Abstract
DNA methylation can be environmentally modulated and plays a role in phenotypic plasticity. To understand the role of environmentally induced epigenetic variation and its dynamics in natural populations and ecosystems, it is relevant to place studies in a real-world context. Our experimental model is the wild potato Solanum kurtzianum, a close relative of the cultivated potato S. tuberosum. It was evaluated in its natural habitat, an arid Andean region in Argentina characterised by spatial and temporal environmental fluctuations. The dynamics of phenotypic and epigenetic variability (with Methyl Sensitive Amplified Polymorphism markers, MSAP) were assayed in three genotypes across three growing seasons. These genotypes were cultivated permanently and also reciprocally transplanted between experimental gardens (EG) differing in ca. 1000 m of altitude. In two seasons, the genotypes presented differential methylation patterns associated to the EG. In the reciprocal transplants, a rapid epigenomic remodelling occurred according to the growing season. Phenotypic plasticity, both spatial (between EGs within season) and temporal (between seasons), was detected. The epigenetic and phenotypic variability was positively correlated. The lack of an evident mitotic epigenetic memory would be a common response to short-term environmental fluctuations. Thus, the environmentally induced phenotypic and epigenetic variation could contribute to populations persistence through time. These results have implications for understanding the great ecological diversity of wild potatoes.
Similar content being viewed by others
Introduction
Natural environments can be subjected to modifications by anthropic interventions, natural disasters, and climate changes, among other factors. In particular, climate change will likely have an impact on plant ecophysiology, distribution, and community interactions (IPCC 2014). To face climate challenges, the presence of variability within and among natural populations is fundamental because it can provide genetic flexibility in changing environments (Nicotra et al. 2010). In sessile organisms such as plants, individual genotypes could optimise their performance through phenotypic plasticity, a phenomenon in which alternative phenotypes are expressed by the same genotype in diverse environments, and whose potential in plant evolution is being actually considered (Nicotra et al. 2010; Cooper et al. 2019; Kelly 2019). Phenotypic plasticity allows individuals to rapidly cope with environmental fluctuations (Kooke et al. 2015; Nicotra et al. 2015), which could result from climate change.
Phenotypic diversity originates from genetic variation (i.e., allele diversity as a result of differences in DNA sequence; Medrano et al. 2014) as well as from epigenetic variation (phenotypic changes unrelated to variation in DNA sequence; Richards 2006; Medrano et al. 2014). Cytosine DNA methylation, the most widely studied epigenetic mechanism, can be modulated in response to stressful conditions (Verhoeven et al. 2010; 2016) and can impact ecologically important phenotypic traits (Cubas et al. 1999; Marfil et al. 2009, 2012; Zhang et al. 2013; Cortijo et al. 2014).
In numerous plant studies carried out under controlled artificial conditions, the role of epigenetic variation has been evaluated as a source of intraspecific diversity (Tricker et al. 2012; Latzel et al. 2013; Zhang et al. 2013). The ecological consequences of this phenomenon are now being considered by performing studies under natural conditions. Epigenetic diversity in natural populations may contribute to affront diverse environmental conditions (Lira-Madeiros et al. 2010; Herrera and Bazaga 2011; Dubin et al. 2015; Richards et al. 2017) and the phenotypic variation caused by epigenetic changes could be a rapid acclimation mechanism to environmental fluctuations (Gao et al. 2010; Nicotra et al. 2015). In higher plants, however, there is little evidence on environmentally induced epigenetic changes and phenotypic plasticity during the lifespan of a single organism as well as on the dynamics of the phenomenon in natural conditions.
One of the most important non-cereal food crop worldwide (FAO UN 2018), the common potato Solanum tuberosum L., has hundreds of wild tuber-bearing relatives (Solanum spp., section Petota subsection Potatoe) distributed throughout the America, which are collectively known as ‘wild potatoes’. Wild potatoes present a great ecological diversity and can be found in even contrasting ecosystems such as deserts, humid forests, high Andean valleys or coastal regions (Spooner 2009). Phenotypic plasticity has been acknowledged as an essential attribute of this plant group (Correll 1962; Hawkes and Hjerting 1969; Masuelli et al. 2009; Camadro 2012), and numerous studies have attempted to shed light on the mechanisms that make this diversity possible. However, studies—carried out with germplasm bank accessions—have revealed that there is a lack of correlation between geographical distribution and genetic and phenotypic variability (McGregor et al. 2002; Jansky et al. 2009). Also, genotypes with different levels of pest and disease resistance coexist in the same populations (Jansky et al. 2006; Marfil and Masuelli 2014). These results would indicate that genetic variability and, possibly, other mechanisms contribute to generate the extensive natural variability of wild potatoes (Marfil et al. 2009; 2012; 2018; Cara et al. 2013).
Solanum kurtzianum Bitter and Wittm. is a diploid wild potato that grows in an arid region in western Argentina ranging in altitude from 1082 to 2490 m a.s.l. (Hijmans et al. 2002). Listed in the Global Priority Crop Wild Relative Inventory, and successfully used in potato breeding (Marfil et al. 2015), it is considered the best-adapted of all Argentinian taxonomic species to dry environments. As other species in subsection Potatoe, it has two alternative modes of reproduction with important roles in natural populations: sexual, through botanical seeds, relevant for populations establishment and as a source of genetic variability; and, asexual, through stolons and tubers, which play a role in population maintenance under several environmental conditions (Marfil and Masuelli 2014; Marfil et al. 2015).
One approach to address whether natural populations are able to respond to climate change is to perform common garden experiments under natural conditions (Berend et al. 2019). Along this line, it is advisable to use genetically uniform backgrounds to obtain evidence about the effect of the environment on both epigenetic variability and phenotypic plasticity. Hence, clonal plants established in experimental gardens (EGs) represent an excellent experimental model for this type of studies. In accordance with these considerations, the present work was performed with three genotypes of S. kurtzianum growing in its natural habitats: the Villavicencio Natural Reserve (VNR). This reserve is an Andean Protected Area in Western Argentina with contrasting environmental conditions, and it has been the centre of previous studies by our research group (Ibañez et al. 2017). The aim of the present work was to evaluate the phenotypic plasticity and the possible establishment of differential DNA methylation patterns and their dynamics in biological replicates (clones) growing at contrasting altitudes along three growing seasons. We aspire to answer: what is the extent at which environmental factors imposed by in situ growing conditions can induce phenotypic and epigenetic changes and, moreover, the stability of these changes if they actually occur.
