Abstract
The root-knot nematode Meloidogyne chitwoodi is a pest that affects potato production in the Pacific Northwest of the United States. Here, to develop new strategies against M. chitwoodi infection of potato, we engineered Bacillus subtilis to secrete the plant-defence elicitor peptide StPep1. Pre-treatment of potato roots with the bacteria secreting StPep1 substantially reduced root galling, indicating that a bacterial secretion of a plant elicitor is an effective strategy for plant protection.
Similar content being viewed by others
Main
Root-knot nematodes (RKN, Meloidogyne spp.) are major agricultural pests on a global scale and can cause billions of dollars in yield losses every year. The temperate root-knot nematode M. chitwoodi is endemic to the potato-growing regions of the Pacific Northwest of the United States, as well as areas of Europe, Asia and Africa1,2,3. This species is important in potato production because it can infect the tubers, causing unsightly galling in addition to brown spots that form under the tuber skin. Commercially available potato cultivars do not have natural genetic resistance to M. chitwoodi. Growers rely on costly nematicides to manage root-knot nematodes in potato fields, and therefore, it would be prudent to develop alternative methods for root-knot nematode control.
One alternative control strategy could be to use synthetic or naturally derived molecules to activate or enhance the plant immune system4. Plants detect tissue damage or injury by wounding or pathogen attack and release endogenous molecules called damage-associated molecular patterns. Plant elicitor peptides (Peps) are one example of damage-associated molecular patterns5,6,7,8,9,10. Pep treatments have been shown to trigger both local and systemic defence responses in plants such as Arabidopsis thaliana and maize, but its effects have not been extensively studied in potato11. A recent paper showed that pre-treatment of soybean seed with soybean Peps (GmPeps) can enhance their resistance to root-knot nematodes, suggesting that defence elicitor peptides could be useful for root-knot nematode control12. Of note, a study showed that B. subtilis can be engineered to secrete a nematode-specific neuropeptide13. Here we combine these two ideas—defence elicitors and bacterial secretion—to develop a B. subtilis delivery system for a plant-defence elicitor used for nematode control, and we highlight its potential as a highly adaptable and novel strategy for plant protection against pathogens.
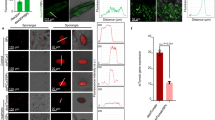
To determine whether the defence elicitor peptide found in potato9, StPep1, is able to protect potato roots from root-knot nematode infection, we analysed M. chitwoodi infections of Russet Burbank potato plants that had been pre-treated with StPep1. For these experiments, potato plants were watered with either 1 µM StPep1 or a mock solution 2 d before nematode inoculation. Nematode juveniles were applied to the roots, and 4 d after M. chitwoodi inoculation, the number of nematodes in the roots were counted. There was no marked difference in the number of juveniles inside the potato roots at 4 dpi between the StPep1- and mock-treated plants, indicating that StPep1 has no marked effect on nematode penetration of the roots (Supplementary Fig. 1). However, the StPep1 pre-treatment considerably reduced M. chitwoodi galling at 12 dpi (Fig. 1a,b) and the number of egg masses at 35 dpi (Fig. 1e,f) produced on potato roots compared with the control. Because of the potential trade-off between plant growth and defence14, we measured the root and shoot biomasses of infected potato plants and found no negative effects on potato aboveground tissue or belowground root weights by the StPep1 treatment (Fig. 1c,d). The StPep1 solution did not directly kill the nematodes (Supplementary Fig. 2), suggesting that the decreased galling was due to enhanced plant resistance to the nematodes. Thus, a single watering with an StPep1 solution stimulated nematode resistance in plants without measurable adverse effects on plant growth.
a–d, Potatoes were watered with 1 µM StPep1 solution, and 2 d later the plants were inoculated with M. chitwoodi juveniles. The number of galls on potato roots was counted at 12 d post-inoculation (dpi) to evaluate the nematode infections. StPep1 pre-treatment significantly reduced the number of galls per plant (a) and galls per g root (b). c,d, The StPep1 pre-treatment had no marked effect on the root or shoot weights of the nematode-inoculated plants. e,f, Egg masses on potato roots were stained with phloxine B and counted at 35 dpi. StPep1 pre-treatment significantly reduced the number of egg masses per plant (e) and egg masses per g root (f). Data are mean ± s.e.m.; n (number of plants) = 10; **P < 0.01, two-tailed Student’s t-test.
