Abstract
While nuclear lamina abnormalities are hallmarks of human diseases, their interplay with epigenetic regulators and precise epigenetic landscape remain poorly understood. Here, we show that loss of the lysine acetyltransferase MOF or its associated NSL-complex members KANSL2 or KANSL3 leads to a stochastic accumulation of nuclear abnormalities with genomic instability patterns including chromothripsis. SILAC-based MOF and KANSL2 acetylomes identified lamin A/C as an acetylation target of MOF. HDAC inhibition or acetylation-mimicking lamin A derivatives rescue nuclear abnormalities observed in MOF-deficient cells. Mechanistically, loss of lamin A/C acetylation resulted in its increased solubility, defective phosphorylation dynamics and impaired nuclear mechanostability. We found that nuclear abnormalities include EZH2-dependent histone H3 Lys 27 trimethylation and loss of nascent transcription. We term this altered epigenetic landscape “heterochromatin enrichment in nuclear abnormalities” (HENA). Collectively, the NSL-complex-dependent lamin A/C acetylation provides a mechanism that maintains nuclear architecture and genome integrity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The MS proteomics data for Mof KO have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the PRIDE partner repository (accession number: PXD008539). MALBAC single-cell genomic sequencing data have been deposited to ENA (accession number: PRJEB27801). Source data for all figures in the main paper and its Supplementary Information where statistics were conducted have been provided as Supplementary Table 7. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Change history
27 February 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41556-023-01106-y
References
Allis, C. D. & Jenuwein, T. The molecular hallmarks of epigenetic control. Nat. Rev. Genet. 17, 487–500 (2016).
Burke, B. & Stewart, C. L. The nuclear lamins: flexibility in function. Nat. Rev. Mol. Cell Biol. 14, 13–24 (2013).
Van Steensel, B. & Belmont, A. S. Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 169, 780–791 (2017).
Raices, M. & D’Angelo, M. A. Nuclear pore complex composition: a new regulator of tissue-specific and developmental functions. Nat. Rev. Mol. Cell Biol. 13, 687–699 (2012).
Hutchison, C. J. & Worman, H. J. A-type lamins: guardians of the soma? Nat. Cell Biol. 6, 1062–1067 (2004).
Liu, B. et al. Genomic instability in laminopathy-based premature aging. Nat. Med. 11, 780–785 (2005).
Dittmer, T. A. et al. The lamin protein family. Genome Biol. 12, 222 (2011).
Bell, E. S. & Lammerding, J. Causes and consequences of nuclear envelope alterations in tumour progression. Eur. J. Cell Biol. 95, 449–464 (2016).
Akhtar, A. & Becker, P. B. Activation of transcription through histone H4 acetylation by MOF, an acetyltransferase essential for dosage compensation in Drosophila. Mol. Cell 5, 367–375 (2000).
Samata, M. & Akhtar, A. Dosage compensation of the X chromosome: a complex epigenetic assignment involving chromatin regulators and long noncoding RNAs. Annu. Rev. Biochem. 87, 323–350 (2018).
Keller, C. I. & Akhtar, A. The MSL complex: juggling RNA–protein interactions for dosage compensation and beyond. Curr. Opin. Genet. Dev. 31, 1–11 (2015).
Chelmicki, T. et al. MOF-associated complexes ensure stem cell identity and Xist repression. Elife 3, e02024 (2014).
Gupta, A. et al. The mammalian ortholog of Drosophila MOF that acetylates histone H4 lysine 16 is essential for embryogenesis and oncogenesis. Mol. Cell. Biol. 28, 397–409 (2008).
Thomas, T., Dixon, M. P., Kueh, A. J. & Voss, A. K. Mof (MYST1 or KAT8) is essential for progression of embryonic development past the blastocyst stage and required for normal chromatin architecture. Mol. Cell. Biol. 28, 5093–5105 (2008).