Materials and methods
Plant material
We evaluated morphological, biochemical, and phenological phenotypes and analysed the dynamics of DNA methylation patterns in clones of three S. kurtzianum genotypes. These clones were cultivated during 3 successive years, both permanently and as reciprocally transplanted, in two Andean EGs, located at 1141 and 2113 m a.s.l. in the VNR. To establish the EGs, and according to the number of available tubers, one genotype from each of the three populations collected in the VNR and studied by Marfil and Masuelli (2014)—1228M, QH5 and 2166M—was selected and labelled as G-1, G-2, and G-3, respectively. Twelve tubers (clones) per genotype were cultivated in pots at the campus of Facultad de Ciencias Agrarias, UNCuyo (33° 00′24″ S, 68° 52′19″ W, 940 m a.s.l.) from November 2013 to January 2014.
EGs and reciprocal transplant approach
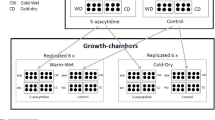
When the pot-grown plants presented three fully expanded leaves, they were transplanted into 10-L pots with a mixture of autoclaved local soil and organic compost (3:1), to prevent seed contamination. These pots were used to establish the field experiment in January 2014, in two EGs located in the VNR. One EG (hereinafter called ‘EG-1100’) was located at 1141 m a.s.l. (S32° 34′39.57″ S, 68° 56′45.65″ W), and the other (hereinafter called ‘EG-2100’) at 2113 m a.s.l. (32° 35′06.47″ S, 69° 05′56.85″ W). Half of the clones of each genotype were placed in each EG following a completely random design (Ibañez et al. 2017). Overall, six clones × three genotypes × two EG were evaluated. Plants were weekly irrigated with KSC II PHYT-actyl fertiliser (Timac Agro, USA). In September 2014, tubers formed during the growing season (November 2013 to April–May 2014) were collected and conserved in the laboratory at 4 °C until dormancy breakdown. The sprouted tubers were used as starting material for the next growing season. The cultivation conditions were repeated with the modifications detailed below (Fig. 1).
Three Solanum kurtzianum genotypes were cultivated in two experimental gardens located within an Andean Protected Area at 1100 and 2100 m a.s.l. (EG-1100 and EG-2100, respectively) during three successive seasons (2015, 2016 and 2017). As an example, only one genotype is shown. For each growing season, tubers (clonal propagules) obtained from the previous growing season were used as starting plant material, and two groups of clones were generated: permanent and transplanted clones.
The cultivation conditions in 2015, 2016 and 2017 were modified by incorporating reciprocal transplants between EGs (Fig. 1). In each season, clones were divided into two groups. Three clones per genotype were cultivated repeatedly in the same EG as in the previous season (permanent clones). For example, clones of G-1, G-2 and G-3 cultivated in EG-1100 in 2014 were cultivated again in EG-1100 in 2015. The other three clones were transplanted (transplanted clones) into the other EG (reciprocal transplants). For example, clones of G-1, G-2 and G-3 cultivated in EG-1100 in 2014 were transplanted into EG-2100 in 2015, whereas those grown in EG-2100 in 2014 were transplanted into EG-1100 in 2015 (Fig. 1). In summary, in 2015, 2016 and 2017, three biological replicates of each of the three genotypes and condition (permanent or transplanted) were cultivated in each EG, that is, three permanently grown clones and three transplanted clones of each of the three genotypes were cultivated in each EG, totalling 18 plants per EG.
Environmental characterisation
Two data loggers (iButton Hygrochron, Maxim Integrated, USA) were placed in each EG at 30 cm from ground to register atmospheric temperature (T°) and relative humidity (RH). One additional data logger (iButton Thermochron, Maxim Integrated, USA) was placed in a pot in each EG to register soil temperature. Data were registered every hour from the beginning of the 2016 season until the end of the 2017 season. Also, to characterise each season (from 2015 to 2017), T°, RH and precipitation data from the nearest weather station at “El Plumerillo”, located ~40 km away from the VNR, were used (data provided by the Meteorological Information Centre of Argentina).
Phenotypic characterisation
All variables described below were measured for all the plants cultivated in 2015, 2016 and 2017 at both EGs.
Leaves
The 6th and 7th fully expanded leaves were removed from plants in anthesis, wrapped with aluminium foil and conserved on ice. Once in the laboratory, leaves were photographed and the total leaf area, terminal leaflet area and rachis length were measured with AxioVison software (Carl Zeiss MicroImaging GmbH, Jena, Deutschland); leaflet number was also registered. Leaflet thickness was estimated through the area/weight ratio in two 2 cm2 leaflet discs dried at 40 °C until reaching a constant weight. Photoprotective pigments as ultraviolet absorbing compounds (UVAC) and anthocyanins were determined in the leaflet discs according to Ibañez et al. (2017).
Tubers
At harvest, the total tuber number (TN) and total and mean tuber weight were recorded for each plant, except in the 2017 season in which data from EG-2100 had to be excluded due to the harmful action of stray animals on both substrate and tubers. Harvested tubers were stored at 4 °C and periodically examined for sprouting (dormancy breakdown). Sprouted tubers were maintained at room temperature in darkness, and the sprouting percentage (S%) and sprout length (SL) were recorded every 2 weeks for 60 days.
DNA extraction
Leaves from each genotype were collected when anthesis began. Leaf collection was done at the same time from plants growing in both EGs and repeated in each of the three growing seasons. Sample preparation and DNA extraction and quantification were done according to Marfil et al. (2015).
MSAP analysis in EG approach
MSAP markers were used to estimate epigenetic variability according to Cara et al. (2013). Three selective primer combinations without fluorescence were assayed: EcoRI + AGC/HpaII-MspI + ATC, EcoRI + AGA/HpaII-MspI + ATC and EcoRI + ACA/HpaII-MspI + ATC. The amplification products were separated by electrophoresis on a 6% denaturing polyacrylamide gel, silver stained and scanned for manual scoring. To calculate the error rate, a subset of 12 samples were analysed in duplicate for each season, using the same extracted DNA.