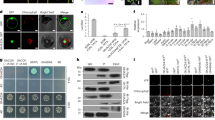
Building on a previous study that showed that Bacillus could be engineered to secrete a plant-parasitic-nematode-specific protein13, we engineered B. subtilis to secrete His-tagged StPep1 (StPep1–His). The fusion protein was detected in B. subtilis protein extracts by western blot (Supplementary Fig. 3a). Secretion of StPep1 was confirmed by enzyme-linked immunosorbent assay, which detected approximately 12.89 ng ml−1 of StPep1 in the culture broth (Supplementary Fig. 3b,c). To determine how long the B. subtilis that secretes StPep1 remains associated with the potato roots during our experiments, we quantified the bacterial colony-forming units associated with potato roots at 1, 4 and 12 d after inoculation. We found that the bacteria remained in association with the roots for at least 12 d after inoculation (Supplementary Fig. 4).
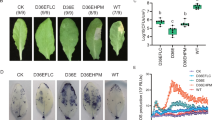
We next assessed the ability of the StPep1-secreting B. subtilis to improve potato resistance against M. chitwoodi. Potato plants were subjected to pre-treatments with water, a B. subtilis culture transformed with the empty vector pBE-S or B. subtilis cultures engineered to express either the C terminus, His-tagged or untagged StPep1. We counted the number of galls per plant 12 d after nematode inoculation. Galling was notably reduced on the potatoes pre-treated with the B. subtilis cultures secreting StPep1–His or untagged StPep1 (Fig. 2a,b). There was no marked difference in galling between pre-treatments with water and B. subtilis transformed with empty vector (Fig. 2a,b), indicating that this strain of B. subtilis does not induce resistance on its own or have nematicidal activity. Infected plant root and shoot biomasses were not substantially affected by any of the B. subtilis treatments (Fig. 2c,d).
a,b, Russet Burbank potato plants were pre-treated with B. subtilis transformed with StPep1–His or StPep1 before M. chitwoodi inoculation. B. subtilis transformed with empty vector pBE-S (EV) and a water-only treatment were used as controls. Plants watered with B. subtilis transformed with StPep1–His or StPep1 showed reduced numbers of galls per plant (a) and number of galls per gram root (b), compared with the controls. c,d, Pre-treatments with engineered B. subtilis had no marked effects on the root or shoot fresh weights of nematode-inoculated plants. Data are mean ± s.e.m. (n = 10). Statistical comparisons were done using a one-way analysis of variance (ANOVA) followed by Tukey’s multiple-comparison test (set at 5%), and different lowercase letters indicate significant differences between treatments.
The results show that pre-treatments with engineered B. subtilis expressing StPep1–His or StPep1 can effectively improve potato plant resistance against M. chitwoodi. To investigate the mode of action of StPep1 in potato, we treated potato roots with either water (mock) or a 100 nM StPep1 solution for gene-expression analyses. We chose a lower concentration of StPep1 solution because for these experiments we soaked tissue-cultured potato roots directly in the StPep1 solution. This is in contrast to previous experiments with StPep1, in which we watered plants grown in sand Cone-tainers with an StPep1 solution before nematode infections and subsequent waterings. Potato gene expression was measured by real-time quantitative PCR with reverse transcription (RT–qPCR) at 0, 2, 6 and 12 h after treatment. Because A. thaliana Pep1 receptor (PEPR) signalling is mediated through salicylic acid6, jasmonic acid15 and ethylene8,10 pathways, we tested the expression of four potato genes linked to these signalling pathways in potato16. We compared the fold change in gene expression between water and StPep1 treatments in roots and leaves. In leaves, the potato genes StPINII, StOsmotin2, StPR1 and StWRKY40 (ref. 16) showed increased expression during the time-course study, suggesting a systemic response to the StPep1 root treatment (Supplementary Fig. 5a). However, none of these genes were differentially regulated in roots (Supplementary Fig. 5b).