Sheikh, B. N. et al. MOF maintains transcriptional programs regulating cellular stress response. Oncogene 35, 2698–2710 (2016).
Pfister, S. et al. The histone acetyltransferase hMOF is frequently downregulated in primary breast carcinoma and medulloblastoma and constitutes a biomarker for clinical outcome in medulloblastoma. Int. J. Cancer 122, 1207–1213 (2008).
Fraga, M. F. et al. Loss of acetylation at Lys16 and trimethylation at Lys20 of histone H4 is a common hallmark of human cancer. Nat. Genet. 37, 391–400 (2005).
Sheikh, B. N. & Akhtar, A. The many lives of KATs — detectors, integrators and modulators of the cellular environment. Nat. Rev. Genet. 20, 7–23 (2019).
Chatterjee, A. et al. MOF acetyl transferase regulates transcription and respiration in mitochondria. Cell 167, 722–738.e23 (2016).
Vargas, J. D., Hatch, E. M., Anderson, D. J. & Hetzer, M. W. Transient nuclear envelope rupturing during interphase in human cancer cells. Nucleus 3, 88–100 (2012).
Hatch, E. M. Nuclear envelope rupture: little holes, big openings. Curr. Opin. Cell Biol. 52, 66–72 (2018).
Toda, T. et al. Nup153 interacts with Sox2 to enable bimodal gene regulation and maintenance of neural progenitor cells. Cell Stem Cell 21, 618–634.e7 (2017).
Nanni, S. et al. The nuclear pore protein Nup153 associates with chromatin and regulates cardiac gene expression in dystrophic mdx hearts. Cardiovasc. Res. 112, 555–567 (2016).
Schölz, C. et al. Acetylation site specificities of lysine deacetylase inhibitors in human cells. Nat. Biotechnol. 33, 415–423 (2015).
Kim, S. C. et al. Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol. Cell 23, 607–618 (2006).
Turgay, Y. et al. The molecular architecture of lamins in somatic cells. Nature 543, 261–264 (2017).
Kamieniarz, K. & Schneider, R. Tools to tackle protein acetylation. Chem. Biol. 16, 1027–1029 (2009).
Astejada, M. N. et al. Emerinopathy and laminopathy clinical, pathological and molecular features of muscular dystrophy with nuclear envelopathy in Japan. Acta Myol. 26, 159–164 (2007).
Ben-Harush, K. et al. The supramolecular organization of the C. elegans nuclear lamin filament. J. Mol. Biol. 386, 1392–1402 (2009).
Taimen, P. et al. A progeria mutation reveals functions for lamin A in nuclear assembly, architecture, and chromosome organization. Proc. Natl Acad. Sci. USA 106, 20788–20793 (2009).
Laubach, J. P., Moreau, P., San-Miguel, J. F. & Richardson, P. G. Panobinostat for the treatment of multiple myeloma. Clin. Cancer Res. 21, 4767–4773 (2015).
Bantscheff, M. et al. Chemoproteomics profiling of HDAC inhibitors reveals selective targeting of HDAC complexes. Nat. Biotechnol. 29, 255–265 (2011).
Cho, S., Irianto, J. & Discher, D. E. Mechanosensing by the nucleus: from pathways to scaling relationships. J. Cell Biol. 216, 305–315 (2017).
Denais, C. M. et al. Nuclear envelope rupture and repair during cancer cell migration. Science 352, 353–358 (2016).
Zong, C., Lu, S., Chapman, A. R. & Xie, X. S. Genome-wide detection of single-nucleotide and copy-number variations of a single human cell. Science 338, 1622–1626 (2012).
Zhang, C.-Z. et al. Chromothripsis from DNA damage in micronuclei. Nature 522, 179–184 (2015).
Liu, S. et al. Nuclear envelope assembly defects link mitotic errors to chromothripsis. Nature 561, 551–555 (2018).