From the EcoRI/HpaII and EcoRI/MspI raw data, each locus could present multistate information. To construct binary epigenetic matrices, each locus in the multistate matrix was split into three subloci (or subepiloci) following the codification ‘Mixed Scoring 2’ described by Schulz et al. (2013): fragments present in both the HpaII and MspI lanes were codified as Type I, fragments present in the HpaII lane but absent in MspI lane were codified as Type II and fragments present in the MspI lane but absent in HpaII lane were codified as Type III. Then, two types of methylation states were considered: non-methylated (Type I) and methylated (Types II + III). In the methylated state, both amplification patterns (II + III) were clustered in one category according to the findings of Fulneček and Kovařík (2014), who indicated that the hemi-methylated state (fragment presence in the HpaII lane) could not be determined with accuracy. The absence of fragments on both lanes was not used for calculating the methylation state because it could be the result of genetic changes.
Data analysis
Environmental data
In the 2016 and 2017 seasons, each EG was characterised according to the monthly mean values of RH, atmospheric T° and soil T°. Then, a comparison between EGs was carried out by ANOVA and Fisher’s LSD with InfoStat (version 2017; Grupo InfoStat, Córdoba, Argentina). Moreover, in order to understand the seasonal variability between EGs, the monthly environmental data were compared between EGs. To this end, the difference in EG-2100 respect to EG-1100 was calculated for each environmental parameter, with negative or positive values indicating, respectively, lower or higher values in EG-2100. The comparison among seasons was made by ANOVA and Fisher’s LSD with InfoStat. To explore the relationships among environmental factors, EGs and seasons, a principal component analysis (PCA) was performed with variance-scaled and centred data with ‘FactoMineR’ and ‘factoextra’ packages in R (Le et al. 2008; Kassambara and Mundt 2017; R Core Team 2018). Two PCA were constructed: one with data from the data loggers placed in each EG, and the other from data provided by the nearest weather station.
Phenotypic data
S% and SL were analysed with a generalised linear mixed model in InfoStat. Days after harvest and growing season were considered random factors, whereas EG and transplanting were considered fixed factors, with logit used as link function for binomial distribution for S% and identity for SL in InfoStat. TN was analysed by using a generalised model in InfoStat, with EG and transplanting as fixed effects and log as link function for the Poisson distribution. The effects of EG and transplanting in the total leaf area, terminal leaflet area, rachis length, leaflet thickness, UVAC and anthocyanin compounds and total and mean tuber weight were determined by ANOVA and Fisher’s LSD comparison in InfoStat. To determine associations between plants from each EG and phenotypic traits, a PCA was performed with centred and variance-scaled data using ‘FactoMineR’ and ‘factoextra’ packages in R. Phenotypic plasticity was quantified for exploring genotype responses to the environment, and the relative distances plasticity index (RDPI) was calculated according to Valladares et al. (2006). In brief, the distance among each trait value for all pair of genotypes (grown in the two different micro-environments) and defined as the absolute value, was calculated as the difference between the values measured in each EG, and the relative distance was then obtained by dividing this difference by the sum of both values. The RDPI could take values ranging from 0 to 1, corresponding to absence of plasticity to maximal plasticity, respectively. To test the consistency of each evaluated trait over time, Spearman rank correlation coefficients were calculated (Garnier et al. 2001) with the package ‘stats’ in R, which compares whether the ranking for each trait remained similar both temporally (between growing season) and spatially (between EG). The rank was independently established for each evaluated genotype and by considering all replicates cultivated in each EG.
MSAP
Three different binary epigenetic matrices were constructed to analyse DNA cytosine methylation: (i) a combined matrix, with the full set of data (three types of subepiloci) coding the presence (1)/absence (0) of fragment types I, II and III, (ii) non-methylated matrix, in which the non-methylated loci were coded as fragments type I = non-methylated state = 1/fragments type II and III = methylated state = 0 and (iii) methylated matrix, in which the methylated loci were coded as fragments type II and III = methylated state = 1/fragments type I = non-methylated state = 0. The combined matrix contains the full set of data, while the non-methylated and methylated matrices were constructed to test the potential functionality of these specific methylation states (Schulz et al. 2013; Preite et al. 2015). In the three codified matrices, ambiguous type-IV loci were coded as zeros. Monomorphic loci and singletons (i.e., fragments present only in one sample) were excluded, thus, all the analyses were performed with polymorphic markers. From these binary matrices, the epigenetic distance matrices were calculated by using Sorensen–Dice coefficient (Sneath and Sokal 1973). In addition, and in order to test the consistency of the performed analysis, the Jaccard coefficient was used to calculate the epigenetic distance from the combined matrix (Supplementary information, Table S1 and Fig. S1). To determine relationships among plants from the EG based on MSAP polymorphism, a principal coordinates analysis was performed with the ‘vegan’ package (Oksanen 2015) in R. An analysis of variance through distance matrices was made with the adonis function in the vegan package to assess the effects of EG, genotype and transplanting for each evaluated season. This function studies differences in the mean epigenetic distance among plants within each source of variability (EG, genotype and transplanting) and allows to examine the significance of those partitions (Oksanen 2015). DNA methylation levels of plants cultivated in the EGs were computed as the observed proportion of methylated and non-methylated loci in the multistate matrix. Differences in the proportion of each methylation state (methylated or non-methylated) were evaluated by a generalised linear model, with log as link function for binomial distribution with InfoStat Software.
Correlations
A simple Mantel test was performed in order to detect linear correlations between distance matrices from epigenetic (combined matrix) and phenotypic variability in the 2015, 2016 and 2017 seasons by using InfoStat. All phenotypic traits were considered in the analysis of the first two seasons, but only leaf traits were considered for the analysis of the three seasons because the tuber phenotypic data of the 2017 season were incomplete.
Results
Environmental differences in EGs among seasons
The RH, soil and atmospheric T° were registered in 2016 and 2017 with data loggers in each EG (Table S2). The 2016 season was colder and wetter than the 2017 one. In fact, on average and when comparing the 2017 vs. the 2016 seasons, the former had lower RH (66.2% vs. 74.1%) and higher atmospheric (13.1 °C vs. 12.0 °C) and soil T° (17.8 vs. 13.9 °C) than the latter (P value < 0.0001).
In both years, the weather in EG-2100 was colder and wetter than in EG-1100 (Table S2). On average, in EG-2100 the mean annual T° was 36.3% and 33% lower, and RH 12.8% and 17% higher than in EG-1100 for 2016 and 2017, respectively. The differences in mean soil T° between EGs were similar to the observed for the atmospheric T° (Table S2).