To find genes that are differentially regulated by StPep1 in potato roots, we performed a transcriptome analysis on Russet Burbank potato roots incubated in a 100 nM StPep1 solution for 0 and 6 h. Expression profiling revealed that StPep1 treatment led to upregulation of 392 genes and downregulation of 373 genes (at least twofold change at 6 h after inoculation, q < 0.05) (Supplementary Table 1). The results of RNA-seq were also validated by RT–qPCR analysis of a subset of these differentially expressed genes (Supplementary Fig. 6).
We then investigated overrepresentation of gene ontology terms among the differentially expressed genes (Supplementary Fig. 7). We selected eight upregulated genes in two highly enriched gene ontology terms, endopeptidase inhibitor activity and transcription factor activity, to develop StPep1-induced root marker genes. We treated potato plants by soaking roots with either water or 100 nM StPep1. We then measured the expression of these 8 genes by RT–qPCR in both roots and leaves 6 h after treatment. In leaves, the expression of these eight genes varied. However, in the roots, seven of the eight genes were upregulated by StPep1 treatment (Supplementary Fig. 8). Therefore, these seven genes could serve as marker genes in potato roots treated with StPep1.
Next, we aimed to determine whether StPep1 secreted by Bacillus would affect expression of the seven StPep1 marker genes in potato roots. We treated roots with water or the Bacillus cultures (empty vector control or Bacillus secreting StPep1–His) for 6 h. The Bacillus StPep1–His increased expression of five of the seven StPep1 marker genes compared with controls (Supplementary Fig. 9). This indicates that Bacillus secreting StPep1–His can trigger root gene expression similar to StPep1 in solution. When combined with the induced nematode-resistance data, these results suggest that the Bacillus secretes an active defence elicitor.
The jasmonate receptor COI1 and jasmonate signalling are required to some extent for AtPep1 immune responses in A. thaliana15,17. Therefore, we tested whether the seven StPep1-responsive genes were dependent on jasmonate perception in potato roots by using potato plants in which expression of the jasmonate receptor StCOI1 is knocked down using RNA inhibition (referred to StCOI1‐RNAi18) (Supplementary Fig. 10a,b). We measured the fold change in gene expression 6 h after mock treatment and StPep1 treatment in the wild-type and StCOI1‐RNAi roots. We found that gene induction was reduced in the StCOI1‐RNAi plants for 2 of the 7 marker genes tested (Supplementary Fig. 10b). These results indicate that some gene-expression outputs in roots require the jasmonate receptor for StPep1-mediated signalling.
Because some A. thaliana PROPEP genes—which encode Pep precursor proteins—are induced by methyl salicylate treatments11,19 and because AtPep1 activates markers of the salicylic acid pathway17, we tested whether the expression of the StPep1-responsive genes in potato roots required salicylic acid (SA). We obtained potatoes that express the bacterially derived NahG gene (Supplementary Fig. 10c). NahG encodes a salicylate hydroxylase that degrades SA so that the plants cannot accumulate SA. The roots of wild-type potatoes and potatoes expressing the NahG gene (Supplementary Fig. 10c) were treated with water or StPep1 for 6 h, and the expression of the 7 StPep1-responsive genes was measured in the roots. We measured the fold change in gene expression at 6 h in the mock treatment and StPep1 treatment in WT and NahG roots. The results showed that expression of three of the seven marker genes was higher in NahG plants compared with wild-type plants (Supplementary Fig. 10d).
Overall, gene-expression analysis showed that treatment of potato roots with StPep1 caused local and systemic changes in gene expression, and the global changes to potato root gene-expression correlates with resistance to root-knot nematode infections. To develop a delivery method for this defence peptide, we engineered a commercial strain of the rhizobacteria B. subtilis to secrete StPep1. We have demonstrated, as a proof of concept, that non-plant delivery of defence-inducing StPep1 to potato roots can substantially increase the resistance to root-knot nematodes (Fig. 3). The adaptability of the Bacillus system provides great potential for secretion of other types of proteins and peptides and tailoring the secreted immunostimulants for other pathosystems.
