Abstract
Sweet potato (Ipomoea batatas L.) is a major root crop worldwide. Sweet potato weevils (SPWs) pose one of the most significant challenges to sweet potato production in tropical and subtropical regions, causing deleterious economic and environmental effects. Characterizing the mechanisms underlying natural resistance to SPWs is therefore crucial; however, the genetic basis of host SPW resistance (SPWR) remains unclear. Here we obtained two sweet potato germplasm with high SPWR and, by map-based cloning, revealed two major SPW-resistant genes—SPWR1 and SPWR2—that are important regulators of natural defence against SPWs. The SPW-induced WRKY transcriptional factor SPWR1 directly activates the expression of SPWR2, and SPWR2, the conserved dehydroquinate synthase, promotes the accumulation of quinate derivative metabolites that confer SPWR in sweet potato. Generally, our results provide new insights into the molecular mechanism underlying sweet potato–SPW interactions and will aid future efforts to achieve eco-friendly SPW management.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The sequencing data used in this study have been deposited in the Genome Sequence Archive (GSA) database under accession number PRJNA830291 and PRJNA827476. Source data are provided with this paper.
References
Loebenstein, G. & Thottappilly, G. (eds) The Sweetpotato (Springer, 2009).
Sheikha, A. F. E. & Ray, R. C. Potential impacts of bio-processing of sweet potato: review. Crit. Rev. Food Sci. Nutr. 57, 455–471 (2015).
Chalfant, R. B. et al. Ecology and management of sweet-potato insects. Annu. Rev. Entomol. 35, 157–180 (1990).
Nottingham, S. F., Son, K. C., Wilson, D. D., Severson, R. F. & Kays, S. J. Feeding and oviposition preferences of sweet potato weevil, Cylas formicarius elegantulus (Summers), on storage roots of sweet potato cultivars with differing surface chemistries. J. Chem. Ecol. 15, 895–903 (1989).
Okonya, J., Okonya, J. S. & Kroschel, J. Incidence, abundance and damage by the sweet potato butterfly (Acraea acerata Hew. and the African sweet potato weevils (Cylas spp.) across an altitude gradient in Kabale district, Uganda. Int. J. Agric. Sci. 3, 814–824 (2013).
Addo-Bediako, A., Tameru, B., Jackai, L. E. N. & Bonsi, C. K. Assessment of risk of introduction of Cylas formicarius elegantulus (Coleoptera: Brentidae) into weevil-free areas in the southern United States. J. Econ. Entomol. 100, 315–321 (2007).
Korada, R. R., Naskar, S. K., Prasad, A. R., Prasuna, A. L. & Jyothi, K. N. Differential volatile emission from sweet potato plant: mechanism of resistance in sweet potato for weevil Cylas formicarius (Fab.). Curr. Sci. 99, 1597–1601 (2010).
Cockerham, K. L. & Deen, O. T. Resistance of new sweetpotato seedlings and varieties to attack by the sweetpotato weevil. J. Econ. Entomol. 40, 439–441 (1947).
Mullen, M. A. et al. Rapid selection of sweet potato lines resistant to the sweetpotato weevil. HortScience 15, 70–71 (1980).
Nokihara, K., Okada, Y., Ohata, S. & Monden, Y. Transcriptome analysis reveals key genes involved in weevil resistance in the hexaploid sweetpotato. Plants 10, 1535 (2021).
Yang, J. et al. Haplotype-resolved sweet potato genome traces back its hexaploidization history. Nat. Plants 3, 696–703 (2017).
Yan, M. et al. Exploring and exploiting genetics and genomics for sweetpotato improvement: status and perspectives. Plant Comm. 3, 100332 (2022).
Gatehouse, J. A. Plant resistance towards insect herbivores: a dynamic interaction. New Phytol. 156, 145–169 (2002).
Erb, M. & Reymond, P. Molecular interactions between plants and insect herbivores. Ann. Rev. Plant Biol. 70, 527–557 (2019).
MithoFer, A. & Boland, W. Plant defense against herbivores: chemical aspects. Ann. Rev. Plant Biol. 63, 431–450 (2012).
Okada, Y. et al. Genome-wide association studies (GWAS) for yield and weevil resistance in sweet potato (Ipomoea batatas (L.) Lam). Plant Cell Rep. 38, 1383–1392 (2019).