Monthly differences between EG in the environmental parameters were also compared between the 2016 and 2017 seasons (Table S3). In February (summer in the southern hemisphere), when samples were collected for DNA extraction, differences between the EG-2100 respect to the EG-1100 were lower in 2016 than in 2017 for RH (9.04% vs. 14.14%) and for both atmospheric T° (−6.1 °C vs. −7.1 °C) and soil T° (−4.1 °C vs. −6.3 °C).
The environmental data registered in ‘El Plumerillo’ weather station indicated that the 2015 season was drier and warm (54.9% RH and 17.3 °C) than the 2016 (62.0% RH and 16 °C) and 2017 (55.6% RH and 16.8 °C) seasons. Also, in the 2016 season, the highest accumulated precipitation (338.4 mm) was register in comparison with that of the 2015 and 2017 seasons (116.2 and 180.8 mm, respectively).
To summarise, a PCA was performed with the environmental information (Fig. 2). In 2016, EG-1100 was associated with the highest values of atmospheric T° and the minimum RH, and EG-2100 was associated with the highest mean RH (Fig. 2a). In 2017, EG-1100 was associated with the highest soil temperature and EG-2100 with the maximum RH (Fig. 2a). According to the information of the closest weather station (Fig. 2b), the 2016 season was associated with highest values of RH, 2017 with the highest values of atmospheric T°, and 2015 with intermediate values of both RH and T°, and the lowest value of accumulated precipitation.
Biplot representation of principal component analysis for the environmental variables registered in: a two mountain experimental gardens (EG-1100, turquoise dots and EG-2100, pink dots) with an elevation difference of about 1000 m within an Andean Protected Area during two seasons (2016 and 2017); and in b the Protected Area nearest weather station ‘El Plumerillo’ for 2015, 2016 and 2017 seasons. Analysis based on mean (Mean_RH), minimum (Min_RH) and maximum (Max_RH) relative humidity, mean (Mean_T), minimum (Min_T) and maximum (Max_T) atmospheric temperature, mean (S_Mean_T), minimum (S_Min_T) and maximum (S_Max_T) soil temperature and accumulated precipitation (Acc_PP) variables.
Phenotypic variability
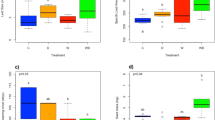
The PCA summarises all phenotypic traits measured for each genotype and EG for the three evaluated seasons (Fig. 3). Leaf traits (Fig. 3a) showed differences among seasons and EGs. The 2015 season was associated with the highest values of leaflet number and UVAC compounds for both EGs, with a small difference between them. In 2016, both EGs were associated with the highest values of total and terminal leaflet area, rachis length and anthocyanin compounds. In 2017, both EGs were associated with the highest values of leaflet thickness. For the tuber traits (Fig. 3b) measured in 2016, EG-2100 was associated with the highest values of mean and total tuber weight and TN. In 2017, EG-1100 was associated with the highest values of S% and SL.
Principal component representation based on phenotypic variability in three S. kurtzianum genotypes cultivated during three successive seasons in two mountain experimental gardens (EG) with an elevation difference of about 1000 m within an Andean Protected Area. a Biplot representation of principal component analysis for leaf variables: leaflet thickness, 6th leaf area (LA6l), 7th leaf area (LA7l), terminal leaflet area (TLA), 6th leaf rachis length (RL6l), 7th leaf rachis length (RL7l), 6th lateral leaflet numbers (LlN6l), 7th lateral leaflet numbers (LlN7l), anthocyanins and UVACs. b Biplot representation of principal component analysis for tuber traits: sprout length (SL), sprouting percentage (S%), mean tuber weight (MTW), total tuber weight (TTW) and tuber numbers (TN). The phenotypes were characterised in plants cultivated in EG-1100 (turquoise diamonds) and EG-2100 (pink diamonds) in 2015, 2016 and 2017 seasons. In 2017, tuber variables were analysed only in EG-1100 because the EG-2100 data were incomplete. Small symbols are individual plants and large symbols indicate EG mean.
A differential response was observed for the same genotypes according to the EG in which they were grown. For example, in the 2015 season, G-1 presented thicker leaves, G-2 presented more tubers and G-3 presented shorter rachis in EG-2100 than in EG-1100 (Table S4). In 2016, G-1 and G-2 presented larger sixth terminal leaflet area, and G-1 and G-3 more anthocyanin compounds in EG-2100 than in EG-1100 (Table S5). In 2017, only G-2 and G-3 presented differences between EGs (Table S6).
Plasticity index values, measured as RDPI, ranged from 0.05 to 0.49, with the only statistical differences among seasons being for SL and S% whereas genotypes also presented different RDPI, ranging from 0.05 to 0.46, with statistical differences detected only for S% and sprout tuber length (Table 1). Leaf traits were less plastic than tuber traits. On average, RDPI for leaf traits was 0.19, 0.16 and 0.21 for the 2015, 2016 and 2017 seasons, respectively. On the other hand, RDPI for tuber traits was 0.19, and 0.33 for the 2015 and 2016 seasons, respectively. The same trend was observed when RDPI was compared among genotypes. RDPI in leaf traits was 0.18, 0.19 and 0.19 for G-1, G-2 and G-3, respectively, whereas RDPI for tuber traits was 0.32, 0.24 and 0.23 for the same genotypes, respectively.
The most frequent result was the lack of consistency of the evaluated traits between seasons, although some of them showed consistency for at least two seasons (e.g., 6th leaf rachis length, leaflet thickness, mean tuber weight and S% for G-1; Table S7).
Epigenetic variability
By using three primer combinations, a total of 182 loci were evaluated, with 0.5% of error rate calculated from 12 duplicated samples. According to the alternative methylation states at each locus (multistate matrix), a total of 306 binary epialleles or subepiloci (i.e., alternative methylation states: non-methylated condition I, and methylated condition II + III) were computed. The number of polymorphic markers varied for each genotype and season (Table S8). In the multivariate analysis using the combined matrix to calculate epigenetic distances, genotypes explained the greatest proportion of the variability in the MSAP profiles (except for 2017). In addition, significant effects of the EGs were detected in two out of three seasons analysed (Table 2). Due to the largest influence of genotypes in the observed epigenetic variability, the three evaluated genotypes were independently analysed, and significant effects of the EGs were also detected in two out of three seasons analysed (Table 3). The three genotypes presented differential methylation patterns (i.e., the combination of multiple methylated and non-methylated states at different loci) according to the EG where they were cultivated in the 2015 and 2017 seasons (Fig. 4). In 2016, however, no epigenetic differentiation between EGs was observed (Fig. 4). These results were further confirmed with the multivariate analysis of the distance matrix (Table 3). The analysis of reciprocal transplants between EGs showed no methylation differences between transplanted and permanent clones for the three genotypes and the three analysed seasons (Table 3). When these analyses were performed separately for the methylated and non-methylated loci, the clustering patterns were similar to those obtained previously (Fig. S2 and Table S9). The significant effects of the EGs on the DNA methylation variability were maintained for the 2015 and 2017 seasons, except for the methylated loci of genotypes 2 and 3 in 2015 (Table S9).