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Korbel, J. O. & Campbell, P. J. Criteria for inference of chromothripsis in cancer genomes. Cell 152, 1226–1236 (2013).
Chen, X. et al. ATAC-see reveals the accessible genome by transposase-mediated imaging and sequencing. Nat. Methods 13, 1013–1020 (2016).
Sato, Y. et al. Genetically encoded system to track histone modification in vivo. Sci. Rep. 3, 2436 (2013).
Verma, S. K. et al. Identification of potent, selective, cell-active inhibitors of the histone lysine methyltransferase EZH2. ACS Med. Chem. Lett. 3, 1091–1096 (2012).
Torvaldson, E., Kochin, V. & Eriksson, J. E. Phosphorylation of lamins determine their structural properties and signaling functions. Nucleus 6, 166–171 (2015).
Ding, Y., Xu, G. K. & Wang, G. F. On the determination of elastic moduli of cells by AFM based indentation. Sci. Rep. 7, 1–8 (2017).
Shogren-Knaak, M. et al. Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science 311, 844–847 (2006).
Oppikofer, M. et al. A dual role of H4K16 acetylation in the establishment of yeast silent chromatin. EMBO J. 30, 2610–2621 (2011).
Falk, M. et al. Heterochromatin drives compartmentalization of inverted and conventional nuclei. Nature 570, 395–399 (2019).
Wolberg, W. H., Street, W. N. & Mangasarian, O. L. Importance of nuclear morphology in breast cancer prognosis. Clin. Cancer Res. 5, 3542–3548 (1999).
Peleg, S., Feller, C., Ladurner, A. G. & Imhof, A. The metabolic impact on histone acetylation and transcription in ageing. Trends Biochem. Sci. 41, 700–711 (2016).
López-Otín, C., Blasco, Ma, Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, (2019).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Herrmann, H., Kreplak, L. & Aebi, U. Isolation, characterization, and in vitro assembly of intermediate filaments. Methods Cell Biol. 78, 3–24 (2004).
Makarov, A. A., Rizzotto, A., Meinke, P. & Schirmer, E. C. Purification of Lamins and Soluble Fragments of NETs. Methods Enzymol. 569, 79–100 (2016).
Kubben, N. et al. Identification of differential protein interactors of lamin A and progerin. Nucleus 1, 513–525 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Tischler, G. & Leonard, S. biobambam: tools for read pair collation based algorithms on BAM files. Source Code Biol. Med. 9, 13 (2014).
Rausch, T., Hsi-Yang Fritz, M., Korbel, J. O. & Benes, V. Alfred: interactive multi-sample BAM alignment statistics, feature counting and feature annotation for long- and short-read sequencing. Bioinformatics 35, 2489–2491 (2019).
Stegle, O., Parts, L., Piipari, M., Winn, J. & Durbin, R. Using probabilistic estimation of expression residuals (PEER) to obtain increased power and interpretability of gene expression analyses. Nat. Protoc. 7, 500–507 (2012).
Olshen, A. B., Venkatraman, E. S., Lucito, R. & Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, 333–339 (2012).
Rausch, T. et al. Genome sequencing of pediatric medulloblastoma links catastrophic DNA rearrangements with TP53 mutations. Cell 148, 59–71 (2012).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Jao, C. Y. & Salic, A. Exploring RNA transcription and turnover in vivo by using click chemistry. Proc. Natl Acad. Sci. USA 105, 15779–15784 (2008).
Rapsomaniki, M. A. et al. EasyFRAP: an interactive, easy-to-use tool for qualitative and quantitative analysis of FRAP data. Bioinformatics 28, 1800–1801 (2012).
Koulouras, G. et al. EasyFRAP-web: a web-based tool for the analysis of fluorescence recovery after photobleaching data. Nucleic Acids Res. 46, W467–W472 (2018).
Rog-Zielinska, E. A. et al. Species differences in the morphology of transverse tubule openings in cardiomyocytes. EP Europace 20, 120–124 (2018).