Yada, B. et al. Identification of simple sequence repeat markers for sweetpotato weevil resistance. Euphytica 213, 129 (2017).
Anyanga, M. O. et al. Resistance to the weevils Cylas puncticollis and Cylas brunneus conferred by sweetpotato root surface compounds. J. Agric. Food Chem. 61, 8141–8147 (2013).
Anyanga, M. O. et al. Segregation of hydroxycinnamic acid esters mediating sweetpotato weevil resistance in storage roots of sweetpotato. Front. Plant Sci. 8, 1011 (2017).
Liao, Y. et al. Induced biosynthesis of chlorogenic acid in sweetpotato leaves confers the resistance against sweetpotato weevil attack. J. Adv. Res. 24, 513–522 (2020).
Stathers, T. E. et al. Sweetpotato infestation by Cylas spp. in East Africa: II. Investigating the role of root characteristics. Int. J. Pest Manag. 49, 141–146 (2003).
Sutherland, J. A. Damage by Cylas formiearius (Fab.) to sweet potato vines and tubers, and the effect of infestations on total yield in Papua New Guinea. Trop. Pest Manag. 32, 316–323 (1986).
Hahn, S. K. & Leuschner, K. Resistance of sweet potato cultivars to African sweet potato weevil. Crop Sci. 21, 499–503 (1981).
Reinhard, H. J. The sweet potato weevil. Tex. Agric. Exp. Stn Bull. 308, 90 (1923).
Sherman, M. Chemical control of sweet potato insects in Hawaii. J. Econ. Entomol. 44, 652–656 (1951).
Nottingham, S. F., Son, K. C., Wilson, D. D., Severson, R. F. & Kays, S. J. Feeding by adult sweet potato weevils, Cylas formicarius elegantulus, on sweet potato leaves. Entomol. Exp. Appl. 48, 157–163 (1988).
Chen, M. et al. Genome-wide identification of agronomically important genes in outcrossing crops using OutcrossSeq. Mol. Plant 14, 556–570 (2021).
Carpenter, E. et al. Structure of dehydroquinate synthase reveals an active site capable of multistep catalysis. Nature 394, 299–302 (1998).
Hue, S.-M. & Low, M.-Y. An insight into sweet potato weevils management: a review. Psyche 2015, 849560 (2015).
Jiang, J. et al. WRKY transcription factors in plant responses to stresses. J. Integr. Plant Biol. 59, 86–101 (2017).
Harrison, H. F. et al. Contents of caffeoylquinic acid compounds in the storage roots of sixteen sweetpotato genotypes and their potential biological activity. J. Am. Soc. Hortic. Sci. 133, 492–500 (2008).
Kirsten, A. et al. Identification of chlorogenic acid as a resistance factor for thrips in chrysanthemum. Plant Physiol. 150, 1567–1575 (2009).
Cole, R. A. Relationship between the concentration of chlorogenic acid in carrot roots and the incidence of carrot fly larval damage. Ann. Appl. Biol. 106, 211–217 (1985).
Jassbi, A. R. Secondary metabolites as stimulants and antifeedants of Salix integra for the leaf beetle Plagiodera versicolora. Z. Naturforsch. C 58, 573–579 (2003).
Cole, R. A. Phenolic acids associated with the resistance of lettuce cultivars to the lettuce root aphid. Ann. Appl. Biol. 105, 129–145 (1984).
Sun, X. et al. SLAF-seq: an efficient method of large-scale de novo SNP discovery and genotyping using high-throughput sequencing. PLoS ONE 8, e58700 (2013).
Li, R. et al. SOAP: short oligonucleotide alignment program. Bioinformatics 24, 713–714 (2008).
Zhang, J. et al. High-density genetic map construction and identification of a locus controlling weeping trait in an ornamental woody plant (Prunus mume Sieb. et Zucc). DNA Res. 22, 183–191 (2015).
Liu, D. et al. Construction and analysis of high-density linkage map using high-throughput sequencing data. PLoS ONE 9, e98855 (2014).
Van Ooijen, J. W. MapQTL® 6. Software for the mapping of quantitative trait loci in experimental populations of diploid species (Kyazma, 2009).
Royo, C. et al. The major origin of seedless grapes is associated with a missense mutation in the MADS-box gene VviAGL11. Plant Physiol. 177, 1234–1253 (2018).