Principal coordinate representation based on a combined matrix of methylated and non-methylated loci derived from methylation-sensitive amplified polymorphism (MSAP) markers evaluated in clones from genotypes 1, 2 and 3 cultivated in two mountain experimental gardens (EG) with an elevation difference of about 1000 m: EG-1100 (turquoise) and EG-2100 (pink) for the 2015, 2016 and 2017 seasons. Each group presented a 0.90 confidence interval.
When the DNA methylation levels were analysed, the non-methylated state was the most common in the three assayed genotypes and during the three seasons (Table S10). Moreover, no differences in DNA methylation levels induced by the EG were observed (Table S11).
Epigenetic and phenotypic variability correlations
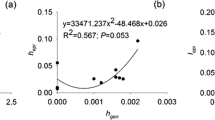
The simple Mantel test indicated that the epigenetic variability was associated with the phenotypic variability for all the considered traits in the 2015 and 2016 seasons (R = 0.36, P value = 0.0001; Fig. S3a). Also, for the 2015, 2016 and 2017 seasons, the epigenetic variability was associated with the leaf phenotypic variability (R = 0.37, P value = 0.0020; Fig. S3b).
Discussion
The understanding of the molecular basis of environmentally induced changes in the phenotype is of great interest in breeding programs. In addition, the study of these changes in wild species could improve our understanding and management of the consequences of climate change. Our work is an initial cross-disciplinary study between ecology and molecular biology, which aims to understand the effects of a real-world context on morphological, biochemical and phenological phenotypes and epigenetics. Epigenetic modifications can contribute to phenotypic plasticity with particular implications in fluctuating environments (Kooke et al. 2015). Species with asexual reproduction (mitotic inheritance) have been proposed as ideal experimental models to study the interplay of phenotypic plasticity, epigenetic variability and adaptation (Castonguay and Angers 2012; González et al. 2018). However, the temporal and spatial phenotypic plasticity of a given genotype and the epigenetic dynamics during successive cycles of agamic reproduction have been overlooked. The present study was set out in this conceptual framework. First, we worked with a species with, presumably but unmeasured, high phenotypic variability, with agamy as an alternative mode of reproduction (Marfil et al. 2015). Second, we performed the assay in a mountainous region, a scenario that offers pronounced environmental gradients. This in situ experiment enabled us to describe the spatial and temporal dynamics of phenotypic variability and DNA methylation patterns during three cycles of clonal reproduction.
In order to obtain evidence about the environmentally induced epigenetic and phenotypic variation in a context of ecological realism, two EGs with differences of ~1000 m in altitude were used. Depending on the season, these two EGs presented differences between 33% and 36.6% in atmospheric mean T° and between 12.8% and 17% in atmospheric mean RH. With this experiment, we showed that clones from different genotypes underwent significant DNA methylation changes and, also, the expression of significant phenotypic plasticity in responses to the two different natural micro-environments. Also, we found that the methylation patterns were greatly influenced by the genotype, indicating that the environmentally induced methylation changes are genotype-dependent, in agreement with previous results (Dubin et al. 2015).
The phenotypic evaluation showed the key influence of the growing seasons. The 2015 season was characterised by highest levels of UVACs. These photoprotective pigments are a common plant response to increased levels of ultraviolet-B radiation (UV-B; Berli et al. 2013). However, Ibañez et al. (2017) observed that the high levels of UVACs in S. kurtzianum could not only be explained by high UV-B levels. In addition to UV-B, UVACs accumulate in response to cold and are thought to be important for the expression of tolerance against this stress (Schulz et al. 2015). In February 2015, the mean temperature was 10% lower than in the 2016 and 2017 seasons (23.7, 26.3 and 26.4 °C, respectively; P value = 0.0014), likely determining the higher UVAC levels in 2015.
Leaf area of higher plants has been widely studied in relation to diverse environmental factors (Peppe et al. 2011). Wright et al. (2017) found that the combination of climatic factors better explained differences in leaf area than individual climatic factors. These authors observed that in the driest sites, leaf area decreased with temperature during the warmest months. Also, Peppe et al. (2011) observed that leaf size varied in a sensitive way according to water availability more than with temperature differences. In the present work, the wetter season (2016, accumulated precipitation ~350 mm) and the wetter EG (EG-2100) were associated with bigger leaf size, whereas the 2017 season, which was the warmest and driest season, was associated with thicker leaves, a trait that has been correlated with dry ecosystems (Coneva and Chitwood 2018). In addition, the unique and unpredictable effects of the growing season on the phenotype were supported by the lack of consistency in the evaluated traits throughout seasons for each EG.
The differential epigenetic patterns between clones cultivated in the two EGs were observed in two out of the three evaluated seasons. There were several environmental differences among seasons indicating that each year had a distinct effect on the DNA methylation variability. The high precipitation regime in 2016 could likely have determined the lack of epigenetic differentiation between EGs for the three evaluated genotypes. The 2016 season registered the highest precipitation in the previous 4 years (412-mm annual accumulated precipitation). In addition, we found that the differences in RH between EGs were lower in the 2016 than in the 2017 sampling collection month (in February, the differences in RH between EGs were 36% lower in 2016 than in 2017). In the desert ecosystem in which the experiment was done, it is likely that water availability could be the principal factor inducing differential environmental conditions, which in turn, would determine differential epigenetic patterns in the assayed genotypes.
