Abstract
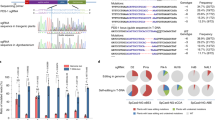
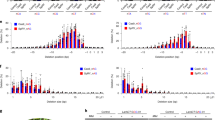
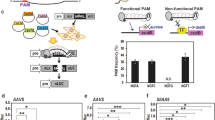
Streptococcus pyogenes Cas9 (SpCas9) is widely used for genome editing and requires NGG as a protospacer adjacent motif (PAM). Here, we show that the engineered SpCas9 (SpCas9-NGv1) can efficiently mutagenize endogenous target sites with NG PAMs in the rice and Arabidopsis genomes. Furthermore, we demonstrate that the SpCas9-NGv1 nickase fused to cytidine deaminase mediates C-to-T substitutions near the 5′ end of the target sequence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The remaining data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Sander, J. D. & Joung, J. K. Nat. Biotechnol. 32, 347–355 (2014).
Doudna, J. A. & Charpentier, E. Science 346, 1258096 (2014).
Jinek, M. et al. Science 337, 816–821 (2012).
Mojica, F. J., Diez-Villasenor, C., Garcia-Martinez, J. & Almendros, C. Microbiology 155, 733–740 (2009).
Shah, S. A., Erdmann, S., Mojica, F. J. & Garrett, R. A. RNA Biol. 10, 891–899 (2013).
Sternberg, S. H., Redding, S., Jinek, M., Greene, E. C. & Doudna, J. A. Nature 507, 62–67 (2014).
Nishimasu, H. et al. Science 361, 1259–1262 (2018).
Hu, J. H. et al. Nature 556, 57–63 (2018).
Kim, D. et al. Nat. Methods 12, 237–243 (2015).
Nishida, K. et al. Science 353, aaf8729 (2016).
Shimatani, Z. et al. Nat. Biotechnol. 35, 441–443 (2017).
Zong, Y. et al. Nat. Biotechnol. 35, 438–440 (2017).
Gaudelli, N. M. et al. Nature 551, 464–471 (2017).
Toki, S. Plant Mol. Biol. Rep. 15, 16–21 (1997).
Wu, F. H. et al. Plant Methods 5, 16 (2009).
Acknowledgements
The plant CRISPR vector (pDeCas9) harbouring the SpCas9-WT gene was kindly provided by F. Fauser, S. Schiml and H. Puchta, Karlsruhe Institute of Technology (Karlsruhe, Germany). The Target-AID vector (nCas9Os-PmCDA1At) was kindly provided by A. Kondo and K. Nishida, Kobe University (Kobe, Japan). We are grateful to S. Gelvin, Purdue University (West Lafayette, IN, USA), for critical reading of the manuscript and helpful suggestions. Digenome sequencing was accomplished by the support of the Advanced Analysis Center, National Agriculture and Food Research Organization (Tsukuba, Japan). This work was supported by the Cabinet Office, Government of Japan, Cross-ministerial Strategic Innovation Promotion Program (SIP), “Technologies for creating next-generation agriculture, forestry and fisheries” (funding agency: Biooriented Technology Research Advancement Institution, NARO).
Author information
Authors and Affiliations
Contributions
S.T. and O.N. conceived the idea for the study. H.N. conducted the in vitro DNA cleavage assay. M.E., M.M., A.E. and H.K. constructed the plasmids. M.E. and M.M. analysed the rice mutants. A.E. and H.K. analysed the transfected Arabidopsis protoplast. A.E., H.K. and T.I. conducted the Digenome sequencing. M.E. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Text, Supplementary Figures 1–9, Supplementary Tables 1–2 and online methods.
Rights and permissions
About this article
Cite this article
Endo, M., Mikami, M., Endo, A. et al. Genome editing in plants by engineered CRISPR–Cas9 recognizing NG PAM. Nature Plants 5, 14–17 (2019). https://doi.org/10.1038/s41477-018-0321-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-018-0321-8
This article is cited by
-
Engineering disease-resistant plants with alternative translation efficiency by switching uORF types through CRISPR
Science China Life Sciences (2024)
-
The applications of CRISPR/Cas-mediated genome editing in genetic hearing loss
Cell & Bioscience (2023)
-
Speed Breeding Opportunities and Challenges for Crop Improvement
Journal of Plant Growth Regulation (2023)
-
Genome Editing Technology and Its Application to Metabolic Engineering in Rice
Rice (2022)
-
SpG and SpRY variants expand the CRISPR toolbox for genome editing in zebrafish
Nature Communications (2022)