Kremer, J. R., Mastronarde, D. N. & R. M, J. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 76, 71–76 (1996).
Ding, Y., Xu, G. & Wang, G. On the determination of elastic moduli of cells by AFM based indentation. Sci. Rep. 7, 45575 (2017).
Acknowledgements
We are grateful to B. Sheikh, M. Shvedunova, M. Buck, M. Samata, C. Pessoa Rodrigues, S. Lefkopoulos and G. Semplicio for critical reading of the manuscript. We thank E. Trompouki and R. Sawarkar for fruitful discussions and suggestions. We also thank P. Rawat for help with FRAP and M.-F. Basilicata for help with the generation of the Moffl/-, CreT/+ ESCs. The MPI-IE core facilities (for fluorescence-activated cell sorting, deep sequencing, imaging and mouse care), the EMBL IT facilities (for computing) and the EMBL EMCF (for electron microscopy) have been invaluable for this project. This work was supported by CRC992 (A02) and CRC1381 (B3) awarded to A.A., by an ERC Starting Grant awarded to J.O.K. (336045) and the Swiss National Science Foundation (SNSF 31003A_179418, to O.M.). E.A.R.-Z. is an Emmy Noether Fellow (DFG no. 396913060); P.K. acknowledges the ERC Advanced Grant CardioNect (201203).
Author information
Authors and Affiliations
Contributions
A.K. and A.A. conceptualized and designed the experiments and A.K. performed the majority of the experiments with the exception of the following. The in vitro lamin assembly experiments and analysis were performed by R.d.L. and O.M; MS analysis and single-cell genomic DNA bioinformatic analysis were carried out by W.S. and G.M. and by T.R. and J.O.K., respectively; the Kansl2 and Kansl3 conditional KO mice were generated by B.H.; S.G. and H.-R.C. helped with the imaging analysis of the Mof, Kansl2 and Kansl3 KO MEFs; H.K. provided the H3K27me3-mintbody plasmid; J.S. helped with the cell culture and biochemistry experiments; nanoindentation and electron tomography imaging were performed by R.P. and E.A.R.-Z. respectively, together with P.K.; A.K. wrote the manuscript with input from all of the authors; O.M., J.O.K. and A.A. acquired funding; A.A. supervised all aspects of the study. All authors reviewed, edited and approved the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Further characterization of the Mof, Kansl2 and Kansl3 KO nuclear abnormalities.
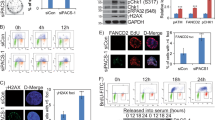
a) Relative Mof mRNA expression (against Rplp0) in control and Mof KO MEFs (4 days). Two-tailed unpaired t-test on n=3 independent biological replicates. Data shown are the mean ± s.e.m. b) Quantification of nuclear abnormalities of Mof fl/-, CreT/- MEFs in the presence or absence of 4OHT; n=2 biological replicates. c) Immunoblotting on whole cell extracts of control and Mof KO MEFs expressing the indicated FLAG-MOF constructs. d) Immunoblotting on whole cell extracts of control and Mof KO MEFs at the indicated time points. e) Immunoblotting on whole cell extracts of control, Kansl2 and Kansl3 KO MEFs. f) Immunoblotting on subcellular fractions of control and Mof KO (2 days) MEFs, where the nucleoplasmic p53 is depicted. Cytopl;Cytoplasmic fraction, Memb;Membrane bound proteins fraction, Nucleopl; Nucleoplasmic fraction, Chrom; Chromatin bound fraction. g) (Same as Fig. 1d) Representative time-lapse imaging snapshots of control, Mof and Kansl2 KO MEFs, transduced with GFP-NLS or GFP-H2B as indicated. 6 independent experiments. The white arrowheads indicate nuclear abnormalities. h) Representative immunostaining of lamin B1 (red) and NUP153 (green) in control, Mof, Kansl2 and Kansl3 KO (4 days) MEFs. dsDNA was counterstained using DAPI. Scale bars, 5μm. i) Representative immunostaining of lamin A (green) and γH2A.X (red) in control and Mof KO (4 days) MEFs. dsDNA was counterstained using DAPI. Scale bars, 5μm. j) Quantification of γH2A.X positive and negative micronuclei on the indicated KO MEFs; Two-tailed unpaired t-test on n=5 (Mof KO) and n=3 (Kansl2 and Kansl3 KO) independent biological replicates. Data shown are the mean ± s.e.m. Uncropped blots of c, d, e, f can be found in Supplementary Fig. 9.