As asexual reproduction circumvents meiosis, a reduced epigenetic resetting would be expected in each cycle (Verhoeven and Preite 2014). However, we found an extensive mitotic reprogramming of methylation patterns principally associated with the growing season. A key question that emerges from this result is whether the documented epigenetic changes were environmentally directed or stochastically induced (see Verhoeven and Preite 2014). If the stochastic changes would have been predominant, a high dispersion among replicates and genotypes cultivated in each season would have been expected. The lack of discrimination between transplanted vs. permanent clones indicates that a programmed epigenetic response was activated in each year. In addition, the close epigenetic similarity between transplanted and permanent clones cultivated in the same EG indicates that the previously established methylation patterns were mostly reprogrammed according to the current environmental conditions. It is important to mention that the methylation reprogramming was induced in a short-time scale. Gao et al. (2010) working with Alternanthera philoxeroidesals also found a rapid restructuring in DNA methylation states and reversible phenotypic responses to particular environmental factors. This lack of stress-memory could be an adaptive response as the stressful conditions are mainly transient and unpredictable (Crisp et al. 2016). The results obtained in the present work suggest that the epigenetic mitotic reprogramming contributes in the potato asexual propagation mode. The rapid response to environmental changes through phenotypic plasticity and methylation reprogramming may possibly allow persistence of potato populations over time (Erazzú et al. 2009; Marfil et al. 2015). This could explain why potatoes were considered very difficult to eradicate in most regions (Hawkes 1954).
We found significant correlations between environmentally induced DNA methylation changes and phenotypic variation. This result indicates that methylation changes in wild potatoes play a role in phenotypic plasticity induced by specific environmental differences. Even though phenotypic plasticity in potatoes is a quality that has been recognised for decades by multiple authors, systematic studies have not addressed this characteristic (see Hawkes and Hjerting 1969). By comparing the RDPI, a significant difference between seasons and among genotypes was associated with tuber traits. In addition, higher values of phenotypic plasticity were found in sprouting behaviour, which is a crucial life-history stage influencing the survival and maintenance of wild S. kurtzianum populations (Marfil et al. 2015). Thus, this high tuber plasticity could lead to the successful establishment and persistence of natural populations under fluctuating environmental conditions. Notwithstanding, the evaluated genotypes also presented differences in tuber plasticity that indicated a possible local selection pressure on this trait. Using RDPI, phenotypic plasticity has been characterised in traits of different species (e.g., Godoy et al. 2012; Wei et al. 2019, Hiatt and Flory 2020). The significance of phenotypic plasticity to different environmental conditions, have often yielded species‐specific conclusions. However, under certain conditions, higher trait plasticity could contribute to the successful spreading of invasive plants (Hiatt and Flory 2020). In our study, because the clustering differentiation was comparable regardless the epigenetic distance matrix used (combined, methylated or non-methylated), the environmentally induced methylation patterns associated to phenotypic variation could not be attributed to a specific methylation state. Thus, it is not advisable to make inferences of functionality based on the different observed methylation states.
Medrano et al. (2014), who worked with the perennial herb Helleborus foetidus, proposed that the epigenetic variation possibly confers this species with the capacity to exploit a broad range of ecological conditions. Similarly, in wild-potato populations, it has been reported that new epigenetic patterns, established after interspecific hybridisation, would enable the hybrids to cope with new environments (Cara et al. 2013). Our results suggest that the epigenetic response to environment experiencing short-term fluctuations allows genotype persistence and promote the maintenance of the genetic diversity of populations. The evolution conducted by selective pressures tends to reduce the within population genetic diversity. However, one of the fundamental characteristics of wild-potato populations is their great genetic variability within populations (Jansky et al. 2006, 2009; Spooner et al. 2009; Marfil and Masuelli 2014; Marfil et al. 2015). The environmentally induced phenotypic and epigenetic variation could be contributing a buffering effect against selective processes in nature (O’Dea et al. 2016; Donelson et al. 2017). Also, our study contributes to understand how environmental exposure induces reversible epigenetic modifications that alter plant phenotypes, which not necessarily lead to genomic fixation of environmentally induced traits in later generations. However, since the most common methylation state found in this work was non-methylated, it is possible that transposable elements are activated, inducing genotypic changes that are inherited through the germ line with an increment of genetic variability (see Pimpinelli and Piacentini 2019).
In summary, we demonstrated that in an herbaceous perennial species with asexual reproduction and in an Andean desert with stochastic environmental changes principally associated with precipitation levels, the phenotype and methylation patterns are rewritten every year. The results obtained over three seasons did not show any evident memory effect of the previous environment, but a reprogramming of methylation patterns and high phenotypic variation according to the growing conditions. These reprogramming in methylation patterns and phenotype, in turn, could contribute to genotype persistence, preventing the loss of genetic variability in potato natural populations.
Data availability
The raw data of MSAP markers and phenotypic analyses obtained in this study can be found under https://doi.org/10.5061/dryad.ksn02v729 in the Dryad Data Repository.
References
Berend K, Haynes K, MacKenzie CM (2019) Common garden experiments as a dynamic tool for ecological studies of alpine plants and communities in northeastern north America. Rhodora 121:174–212
Berli FJ, Alonso R, Bressan-Smith R, Bottini R (2013) UV-B impairs growth and gas exchange in grapevines grown in high altitude. Physiol Plant 149:127–140
Camadro EL (2012) Relevance of the genetic structure of natural populations, and sampling and classification approaches for conservation and use of wild crop relatives: potato as an example. Botany 90(11):1065–1072
Cara N, Marfil CF, Masuelli RW (2013) Epigenetic patterns newly established after interspecific hybridization in natural populations of Solanum. Ecol Evol 3:3764–3779
Castonguay E, Angers B (2012) The key role of epigenetics in the persistence of asexual lineages. Genet Res Int 2012:1–9
Coneva V, Chitwood DH (2018) Genetic and developmental basis for increased leaf thickness in the Arabidopsis cvi ecotype. Front Plant Sci 9:1–10.
Cooper HF, Grady KC, Cowan JA, Best RJ, Allan GJ, Whitham TG (2019) Genotypic variation in phenological plasticity: Reciprocal common gardens reveal adaptive responses to warmer springs but not to fall frost. Glob Change Biol 25:187–200
Correll D (1962) The potato and its wild relatives. Texas Research Foundation, Renner, TX
Cortijo S, Wardenaar R, Colomé-Tatché M, Gilly A, Etcheverry M, Labadie K et al. (2014) Mapping the epigenetic basis of complex traits. Science 343:1145–1148
Crisp PA, Ganguly D, Eichten SR, Borevitz JO, Pogson BJ (2016) Reconsidering plant memory: intersections between stress recovery, RNA turnover, and epigenetics. Sci Adv 2:1–14.