Methods
Supplementary Methods includes the sections ‘Nematode cultures’, ‘Nematode viability assays’, ‘RNA-seq experiment and analysis’ and ‘Isolation and enumeration of B. subtilis on potato roots in sand.’
Gene-expression analysis by RT–qPCR
For gene-expression analysis, 3-week-old potato tissue culture plants were first incubated by immersing the roots in deionized H2O for 24 h. The water was replaced by 100 nM StPep1 solution or water. StPep1 (amino acid sequence ATERRGRPPSRPKVGSGPPPQNN) was synthesized by GenScript. Root or leaf samples were collected at indicated treatment times. The Russet Burbank potato cultivar was used in all experiments, unless stated otherwise. The StCOI1-RNAi potato and its corresponding wild-type lines were in the Désirée background18. The NahG-expressing potato line and its corresponding wild-type line are in the Bannock Russet background. Total RNA was extracted from frozen leaf or root samples using RNeasy Plant Mini Kit (QIAGEN) according to the manufacturer’s instructions. cDNA was produced using ProtoScript II First Strand cDNA Synthesis Kit (New England Biolabs) with an oligo-dT primer. RT–qPCR was performed using SsoAdvanced Universal SYBR Green Supermix on a CFX96 Real-Time PCR Detection System (Bio-Rad). The PCR conditions were: 95 °C for 3 min, 40 cycles of 95 °C for 15 s, 53 °C for 15 s and 72 °C for 20 s, followed by a melting curve analysis from 65 °C to 95 °C with 0.5 °C incremental increases. Expression levels of genes of interest in potato were normalized to the expression of the potato housekeeping gene elongation factor 1-alpha (StEF1α). The relative expression levels of each target gene following StPep1 treatment were calculated by comparing with those of corresponding mock treatments using the \(2^{-\Delta \Delta C_{\mathrm{t}}}\) method20. The primer pairs used for RT–qPCR are summarized in Supplementary Table 2. Each RT–qPCR experiment consisted of two technical replicates per treatment and gene, and the average of three biological replicates is shown.
Statistical tests were performed using GraphPad Prism v.8.00 for Windows (GraphPad Software). Student’s t-tests were performed unless the data had unequal variances based on the Brown-Forsythe test, in which case Welch’s t-test was used. One-way ANOVA followed by Tukey’s test was also performed using GraphPad Software. Exact P values are listed in Supplementary Table 3.
Nematode infection assays
Three- to four-week-old Russet Burbank tissue culture plantlets were transferred to 500 ml cones filled with sand and grown in growth chambers (14 h:10 h light:dark cycle, 23 °C) for 14 d. Subsequently, the potato plants were watered with 10 ml of StPep1 solution (1 µM). Two days after peptide treatment, the plants were inoculated with 500 freshly-hatched M. chitwoodi (McRoza) juveniles. The plants were collected at 12 dpi and the number of galls was recorded. The fresh weights of roots and shoots were also measured. To count egg masses, plants were collected at 35 dpi and roots were stained with phloxin B solution (0.15 g l−1) to visualize egg masses. To visualize parasitic J2s inside potato roots, plants were collected at 4 dpi, and roots were stained with acid fuchsin solution (0.35% acid fuchsin, 25% acetic acid).
For B. subtilis pre-treatment, B. subtilis strains were grown overnight in Luria–Bertani (LB) medium with kanamycin (10 µg ml−1) with shaking at 29 °C. The cultures were then diluted by 1:100 in LB medium with kanamycin (10 µg ml−1) and incubated with shaking until the optical density at 600 nm (OD600) was 0.6. The bacterial cultures were spun down and pellets were resuspended in one volume of deionized H2O. Potato tissue culture plant roots were immersed in suspensions of B. subtilis strains for 2 h before the plants were transferred to sand Cone-tainers. Ten millilitres of the B. subtilis suspensions (OD600 = 0.6) were used to treat the Russet Burbank plants in cones with sand 10 d after transfer. Four days after treatment, the potato plants were inoculated with 500 M. chitwoodi (McRoza) J2s as described above.
Each assay for galling or egg masses used ten potato plants and was repeated three times. The nematode invasion assay was repeated twice.
