the CF transmembrane conductance regulator (CFTR) gene induce a multisystemic disease affecting several organs such as the lungs, pancreas, intestine, and liver.1 A broad spectrum of hepatobiliary abnormalities are covered under the term CF-related liver disease (CFLD). Focal biliary cirrhosis is the most clinically important form of CFLD, since extension of the initially focal fibrogenic process may cause multilobular biliary cirrhosis followed by portal hypertension and associated complications.2,3 Multilobular cirrhosis ranks nowadays as the third leading cause of death in patients with CF, after respiratory failure and transplantation-related complications.4
Although CF is recognized as a monogenic disease, a large interindividual variability in phenotype exists among patients with the same CFTR pathogenic variants.5,6,7 In addition to environmental factors, genetic modifiers are recognized to contribute to this variability.5,6 We have assembled a large cohort of patients as part of the French CF Modifier Gene Study and recently reported the incidence of CFLD and severe CFLD in these patients.8 We found that the risk of CFLD increases with age, with a frequency up to 32% by age 25, and is associated with several risk factors, including male sex, CFTR F508del homozygosity, and a history of meconium ileus at birth.
The SERPINA1 gene has been implicated in the development and progression of CFLD.9 SERPINA1 encodes the ɑ-1 antitrypsin (AAT) protein synthesized by the liver. Several pathogenic variants of SERPINA1 have been determined to cause an AAT deficiency that predisposes an individual to liver disease and early-onset emphysema. The most common variants involve the Z and S alleles, each caused by single-nucleotide polymorphisms.10 The Z variant is the allele overwhelmingly associated with liver disease.11 Indeed, during biogenesis the Z-type AAT protein folds abnormally in the endoplasmic reticulum of hepatocytes and is retained intracellularly instead of being efficiently secreted, resulting in low serum levels of AAT.12 The intracellular accumulation of AAT mutant Z proteins within hepatocytes can lead to liver injury, cirrhosis, or hepatocellular carcinoma.13 Individuals heterozygous for AAT that carry one normal protease inhibitor M allele and one pathogenic variant Z allele (PiMZ or MZ) are asymptomatic with regard to liver disease. Compound heterozygotes for the S and the Z alleles of AAT (PiSZ) may develop liver disease with identical manifestations to those of PiZZ patients, whereas liver disease is absent in PiSS homozygote individuals.13 A CF case–control study revealed that carriers of the SERPINA1 Z allele are more common among patients with severe CFLD, with a large odds ratio of 5 (ref. 9). However, the study focused on patients with severe liver disease defined as cirrhosis showing signs of portal hypertension, which prevented the determination of the actual risk associated with the variant for developing CFLD. Therefore, in the present study, we estimated the cumulative incidence of CFLD based on the SERPINA1 genotype in a cohort of French CF patients of unprecedented size (n = 3328).
MATERIALS AND METHODS
As previously described, we assembled the French CF Modifier Gene Study cohort that included CF patients treated at French CF centers since 2004 (see also Supplementary Information).8 In brief, 4798 patients with CF were recruited for the study, corresponding to approximately 80% of all French patients with CF.14 Among this cohort, 3328 CF patients with pancreatic insufficiency born after 1985 were available for evaluation of CFLD incidence and risk factors, including patients with severe CFLD.8 CFLD was defined according to the European Best Practice Guidance by Debray et al.2 Patients with cirrhosis, portal hypertension, and/or esophageal varices were classified as having severe CFLD (see also Supplementary Information).3,15 The study was approved by the French ethical committee (CPP number 2004/15) and the information collection was approved by the Commission Nationale de L’informatique et des Libertés (number 04.404). Written informed consent was obtained from each patient and/or guardian.
Genotyping of the SERPINA1 Z (rs28929474) and S (rs17580) alleles was carried out using Kompetitive Allele Specific PCR (KASP) genotyping chemistry (LGC, Teddington, UK). In the dbSNP database (https://www.ncbi.nlm.nih.gov/projects/SNP/), rs28929474 is identified as an A/G variant with G being the ancestral allele, and rs17580 is identified as an A/T variant with A being the ancestral allele.
Descriptive statistics were compiled as the mean ± standard deviation (SD) or percentages as appropriate. All patients were considered to be at risk for CFLD since birth and were censored at the time of the last visit without a CFLD diagnosis before January 2017. For other patients, the age at CFLD diagnosis was determined using the date of the first report in the medical records, allowing for “interval censoring” between birth and the age of the first report when the date of onset was not precisely known. This was the case for 142 of 605 patients with CFLD (24%). Likewise, severe CFLD onset was defined as the first date that cirrhosis, portal hypertension, and/or esophageal varices were reported and uncertainty regarding this date was considered as described above. The date of severe CFLD was interval censored in 19 of 175 patients (11%). We used the log-rank test adapted for interval-censored data for comparisons between the cumulative incidences curves16 and Cox regression adapted to interval-censored data to determine the association of factors linked to age at CFLD onset.17 Confidence intervals were computed by the bootstrap method. Bonferroni correction was used for multiple comparisons.