Cubas P, Vincent C, Coen E (1999) An epigenetic mutation responsible for natural variation in floral symmetry. Nature 401:157–161
Donelson JM, Salinas S, Munday PL, Shama LN (2017) Transgenerational plasticity and climate change experiments: Where do we go from here? Glob Change Biol 24:13–34
Dubin MJ, Zhang P, Meng D, Remigereau M-S, Osborne EJ, Casale FP et al. (2015) DNA methylation in Arabidopsis has a genetic basis and shows evidence of local adaptation. Elife 4:e05255
Erazzú LE, Camadro EL, Clausen AM (2009) Persistence over time, overlapping distribution and molecular indications of interspecific hybridization in wild potato populations of Northwest Argentina. Euphytica 168:249–262
FAO UN (2018) FAOstat. http://www.fao.org/faostat/en/#data/QC. Accessed 29 Dec 2018.
Fulneček J, Kovařík A (2014) How to interpret methylation sensitive amplified polymorphism (MSAP) profiles? BMC Genet 15:1–9
Gao L, Geng Y, Li B, Chen J, Yang J (2010) Genome-wide DNA methylation alterations of Alternanthera philoxeroides in natural and manipulated habitats: Implications for epigenetic regulation of rapid responses to environmental fluctuation and phenotypic variation. Plant Cell Environ 33:1820–1827
Garnier E, Laurent G, Bellmann A, Debain S, Berthelier P, Ducout B, Roumet C, Navas ML (2001) Consistency of species ranking based on functional leaf traits. N Phytologist 152:69–83
Godoy O, Valladares F, Castro‐Díez P (2012) The relative importance for plant invasiveness of trait means, and their plasticity and integration in a multivariate framework. N Phytologist 195:912–922
González APR, Preite V, Verhoeven KJF, Latzel V (2018) Transgenerational effects and epigenetic memory in the clonal plant Trifolium repens. Front Plant Sci 9:1–11
Hawkes JG (1954) The ecology of wild potato species and its bearing on the origin of potato cultivation. J d’Agriculture Traditionnelle et de Botanique Appliquée 1:356–358
Hawkes JG, Hjerting JP (1969) The potatoes of Argentina, Brazil, Paraguay and Uruguay. A biosystematic study. Clarendon Press, Oxford, UK
Herrera CM, Bazaga P (2011) Untangling individual variation in natural populations: ecological, genetic and epigenetic correlates of long-term inequality in herbivory. Mol Ecol 20:1675–1688
Hijmans RJ, Spooner DM, Salas AR, Guarino L, de la Cruz J (2002) Atlas of wild potatoes. International Plant Genetic Resources Institute, Rome
Hiatt D, Flory SL (2020) Populations of a widespread invader and co‐occurring native species vary in phenotypic plasticity. N Phytologist 225:584–594
Ibañez VN, Berli FJ, Masuelli RW, Bottini RA, Marfil CF (2017) Influence of altitude and enhanced ultraviolet-B radiation on tuber production, seed viability, leaf pigments and morphology in the wild potato species Solanum kurtzianum Bitter and Wittm collected from an elevational gradient. Plant Sci 261:60–68
IPCC (2014) Climate Change 2014: Synthesis report. Contribution of Working Groups I, II and III to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change, In: Core Writing Team, RK Pachauri, LA Meyer (eds). IPCC, Geneva, Switzerland, p 151
Jansky SH, Simon R, Spooner DM (2006) A test of taxonomic predictivity: resistance to white mold in wild relatives of cultivated potato. Crop Sci 46:2561–2570
Jansky SH, Simon R, Spooner DM (2009) A test of taxonomic predictivity: resistance to the colorado potato beetle in wild relatives of cultivated potato. J Economic Entomol 102:422–431
Kassambara A, Mundt F (2017) Factoextra: extract and visualize the results of multivariate data analyses. R package version 1.0.5.
Kelly M (2019) Adaptation to climate change through genetic accommodation and assimilation of plastic phenotypes. Philos Trans R Soc B 374:20180176
Kooke R, Johannes F, Wardenaar R, Becker F, Etcheverry M, Colot, Vreugdenhil D, Keurentjes JJB (2015) Epigenetic basis of morphological variation and phenotypic plasticity in Arabidopsis thaliana. Plant Cell 27:337–348
Latzel V, Allan E, Bortolini Silveira A, Colot V, Fischer M, Bossdorf O (2013) Epigenetic diversity increases the productivity and stability of plant populations. Nat Commun 4:1–7
Le S, Julie J, Husson F (2008) FactoMineR: an R package for multivariate analysis. J Stat Softw 25:1–18
Lira-Madeiros CF, Parisod C, Fernandes RA, Mata CS, Cardoso MA, Ferreira PCG (2010) Epigenetic variation in mangrove plants occurring in contrasting natural environment. PLoS ONE 5:1–8
Marfil CF, Camadro EL, Masuelli RW (2009) Phenotypic instability and epigenetic variability in a diploid potato of hybrid origin, Solanum ruiz-lealii. BMC Plant Biol 9:1–16.
Marfil CF, Asurmendi S, Masuelli RW (2012) Changes in micro RNA expression in a wild tuber-bearing Solanum species induced by 5-Azacytidine treatment. Plant Cell Rep 31:1449–1461
Marfil CF, Masuelli RW (2014) Reproductive ecology and genetic variability in natural populations of the wild potato, Solanum kurtzianum. Plant Biol 16:485–494
Marfil CF, Hidalgo V, Masuelli RW (2015) In situ conservation of wild potato germplasm in Argentina: example and possibilities. Glob Ecol Conserv 3:461–476
Marfil CF, Duarte P, Masuelli RW (2018) Phenotypic and epigenetic variation induced in newly synthesized allopolyploids and autopolyploids of potato. Scientia Horticulturae 234:101–109
Masuelli RW, Camadro EL, Erazzú LE, Bedogni MC, Marfil CF (2009) Homoploid hybridization in the origin and evolution of wild diploid potato species. Plant Syst Evol 277:143–151
McGregor CE, van Treuren R, Hoekstra R, van Hintum TJL (2002) Analysis of the wild potato germplasm of the series Acaulia with AFLPs: implications for ex situ conservation. TAG. Theor Appl Genet 104:146–156
Medrano M, Herrera CM, Bazaga P (2014) Epigenetic variation predicts regional and local intraspecific functional diversity in a perennial herb. Mol Ecol 23:4926–4938
Nicotra AB, Atkin OK, Bonser SP, Davidson AM, Finnegan EJ, Mathesius U, Poot P, Purugganan MD, Richards CL, Valladares F, van Kleunen M (2010) Plant phenotypic plasticity in a changing climate. Trends plant Sci 15:684–692
Nicotra AB, Segal DL, Hoyle GL, Schrey AW, Verhoeven KJF, Richards CL (2015) Adaptive plasticity and epigenetic variation in response to warming in an Alpine plant. Ecol Evolution 5:634–647
O’Dea RE, Noble DW, Johnson SL, Hesselson D, Nakagawa S (2016) The role of non-genetic inheritance in evolutionary rescue: epigenetic buffering, heritable bet hedging and epigenetic traps. Environ Epigenetics 2:1–12.