Supplementary Figure 2 Peptide fractionation and SILAC analysis scheme for MEFs and ESCs.
a) Schematic representation of the experimental design used for total proteome and acetylome identification by LC-MS. Cells were labeled with heavy lysine (K8) and arginine (R10) and reciprocal labeling setup (“label swap”) was implemented (Methods). Tryptic peptides were fractionated on a C18 (solid phase extraction) column and underwent two rounds of anti-acetyl (Lys) immunoprecipitation procedures using two different antibody resins. Samples were measured by nano LC-MS/MS (Methods). Additionally, IP input samples were measured. b) Correlation matrix scatter plots with Pearson-r correlation values for acetyl (Lys) sites in replicates from MEFs and ESCs control and Mof KO. Raw ratios were log2 transformed and ratio distributions were adjusted by quartile normalization. c) Histograms representing ratio distributions across replicates in anti-acetyl(Lys) IP samples in MEFs and ESCs. Raw ratio values were log2 transformed (white) and subsequently adjusted by quartile normalization (blue). d) Numerical UpSet diagram representing the overlap between identified acetyl(K)sites within all replicates in MEFs (left panel) and ESCs (right panel). From the total number of 6534 unique acetyl(K)sites in MEFs, 4048 were found in at least 3 replicates. For ESCs, 1964 acetyl(K)sites were found in at least 3 replicates out of 3020 unique acetyl(K)sites. For b, c and d n=6 independent biological replicates.
Supplementary Figure 3 Characterization of the MOF dependent proteome and acetylome deregulation on MEFs and ESCs.
a) Adjusted liquid chromatography profiles for particular IP fractions that exhibit different hydrophobic properties (Methods). b) Graphic representation of histone multiplicity (Supplementary Tables 3, 4 and 6). c) Correlation between different multiplicities of acetylated sites identified in MEFs and ESCs Mof KO. Log2 fold changes of a two-sided t-test performed on n=6 SILAC ratios. Histone acetyl (Lys) sites are highlighted in red and H4K16ac in green. The red line and gray field correspond to a linear smooth fit regression with a 0.95 confidence interval respectively. d) Immunoblotting on whole cell extracts of control and Mof KO ESCs. Uncropped blots in Supplementary Fig. 9. e) Overlap of the MEFs and ESCs Mof KO proteome and acetylome datasets. f) Heatmap illustration of the quantitative proteome changes of Mof KO MEFs and ESCs. Dendrogram illustrates Euclidean distance clustering with 4 identified clusters. Color range indicates the log2 fold changes. g) Enriched gene sets of Supplementary Fig. 3f. (Supplementary Table 5). h) Same as Supplementary Fig. 3f for the commonly detected acetyl (Lys) sites. i) Enriched gene sets of Supplementary Fig. 3h. (Supplementary Table 5). j) Enriched gene sets of the MEFs and ESCs-only acetylome. (Supplementary Table 5). k) Heatmap illustration of all quantified histone acetyl (Lys) sites, in MEFs and ESCs Mof KO. Color range indicates the log2 fold changes. Different multiplicity values were averaged (Supplementary Fig. 3b, c and Supplementary Table 6). Not identified sites are marked as a crossed box. l) Enriched gene sets of the statistically significantly hypoacetylated targets in MEFs and ESCs Mof KO. Color scale depicts the p-value of the statistically enriched terms (Supplementary Table 5). m) Circos plot representation of the GO analysis overlap between significantly hypoacetylated Mof KO targets in MEFs and ESCs. Yellow lines represent similar GO terms (Supplementary Table 5). For panels g, I, j, l, m the n=number of dataset genes within each GO term is defined in Supplementary Table 5.