RESULTS
Clinical characteristics of the 3328 CF patients included in this study, along with the distribution of the SERPINA1 Z and S alleles, are shown in Table 1. The minor allele frequencies were similar in our cohort to those reported for Europeans: SERPINA1 Z (A variant), 1.4% vs. 2% respectively; SERPINA1 S (T variant), 6.4% versus 6% respectively. There were no patients homozygous for SERPINA1 Z. Details on the number of CF patients at risk of developing CFLD and severe CFLD, and the cumulative number of CFLD and severe CFLD events for the entire cohort as well as according to SERPINA1 Z and S genotypes are provided in the Supplementary Information (Table S1, S2, and S3).
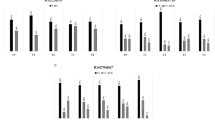
Overall, 3% of the CF patients carried the SERPINA1 Z allele and 13% carried the S allele. The cumulative incidence of CFLD increased more rapidly in patients carrying the SERPINA1 Z allele (hazard ratio [HR] = 1.6; 95% confidence interval [CI] = 1.1–2.4, P = 0.019), reaching 47% by age 25 in the Z allele carriers compared with 30% at age 25 in the others (Table 1 and Fig. 1). The increase in risk was similar for patients with severe CFLD (HR = 1.5, 95% CI = 0.7–3.2, P = 0.31), but this did not reach statistical significance. With respect to severe CFLD, there were only seven cases among the Z allele carriers, making the cumulative incidence curve difficult to accurately estimate. Clinical characteristics such as sex, year of birth, and year of CF diagnosis were not associated with SERPINA1 Z and S alleles (Supplementary Information Table S4). Adjusting on European origin, CFTR variants and meconium ileus did not change the strength of the association (Supplementary Information Table S5). The effect of carrying one SERPINA1 S allele on CFLD risk was not statistically significant (Table 1 and Fig. 1).
Cumulative incidence of cystic fibrosis-related liver disease (CFLD) (a, c) and severe CFLD (b, d) according to SERPINA1 Z (a, b) and S (c, d) alleles. In each graph, the solid line curve indicates the cumulative incidence for patients carrying the normal M allele (G and A for SERPINA1 Z and S, respectively) and the dotted line curve indicates the cumulative incidence for patients carrying the pathogenic variant (A and T for SERPINA1 Z and S, respectively). Below each graph is indicated the number of patients at a specific age who continue to be followed up but have not yet developed CFLD or severe CFLD according to SERPINA1 Z and S alleles.
DISCUSSION
The French CF Modifier Gene Study provided an unprecedentedly large cohort of 3328 pancreatic-insufficient patients with CF born after 1985. This gave an opportunity to obtain a more accurate estimate of the incidence of CFLD and severe CFLD with sufficient power to detect associations of clinical relevance.8 We found that the SERPINA1 Z allele was associated with an increased risk of developing CFLD, although the association was weaker than that previously reported.9 Nevertheless, the incidence of CFLD increased more rapidly in patients carrying the SERPINA1 Z allele, with up to 47% of the Z allele carriers developing liver disease before the age of 25 compared with only 30% for noncarrier patients.
The role of SERPINA1 in CFLD was first identified in a two-stage case–control study including CF patients from several countries worldwide.9 Both the initial and a replicate studies showed that severe CFLD is associated with the SERPINA1 Z allele (odds ratio of 4.72 and 3.42, respectively).9 Analysis of our cohort of patients with CF confirmed this association, but demonstrated a smaller difference in risk than that reported previously. For example, using the cumulative incidence at age 25, the odds ratio for the Z allele was only 2 in our cohort. However, the difference in cumulative risk with age was still clinically relevant because SERPINA1 Z carriers had a 50% greater risk of developing CFLD compared with noncarriers. We did not observe any association of the SERPINA1 S allele with CFLD, which was not surprising because the S allele is recognized to be associated with reduced levels of AAT protein, but not with liver manifestations.18
Our study had limitations related to its design and the use of medical records as a primary source of information. However, we previously reported that selection bias due to differential mortality was likely to be small.8 A second potential issue was the rarity of the SERPINA1 Z allele in our cohort. This reduced the precision of estimation, despite the large cohort size. Indeed, there were only 90 carriers of the SERPINA1 Z allele (PiMZ) and no homozygous PiZZ patients in the cohort. Furthermore, only seven of these patients experienced severe CFLD, making the cumulative incidence difficult to accurately estimate.