Liu, X. et al. The NF-YC–RGL2 module integrates GA and ABA signalling to regulate seed germination in Arabidopsis. Nat. Commun. 7, 12768 (2016).
Tahara, K. et al. Dehydroquinate dehydratase/shikimate dehydrogenases involved in gallate biosynthesis of the aluminum-tolerant tree species Eucalyptus camaldulensis. Planta 253, 3–18 (2021).
Chen, C. et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 13, 1194–1202 (2020).
Kanehisa, M. et al. The KEGG resource for deciphering the genome. Nucleic Acids Res. 32, 277–280 (2004).
Acknowledgements
We thank B. Fang and Z. Wang at Guangdong Academy of Agricultural Sciences for providing Beauregard cultivar, and D. Ma and Q. Cao at Xuzhou Academy of Agricultural Sciences/Sweet Potato Research Institute for providing technical advice. This work was supported by the National Natural Science Foundation of China-Guangdong Natural Science Foundation Joint Project (grant no. U1701234 to X.H.), National Natural Science Foundation of China (grant no. 31970623 to X.L.), Guangdong Special Support Plan Project (grant no. 2019TQ05N140 to X.L.) and the Guangzhou Municipal Science and Technology Project (grant no. 202002030057 to X.L.).
Author information
Authors and Affiliations
Contributions
X.L. and X.H. designed the experiments and managed the projects. X.L., Y.W., H.Z., G.M., Y.L., S.R., S.L., A.C., H.L., L.Z. and Y.X. performed the experiments. X.L., Y.W., H.Z., Y.L., X.Li., G.M., Z.Y. and X.H. performed the data analysis. X.L. and X.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Guangcun He, Youjun Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 An overview of the collecting regions of sweet potato germplasms in this study.
a, Global SPW infestation regions (left panel) and the collecting places of sweet potato germplasms in East Asia (right panel). The worldwide areas of SPW threat are shown in pink. Deep pink indicates the tropical area with more serious SPW infestation than that marked by light pink. Cultivar sources are marked by blue spots. The collecting regions of sweet potato germplasms are marked in red. The maps were drawn using ArcGIS software for desktop (https://desktop.arcgis.com/en/). Map scale = 1500 km. b, Screening and identification of the high SPWR candidate germplasms in the field during 2015 to 2017. Stem base damage is recorded in three grades: abundant wormholes, partly intact, and intact, according to degree of SPW damage. Tuber damage index = [∑ (the number of tubers in each damage grade × its damage grade number)]/(the total number of tubers × the highest-grade number). Tuber damage grades: 1 = 2-5 wormholes; 2 = 6–10 wormholes; 3 > 10 wormholes, which were surveyed by dissecting the tubers (modified according to Stathers et al., 2003)21. Data are presented as means ± SD of ten tubers with randomized blocks of three replicates per year.
Extended Data Fig. 2 . Evaluation of SPWR in stem, tuber, leaf, and storage tuber of sweet potato.
Supplementary Fig. 2a, Evaluation of SPWR in the stems and tubers of G87, N73, and N28 under field conditions. Left panel presents the SPWR phenotype of stems and tubers of sweet potato. Red arrows indicate the damaged stem base and tuber by SPW adults and larvae. Scale bar = 1 cm. Right panel shows the tuber damage index recorded in the field during 2015 to 2017. Data are presented as means ± SD of nine replicates. Asterisks indicate significant difference compared with G87 (Two-tailed Student’s t-test, *** P < 0.001, N73 vs G87, P = 6.022 × 10−10; N28 vs G87, P = 3.105 × 10−10). b, SPWR phenotype of the storage tubers of N73, N28, and G87 described in Fig. 1a. Newly hatched adults from tubers were shown at right side. Scale bar = 1 cm. c, The method of the SPW-fed leaves scanning (left) and feeding area capture (right; black color indicates the SPW damage regions). The feeding areas were calculated using ImageJ software (http://imagej.nih.gov/ij/). Scale bar = 1 cm.