Supplementary Figure 4 Conservation analysis of the lamin A/C acetyl(Lys)sites.
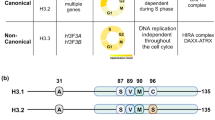
a) Left; Representative immunostaining of lamin B1 (green) and OCT4 (red) in control and Mof KO (4 days) ESCs. dsDNA was counterstained using DAPI. Scale bars, 5μm. Right; Quantification of nuclear abnormalities in control and Mof KO ESCs. n=2 biological replicates. b) Phylogenetic tree of lamin A/C in the indicated organisms based on complete protein sequences. The maximum-likelihood tree was reconstructed using PhyML. (Uniprot annotations; P02545 – Homo sapiens, P48678 - Mus musculus, P13648 - Gallus gallus, B3DKC5 - Danio rerio, Q60HF0 - Macaca fascicularis, Q21443 - Caenorhabditis elegans, F1MYG5 - Bos Taurus, Q03427 - Drosophila melanogaster). c) Vertebrate lamin A/C protein sequence alignment. The lysine (K) residues are highlighted in red. The coiled-coil domain is framed and the detected acetyl(K)sites from the MEFs SILAC are marked with asterisks. Alignment was performed using the SeaView software employing the clustal omega algorithm.
Supplementary Figure 5 Characterization of lamin A/C acetylation in MEFs.
a) Left; immunoblotting on subcellular fractions of WT/K417R OST-lamin A MEFs. Right; Nuclear abnormalities quantification; Two-tailed unpaired t-test on n=3 independent biological replicates. b) Relative mRNA expression (against Rplp0) of the OST-tagged WT, K97R, K108R, K311R and K486R derivatives (n=3 independent biological replicates) of Supplementary Fig. 5c. c) Nuclear abnormalities quantification in MEFs expressing the indicated lamin A derivatives. Statistical comparisons against WT lamin A using ordinary one-way ANOVA on n=3 independent biological replicates. d) Relative mRNA expression (against Rplp0) of the OST-tagged WT, K311R and K311Q lamin A derivatives (n=3 independent biological replicates) of Fig. 3c. e) Immunoblotting on the indicated subcellular fractions of MEFs expressing WT, K311R or K311Q lamin A. f) Left; Immunoblotting on whole cell extracts of immortalized Lmna KO MEFs expressing WT, K311R or K311Q OST-lamin A. Right; Nuclear abnormalities quantification. Two-tailed unpaired t-test on n=3 independent biological replicates. g) Representative immunostaining for OST-tag (green) and lamin B1 (red) on control and Mof KO K319R/Q MEFs. h) Nuclear abnormalities quantification of Supplementary Fig. 5g. Ordinary one-way ANOVA on n=3 independent biological replicates. i) Representative immunostaining for WT/K59R GFP-lamin B2 (green) and lamin B2 (red). j) Nuclear abnormalities quantification in WT/K59R lamin B2 MEFs. Two-tailed unpaired t-test on n=3 independent biological replicates. k) TEM analysis of paracrystalline arrays. K311R and WT lamin C show similar striped patterns and structural organization of the paracrystalline arrays. Scale bar, 20 nm. l) Quantification of total RNA Pol II and RNA Pol II-S2P. Nonparametric one-way ANOVA followed by a Kruskal-Wallis test. n numbers are stated in the graph and represent nuclear structures counted. m) left:Representative immunostaining of NUP153 in WT and K311R OST-lamin A MEFs. Right: Quantification of NUP153 in main nuclei and nuclear abnormalities of K311R MEFs. Two-tailed nonparametric Mann-Whitney test. n numbers are stated in the graph and represent nuclear structures counted. n) H4K16ac intensities on MEFs expressing WT, K311Ror K311Q OST-lamin A. Means were compared by a Kruskal-Wallis test on n=3 independent biological replicates. The lower and upper hinges correspond to the first and third quartiles. The whiskers extends from the hinge to the largest or smallest values respectively, no further than 1.5 * IQR (inter-quartile range) from the hinge. The middle line represents the median (the middle value of the dataset). Data beyond the end of the whiskers are plotted individually. Data shown are the mean ± s.e.m for a, b, c, d, f, h, j, l, m. For g, i, m dsDNA was counterstained using DAPI and scale bars represent 5μm. Uncropped blots of a, e, f can be found in Supplementary Fig. 9.