Obtaining a more accurate prediction of the risk of CFLD in CF patients remains an important issue, especially because ursodeoxycholic acid, the treatment commonly prescribed for its prevention, seems to have little to no effect.8,19 Given this situation, identifying biomarkers to predict the occurrence of CFLD is fundamental to improving the monitoring of disease progression and for assessing the effects of novel therapies, such as bile salt analogs, antifibrotics, and CFTR correctors and potentiators. SERPINA1 Z genotyping at the time of CF diagnosis may help to single out a population of patients who should be more closely screened for liver disease. Gaining a better understanding of the genetic profiles of patients with CFLD will undoubtedly open new therapeutic avenues and help to develop prospective therapies that focus on high-risk groups.
References
Riordan JR, Rommens JM, Kerem B, et al. Identification of the cystic fibrosis gene: cloning and characterization of complementary DNA. Science. 1989;245:1066–1073.
Debray D, Kelly D, Houwen R, Strandvik B, Colombo C. Best practice guidance for the diagnosis and management of cystic fibrosis-associated liver disease. J Cyst Fibros. 2011;10 suppl 2:S29–36.
Stonebraker JR, Ooi CY, Pace RG, et al. Features of severe liver disease with portal hypertension in patients with cystic fibrosis. Clin Gastroenterol Hepatol. 2016;14:1207–1215.
Rowland M, Gallagher C, Gallagher CG, et al. Outcome in patients with cystic fibrosis liver disease. J Cyst Fibros. 2015;14:120–126.
Cutting GR. Cystic fibrosis genetics: from molecular understanding to clinical application. Nat Rev Genet. 2015;16:45–56.
Guillot L, Beucher J, Tabary O, Le Rouzic P, Clement A, Corvol H. Lung disease modifier genes in cystic fibrosis. Int J Biochem Cell Biol. 2014;52:83–93.
McKone EF, Emerson SS, Edwards KL, Aitken ML. Effect of genotype on phenotype and mortality in cystic fibrosis: a retrospective cohort study. Lancet. 2003;361:1671–1676.
Boelle PY, Debray D, Guillot L, Clement A, Corvol H, French CFMGSI. Cystic fibrosis liver disease: outcomes and risk factors in a large cohort of French patients. Hepatology. 30 2018.
Bartlett JR, Friedman KJ, Ling SC, et al. Genetic modifiers of liver disease in cystic fibrosis. JAMA. 2009;302:1076–1083.
Bartels CL, Marchetti AL, Edward Highsmith W, Tsongalis GJ. Real time PCR detection of the PI*Z and PI*S mutations associated with alpha-1 antitrypsin deficiency. Am J Transl Res. 2009;1:406–411.
Zorzetto M, Russi E, Senn O, et al. SERPINA1 gene variants in individuals from the general population with reduced alpha1-antitrypsin concentrations. Clin Chem. 2008;54:1331–1338.
Stoller JK, Aboussouan LS. A review of alpha1-antitrypsin deficiency. Am J Respir Crit Care Med. 2012;185:246–259.
Teckman JH. Liver disease in alpha-1 antitrypsin deficiency: current understanding and future therapy. COPD. 2013;10 suppl 1:35–43.
Vaincre-la-Mucoviscidose, Ined. Registre Français de la mucoviscidose - Bilan des données 2016. National report, 2017.
Debray D, Narkewicz MR, Bodewes F, et al. Cystic fibrosis-related liver disease: research challenges and future perspectives. J Pediatr Gastroenterol Nutr. 2017;65:443–448.
Fay MP. Rank invariant tests for interval censored data under the grouped continuous model. Biometrics. 1996;52:811–822.
Pan W. Extending the iterative convex minorant algorithm to the Cox model for interval-censored data. J Comput Graph Stat. 1999;8:109–120.
Torres-Duran M, Lopez-Campos JL, Barrecheguren M, et al. Alpha-1 antitrypsin deficiency: outstanding questions and future directions. Orphanet J Rare Dis. 2018;13:114.
Cheng K, Ashby D, Smyth RL. Ursodeoxycholic acid for cystic fibrosis-related liver disease. Cochrane Database Syst Rev. 2017;9:CD000222.
Acknowledgements
The authors would like to thank the clinical research assistants involved in the data collection, namely, Pauline Touche, Tania Kombi, and Julie Mésinèle. We would like to especially thank the patients and families who participated in this study and the contributing French CF Centers (see the list of investigators in the Supplementary Information). This work was supported by the Institut National de la Santé et de la Recherche Médicale (INSERM), Sorbonne Université, Assistance Publique-Hôpitaux de Paris (APHP), Agence Nationale de la Recherche (ANR), DGS, Association Vaincre La Mucoviscidose, Cystic Fibrosis Foundation (CORVOL16G0), Association Agir Informer Contre la Mucoviscidose, GIS-Institut des Maladies Rares, and Société Française de la Mucoviscidose.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Co-last authors Loic Guillot and Harriet Corvol
Supplementary information
Rights and permissions
About this article
Cite this article
Boëlle, PY., Debray, D., Guillot, L. et al. SERPINA1 Z allele is associated with cystic fibrosis liver disease. Genet Med 21, 2151–2155 (2019). https://doi.org/10.1038/s41436-019-0449-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-019-0449-6