Extended Data Fig. 3 Comparison of SPWR of F1 individuals between two independent biological replicates.
a, Comparison (left) and correlation (right) of relative feeding areas of F1 progeny between repeat 1 and repeat 2. The feeding area of F1 individual No. 001 and No. 016 was set to 1 in repeat 1 and repeat 2, respectively. b, Distribution of SPWR grades of F1 progeny in repeat 1 (left) and repeat 2 (right). SPW grades were set to I-X (Roman numeral) according to feeding area (grade X represents the biggest feeding area in F1 progeny). Values present the number of F1 individuals per grade of SPWR. Arrows indicate the places of parents.
Extended Data Fig. 4 Genetic map construction and QTL analysis of SPWR trait.
a, Sweet potato genetic map was constructed according to genome resequencing of parents and SLAF-sequencing of F1 progeny based on Taizhong 6 genome. The HighMap software is used to analyze the linear arrangement of the markers in the linkage group marked as a unit. Each chromosome is a linkage group. The number of markers, linkage groups, total map distance, and average map distance are shown in Supplementary Table 2. b, Whole-chromosome scanning of SPWR QTLs of F1 population in two biological replicates. Pink color marks the QTL loci at chromosome 7 and 9.
Extended Data Fig. 5 QTL analysis of leaf shape trait in the G87 × N73 F1 population.
Leaf shape trait was associated with a locus in chromosome 7, which appears to be the same as the leaf morphology locus as described by Chen et al.27. QTL locus is present in a genetic map distance locus/gene. The loci on chromosome 7 identified by two methods are identical.
Extended Data Fig. 6 Fine-mapping of SPWR1 and SPWR2 by G87 × N73 F1 population.
a, Delimitation of the SPWR1 locus to a 0.14 Mb region of chromosome 9. In all used individual recombinants, only the SPWR1 locus was segregated while the SPWR2 locus is the genotype of N73 parent. b, Delimitation of the SPWR2 locus to a 0.07 Mb region of chromosome 7. In all used individual recombinants, only the SPWR2 locus was segregated while the SPWR1 locus is the genotype of N73 parent. Recombinant F1 individuals were marked with black digitals. The genotype of recombinants is presented in two colors: orange indicates G87 parent genotype and green indicates N73 parent genotype. The phenotype and resistance grade are presented at the right side of recombinants. Resistant phenotype is the grades I-III (see Supplementary Table 5). Markers and genome information were obtained according to the Taizhong 6 reference genome.
Extended Data Fig. 7 Transcriptomic analysis of SPWR-related genes in G87 and N73.
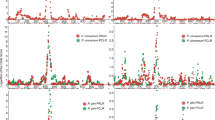
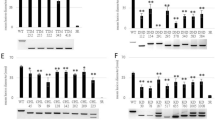
a, Quality control of transcriptome of G87, N73, and OE-SPWR1 leaves with SPW treatment or not (control). The two biological replicates for each sample were marked with −1 and −2, respectively. b, GO analysis of differentially expressed genes (DEGs) involved in metabolic pathways in N73 compared with G87 under SPW treatment. Squares indicate the number of DEGs enriched in the pathway. The color represents the q-value. Red-boxes indicate shikimate pathway-related metabolisms. c,d, Transcriptomic expression of genes located at SPWR1 (c) and SPWR2 (d) find-mapping regions under control and SPW treatment. The values represent RPKM value of genes in transcriptome. Data are presented as means ± SD of two biological replicates.
Extended Data Fig. 8 Fig. 8. Protein sequence alignment and phylogenetic analysis of SPWR1.
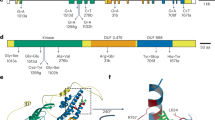
a, Protein sequence alignment of SPWR1 in G87, N73, and N28. Blue box indicates the conserved WRKY domain. Red boxes and phrases indicate the amino acid substitutions and indel site. Sequences were aligned with Clustal X. b, Phylogenetic tree shows the predicted SPWR1 homologs in different plant species. NCBI accessions: SPWR1 (Ipomoea batatas), OM283291; WRKY-like (Ipomoea triloba), XP_031129806.1; WRKY-like (Ipomoea nil), XP_019170409.1; WRKY-like (Nicotiana attenuata), XP_019233122.1; WRKY-like (Nicotiana tabacum), XP_016453841.1; WRKY-like (Solanum tuberosum), XP_006352253.1; WRKY-like (Salvia splendens), XP_042022560.1; WRKY-like (Sesamum indicum), XP_020551475.1; WRKY-like (Olea europaea var. sylvestris), XP_022886946.1; WRKY-like (Actinidia rufa), GFZ16403.1; WRKY-like (Theobroma cacao), XP_007040478.2. Phylogenetic analysis was performed by maximum likelihood (ML) method using MEGA7.1 with 1000 bootstrap replicates to build the ML tree.