Supplementary Figure 6 Sensitivity of lamin A/C acetylation to HDAC inhibition and representative genomic DNA read-depth plots for control, Mof KO and K311R lamin A MEFs.
a) Immunoblotting on whole cell extracts of control and Mof KO upon panobinostat (200nM) and bufexamac (5μM) treatment. b) Quantification of nuclear abnormalities in control and Mof KO MEFs treated with the indicated panobinostat and bufexamac concentrations; Ordinary one-way ANOVA on n=3 independent biological replicates. c–e) Normalized read-depth along the genome for typical WT control primary MEFs. Three different cells are shown. f) Normalized read-depth along the genome for a Mof KO MEF exhibiting chromothripsis, shown in Fig. 4i. g) Normalized read-depth along the genome for a K311R lamin A MEF exhibiting chromothripsis, shown in Fig. 4j. h) Normalized read-depth plots and CBS segmentation results for selected Mof KO cells. CBS segments were further computationally processed (Methods) to generate high confidence segments used in Fig. 4e, f. i) Immunoblotting on subcellular fractions of MEFs expressing the indicated OST-lamin A derivatives. j) Normalized read-depth plots and CBS segmentation results for selected K311R lamin A MEFs. CBS segments were further computationally processed (Methods), to generate high confidence segments used in Fig. 4g, h. Uncropped blots of a, i can be found in Supplementary Fig. 9.
Supplementary Figure 7 In depth characterization of the nuclear abnormalities epigenetic landscape.
a) Quantification of H3K27me3 and H3K27ac normalized to DAPI in the main nuclei, nuclear blebs and micronuclei of Mof KO (4 days); Nonparametric one-way ANOVA followed by a Kruskal-Wallis test. b) Representative immunostaining of H4K8ac (green) in Mof and Kansl2 KO (4 days) MEFs. c) Quantification of H4K8ac/DAPI in the main nuclei, nuclear blebs and micronuclei of Mof and Kansl2 KO (4 days); Nonparametric one-way ANOVA followed by a Kruskal-Wallis test. d) Representative immunostaining of K311R OST-lamin A MEFs for ATAC-see (white) and H3K27me3 (red). Arrows indicate nuclear abnormalities enriched for H3K27me3. e) Quantification of DAPI/dsDNA in the main nuclei and micronuclei of K311R lamin A MEFs; Two-tailed nonparametric Mann–Whitney test. f) Quantification of histone H3/DAPI in the indicated nuclear structures of Mof KO MEFs; Nonparametric one-way ANOVA followed by a Kruskal-Wallis test. g) Quantification of DAPI in the indicated nuclear structures of K311R lamin A MEFs; Nonparametric one-way ANOVA followed by a Kruskal-Wallis test. h) Representative immunostainings upon incubation with blocking peptides indicating specificity of the detected signal. Arrows indicate nuclear abnormalities. i) Quantification of bulk ATAC-see in MEFs expressing the indicated lamin A derivatives. Dots represent mean signal intensity. Nonparametric one-way ANOVA followed by a Kruskal-Wallis test. j) Immunoblotting on whole cell extracts of WT and K311R lamin A MEFs. Uncropped blots in Supplementary Fig. 9. k) Quantification of nuclear abnormalities on control and Mof KO (2 days) MEFs in the presence of the indicated GSK343 concentrations; Ordinary one-way ANOVA on n=3 independent biological replicates. l) Quantification of H3K27me3/DAPI in the main nuclei and nuclear abnormalities of control