Oksanen J,(2015) Multivariate analysis of ecological communities in R: vegan tutorial. R Documentation. 1:11–12
Peppe DJ, Royer DL, Cariglino B, Oliver SY, Newman S, Leight E et al. (2011) Sensitivity of leaf size and shape to climate: global patterns and paleoclimatic applications. N Phytologist 190:724–739
Pimpinelli S, Piacentini L (2019) Environmental change and the evolution of genomes: Transposable elements as translators of phenotypic plasticity into genotypic variability. Funct Ecol 00:1–14
Preite V, Snoek LB, Oplaat C, Biere A, Van Der Putten WH, Verhoeven KJF (2015) The epigenetic footprint of poleward range-expanding plants in apomictic dandelions. Mol Ecol 24:4406–4418
R Core Team (2018) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Richards CL, Alonso C, Becker C, Bossdorf O, Bucher E, Colomé-Tatché M et al. (2017) Ecological plant epigenetics: Evidence from model and non-model species, and the way forward. Ecol Lett 20:1576–1590
Richards EJ (2006) Inherited epigenetic variation revisiting soft inheritance. Nat Rev Genet 7:395–401
Schulz B, Eckstein RL, Durka W (2013) Scoring and analysis of methylation‐sensitive amplification polymorphisms for epigenetic population studies. Mol Ecol Resour 13:642–653
Schulz E, Tohge T, Zuther E, Fernie AR, Hincha DK (2015) Natural variation in flavonol and anthocyanin metabolism during cold acclimation in Arabidopsis thaliana accessions. Plant Cell Environ 38:1658–1672
Sneath PHA, Sokal RR (1973) Numerical taxonomy. The principles and practice of numerical classification. WH Freeman and Company, San Francisco, USA
Spooner DM (2009) DNA barcoding will frequently fail in complicated groups: an example in wild potatoes. Am J Bot 96:1177–1189
Spooner DM, Jansky SH, Simon R (2009) Tests of taxonomic and biogeographic predictivity: resistance to disease and insect pests in wild relatives of cultivated potato. Crop Sci 49:1367–1376
Tricker PJ, Gibbings JG, Rodríguez López CM, Hadley P, Wilkinson MJ (2012) Low relative humidity triggers RNA-directed de novo DNA methylation and suppression of genes controlling stomatal development. J Exp Bot 63:3799–3813
Valladares F, Sanchez-Gomez D, Zavala M (2006) Quantitative estimation of phenotypic plasticity: bridging the gap between the evolutionary concept and its ecological applications. J Ecol 94:1103–1116
Verhoeven KJF, Jansen JJ, van Dijk PJ, Biere A (2010) Stress-induced DNA methylation changes and their heritability in asexual dandelions. N Phytologist 185:1108–1118
Verhoeven KJF, Preite V (2014) Epigenetic variation in asexually reproducing organisms. Evolution 68:644–655
Verhoeven KJF, Von Holdt BM, Sork VL (2016) Epigenetics in ecology and evolution: what we know and what we need to know. Mol Ecol 25:1631–1638
Wei N, Cronn R, Liston A, Ashman TL (2019) Functional trait divergence and trait plasticity confer polyploid advantage in heterogeneous environments. N Phytologist 221:2286–2297
Wright IJ, Dong N, Maire V, Prentice IC, Westoby M, Díaz S et al. (2017) Global climatic drivers of leaf size. Science 357:917–921
Zhang YY, Fischer M, Colot V, Bossdorf O (2013) Epigenetic variation creates potential for evolution of plant phenotypic plasticity. N Phytologist 197:314–322
Acknowledgements
IVN is a fellow of the Consejo Nacional de Investigaciones Científica y Técnicas (CONICET), Argentina. This work was supported by Agencia Nacional de Promoción Científica y Tecnológica (PICT-2014-689 and PICT 201-110) and Universidad Nacional de Cuyo (SECTYP A057), Argentina. We are grateful to the Villavicencio Natural Reserve’s Park Ranger Team for their valuable assistance in the field and Elsa Camadro, INTA EEA Balcarce-CONICET and María Virginia Sanchez-Puerta, IBAM UNCuyo-CONICET, for critical reading of the manuscript and English editing.
Author information
Authors and Affiliations
Contributions
MCF and MRW planned and design the research. IVN performed the molecular and morphological analyses. IVN and MCF monitored the experimental gardens approach and drafted the manuscript. MRW contributed in writing the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Alison Bentley
Supplementary information
Rights and permissions
About this article
Cite this article
Ibañez, V.N., Masuelli, R.W. & Marfil, C.F. Environmentally induced phenotypic plasticity and DNA methylation changes in a wild potato growing in two contrasting Andean experimental gardens. Heredity 126, 50–62 (2021). https://doi.org/10.1038/s41437-020-00355-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-020-00355-z
Keywords
This article is cited by
-
Comparative epigenetic and genetic spatial structure in Mediterranean mountain plants: a multispecies study
Heredity (2024)
-
Activating stress memory: eustressors as potential tools for plant breeding
Plant Cell Reports (2022)
-
Non-Thermal Plasmas Affect Plant Growth and DNA Methylation Patterns in Glycine max
Journal of Plant Growth Regulation (2022)
-
In vitro photomorphogenesis, plant growth regulators, melatonin content, and DNA methylation under various wavelengths of light in Phalaenopsis amabilis
Plant Cell, Tissue and Organ Culture (PCTOC) (2022)
-
Vineyard environments influence Malbec grapevine phenotypic traits and DNA methylation patterns in a clone-dependent way
Plant Cell Reports (2021)