Treatment of potato plants with B. subtilis strains for gene-expression analysis
Potato tissue culture plants were incubated by immersing roots in a 6-well plate with deionized H2O for 24 h with gentle shaking at 80 rpm. Concentrated bacterial suspensions in water of B. subtilis transformed with pBE-S empty vector or pBE-S with the sequence for StPep1–His were then added to wells of 6-well plates to reach an OD600 of 0.8–1.0. The plates were shaken at 80 rpm. Root samples were collected after 6 h of treatment for total RNA extraction and quantitative rtPCR as described above.
Transformation of B. subtilis
The DNA sequence encoding StPep1 was optimized for better expression in the bacterium B. subtilis using the codon-optimization tool (Integrated DNA Technologies). The codon-optimized StPep1 gene was synthesized by PCR using primers with overlapping regions, and two versions of the gene were obtained, one with a stop codon and the other without a stop codon. The PCR products were cloned into the pBE-S vector (Takara) via SacI and XbaI restriction enzyme sites. The StPep1 gene without stop codon was fused with a sequence encoding a 6×His tag at the C terminus. The ligated plasmids were introduced into E. coli One Shot TOP10 chemically competent cells (Thermo Fisher Scientific). The sequences of the plasmids were confirmed by DNA sequencing (Elim Biopharmaceuticals). The verified plasmids were introduced into B. subtilis RIK1285 by the electroporation method. Colony PCR was used to confirm transformation of B. subtilis RIK1285 with the correct plasmids.
Confirmation of StPep1–His secretion by B. subtilis
Colonies of StPep1–His-transformed or wild-type B. subtilis were grown in LB medium with kanamycin (10 µg ml−1) or LB medium only, respectively, at 29 °C overnight. The B. subtilis cultures were diluted in 30 ml LB medium and were kept shaking until OD600 reached 0.8. One millilitre of the bacterial culture was used for crude protein extraction and expression of the StPep1–His fusion protein was detected by western blot using anti-His antibody (Santa Cruz Biotechnology). Supernatants were collected by centrifugation of solid material. These supernatants were used for purification of His-tagged proteins by the Capturem His-Tagged Purification Maxiprep Kit (TaKaRa) according to the manufacturer’s instructions. Concentration of His-tagged peptides was estimated using the His Tag Detection Kit (GenScript), a competitive enzyme-linked immunosorbent assay, following the manufacturer’s protocol, with horseradish peroxidase reading outputs obtained using a SpectraMax Plus384 microplate reader (Molecular Devices).
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The datasets generated in the study are either included in this publication or available from the corresponding author on reasonable request.
References
European and Mediterranean Plant Protection Organization. Meloidogyne chitwoodi and Meloidogyne fallax. EPPO Bull. 39, 5–17 (2009).
Santo, G. S., O’Bannon, J. H., Finley, A. M. & Golden, A. M. Occurrence and host range of a new root-knot nematode (Meoidogyne chitwoodi) in the Pacific Northwest. Plant Dis. 64, 951–952 (1980).
Mojtahedi, H., Brown, C. R., Riga, E. & Zhang, L. H. A new pathotype of Meloidogyne chitwoodi race 1 from Washington state. Plant Dis. 91, 1051 (2007).
Quintana-Rodriguez, E., Duran-Flores, D., Heil, M. & Camacho-Coronel, X. Damage-associated molecular patterns (DAMPs) as future plant vaccines that protect crops from pests. Sci. Hort. 237, 207–220 (2018).
Huffaker, A., Dafoe, N. J. & Schmelz, E. A. ZmPep1, an ortholog of Arabidopsis elicitor peptide 1, regulates maize innate immunity and enhances disease resistance. Plant Physiol. 155, 1325–1338 (2011).
Huffaker, A., Pearce, G. & Ryan, C. A. An endogenous peptide signal in Arabidopsis activates components of the innate immune response. Proc. Natl Acad. Sci. USA 103, 10098–10103 (2006).
Huffaker, A. et al. Plant elicitor peptides are conserved signals regulating direct and indirect antiherbivore defense. Proc. Natl Acad. Sci. USA 110, 5707–5712 (2013).