Extended Data Fig. 9 Protein and promoter sequence alignment and phylogenetic analysis of SPWR2.
a, Protein sequence alignment of SPWR1 in G87, N73, and N28. Blue box indicates the conserved DHQS domain. Red boxes and phrases indicate the amino acid substitutions. b, Promoter sequence alignment of SPWR2 in G87, N73, and N28. Red boxes and phrases indicate the Indels. Blue box indicates the W-box element in Indel1. Sequences were aligned with Clustal X. c, Phylogenetic tree shows the predicted SPWR2 homologs in different species. NCBI accessions: SPWR2 (Ipomoea batatas), OM283290; DHQS-like (Ipomoea nil), XP_031109196.1; DHQS-like (Nicotiana tabacum), XP_016464940.1; DHQS-like (Solanum tuberosum), XP_006340763.1; DHQS-like (Solanum lycopersicum), NP_001233863.1; DHQS-like (Olea europaea var. sylvestris), XP_022885746.1; DHQS-like (Coffea arabica), XP_027073560.1; DHQS-like (Arabidopsis thaliana), NP_001030791.1; DHQS-like (Oryza sativa), XP_015612522.1; DHQS-like /aroB (Escherichia coli), NP_417848.1. Phylogenetic analysis was performed by maximum likelihood (ML) method using MEGA7.1 with 1000 bootstrap replicates to build the ML tree.
Extended Data Fig. 10 Generation and expression identification of the SPWR1 and SPWR2 transgenic sweet potatoes.
a, Southern blot analysis of the transgenic sweet potato lines using a DIG-labeled probe for detection of HPT (Hygromycin Phosphotransferase) gene. M, DIG-labeled lambda/Hind III Marker (TIANDZ, Inc., China); W, the wild type G87; lanes 1-4, OE-SPWR1 transgenic lines; lanes 5-8, OE-SPWR2 transgenic lines; 9-10, SPWR1-RNAi transgenic lines; lanes 11-12, SPWR2-RNAi transgenic lines. All genomic DNA was digested with Hind III. b,c, Gene expression analysis of SPWR1 (b) and SPWR2 (c) in different transgenic lines. d, The phenotype of SPWR1 and SPWR2 transgenic lines at 3 WAC. Scale bar = 1 cm. Data are presented as means ± SD of nine biological replicates. Asterisks indicate significant difference (Two-tailed Student’s t-test, ** P < 0.01, *** P < 0.001). Tukey’s multiple comparison was performed by One-way ANOVA, and statistically significant differences are indicated by different lower-case letters. All exact P values were shown in the relative Source Data Extended Data Fig. 10.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10, Method and List of supplementary files.
Supplementary Tables
Supplementary Table 1 Sweet potato germplasm for field screening. Supplementary Table 2 Information of genome resequencing and SLAF sequencing of the parents and F1 progeny. Supplementary Table 3 Two repeated QTL detections in the F1 population of G87 × N73. Supplementary Table 4 Molecular markers and sequencing primers for fine mapping. Supplementary Table 5 Resistance grades of F1 individuals in two repeats. Supplementary Table 6 Genotype of the selected recombinations for fine mapping. Supplementary Table 7 Predicted genes list in the mapped 0.14 Mb genomic region of SPWR1 locus according to the Taizhong 6 genome. Supplementary Table 8 Predicted genes list in the mapped 0.07 Mb genomic region of SPWR2 locus according to the Taizhong 6 genome. Supplementary Table 9 Primers used in this study.
Supplementary Video
Demonstration of the inhibitory effect of compounds on the movement of SPWs within 20 min after directly feeding metabolites to SPWs. Quinate, quinic acid; CGA, chlorogenic acid; 1-CA, 1-caffeoylquinic acid. The three vertical wells show three replicates for each treatment.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed gels.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, X., Wang, Y., Zhu, H. et al. Natural allelic variation confers high resistance to sweet potato weevils in sweet potato. Nat. Plants 8, 1233–1244 (2022). https://doi.org/10.1038/s41477-022-01272-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-022-01272-1