Liu, Z. et al. BIK1 interacts with PEPRs to mediate ethylene-induced immunity. Proc. Natl Acad. Sci. USA 110, 6205–6210 (2013).
Lori, M. et al. Evolutionary divergence of the plant elicitor peptides (Peps) and their receptors: interfamily incompatibility of perception but compatibility of downstream signalling. J. Exp. Bot. 66, 5315–5325 (2015).
Tintor, N. et al. Layered pattern receptor signaling via ethylene and endogenous elicitor peptides during Arabidopsis immunity tobacterial infection. Proc. Natl Acad. Sci. USA 110, 6211–6216 (2013).
Huffaker, A. & Ryan, C. A. Endogenous peptide defense signals in Arabidopsis differentially amplify signaling for the innate immune response. Proc. Natl Acad. Sci. USA 104, 10732–10736 (2007).
Lee, M. W. et al. Plant elicitor peptides promote plant defences against nematodes in soybean. Mol. Plant Pathol. 19, 858–869 (2018).
Warnock, N. D. et al. Nematode neuropeptides as transgenic nematicides. PLoS Pathol. 13, e1006237 (2017).
Karasov, T. L., Chae, E., Herman, J. J. & Bergelson, J. Mechanisms to mitigate the trade-off between growth and defense. Plant Cell 29, 666 (2017).
Holmes, D. R., Grubb, L. E. & Monaghan, J. The jasmonate receptor COI1 is required for AtPep1-induced immune responses in Arabidopsis thaliana. BMC Res. Notes 11, 555 (2018).
Wiesel, L. et al. A transcriptional reference map of defence hormone responses in potato. Sci. Rep. 5, 15229 (2015).
Poncini, L. et al. In roots of Arabidopsis thaliana, the damage-associated molecular pattern AtPep1 is a stronger elicitor of immune signalling than flg22 or the chitin heptamer. PLoS ONE 2017, e0185808 (2017).
Halim, V. A. et al. PAMP-induced defense responses in potato require both salicylic acid and jasmonic acid. Plant J. 57, 230–242 (2009).
Bartels, S. & Boller, T. Quo vadis, Pep? Plant elicitor peptides at the crossroads of immunity, stress, and development. J. Exp. Bot. 66, 5183–5193 (2015).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Acknowledgements
We acknowledge support by Washington State University and by the United States Department of Agriculture National Institute of Food and Agriculture, Hatch umbrella project no. 1015621. We thank S. Rosahl for the StCOI1‐RNAi lines; R. Navarre for the NahG potato plants; C. Brown for the M. chitwoodi Roza pathotype; C. Xia and Q. Sun for help with the RNA-seq analyses; L. Thomashow and M. Yang for help with B. subtilis extraction and enumeration; S. Izaguirre and M. De Leon for their help with growing potato plants and for their assistance in the laboratory; and Z. Dubois for illustration services.
Author information
Authors and Affiliations
Contributions
L.Z. designed and performed experiments. L.Z. and C.G. analysed data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Shahid Siddique and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods and Figs. 1–10.
Supplementary Tables
Supplementary Table 1: list of differentially regulated genes identified in the RNA-seq analysis in this study (6 h/0 h). Supplementary Table 2: list of primers used in this study. Supplementary Table 3: statistics and reproducibility—exact P values for Figs. 1 and 2, and Supplementary Figs. 1, 2, 5 and 8–10.
Rights and permissions
About this article
Cite this article
Zhang, L., Gleason, C. Enhancing potato resistance against root-knot nematodes using a plant-defence elicitor delivered by bacteria. Nat. Plants 6, 625–629 (2020). https://doi.org/10.1038/s41477-020-0689-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-0689-0
This article is cited by
-
Small secreted peptides (SSPs) in tomato and their potential roles in drought stress response
Molecular Horticulture (2023)
-
Root-Knot Nematodes (Meloidogyne spp.): Biology, Plant-Nematode Interactions and Their Environmentally Benign Management Strategies
Gesunde Pflanzen (2023)
-
Plant immunity inducers: from discovery to agricultural application
Stress Biology (2022)
-
New allies to fight worms
Nature Plants (2020)