and Mof KO (2 days) MEFs in the presence of the indicated GSK343 concentrations; Nonparametric one-way ANOVA followed by a Kruskal-Wallis test. m) Quantification of Pol II-S2P in the main nuclei and nuclear abnormalities of control and Mof KO (2 days) MEFs in the presence of the indicated GSK343 concentrations; Nonparametric one-way ANOVA followed by a Kruskal-Wallis test. The data shown are the mean ± s.e.m. for a, c, e, f, g, k, l, m. dsDNA was counterstained using DAPI and scale bars of 5μm where applied to b, d, h. For a, c, e, f, g, l, m, n numbers are stated in the graphs and represent nuclear structures counted.
Supplementary Figure 8 Defective nuclear lamina architecture and impaired nuclear mechanics upon loss of MOF and lamin A/C acetylation.
a) Representative electron tomographic slices showing preserved nuclear structure in Mof control and WT lamin A MEFs. N; nucleus. b) Quantification of electron tomographic micronuclei heterochromatin density in Mof KO and K311R lamin A MEFs. Data shown are the mean ± s.e.m. n=15 nuclear structures counted. c) Quantification of electron tomographic submembrane (120 nm) heterochromatin density in the 1-μm-vicinity of fuzzy nuclear lamina sites; Two-tailed nonparametric Mann–Whitney test. The data shown are the mean ± s.e.m. n numbers are stated in the panel and represent single cells counted. d) Quantification of electron tomographic global nuclear heterochromatin density measured; Two-tailed nonparametric Mann–Whitney test. Data shown are the mean ± s.e.m. n numbers are stated in the panel and represent single cells counted. e) Relative Emerald-lamin A FRAP over time in Lmna KO MEFs overexpressing WT or K311R lamin A. Two-tailed nonparametric Mann–Whitney test for n=45 WT and n=50 K311R cells. f) Immunoblotting on whole cell extracts of MEFs upon Mof or Kansl2 KO (4 days). Uncropped blots in Supplementary Fig. 9. g) Nuclear and cyto-indentations are performed in triplicate for each cell (left image) and for each isolated nucleus (right image). Pink crosses indicate sites of nanoindentation. Scale bar; 20μm. h) Schematics of the nanoindentation experiment performed and representative force distance curve used to calculate the effective Young’s Modulus. Fit of the indentation curve used to obtain the Young’s Modulus shown in black. i) Cytoindentation quantification on intact control and Mof KO (4 days) MEFs; Two-tailed nonparametric Mann–Whitney test. Data shown are the mean ± s.e.m. n numbers are stated in the panel and represent single cells counted. j) Nanoindentation nuclei stiffness quantification in intact control and Mof KO (4 days) MEFs; Two-tailed nonparametric Mann–Whitney test. Data shown are the mean ± s.e.m. n numbers are stated in the panel and represent single cells counted. k) Nanoindentation stiffness quantification on freshly isolated control and Mof KO (4 days) MEFs nuclei; Two-tailed nonparametric Mann–Whitney test. Data shown are the mean ± s.e.m. n numbers are stated in the panel and represent single nuclei counted.
Supplementary Figure 9 Unprocessed scans of immunoblots.
Red boxes show cropped regions. Molecular weights (kDa) are indicated.
Supplementary information
Supplementary Information
Supplementary Figures 1–9 and Supplementary Video and Table titles/legends.
Supplementary Table 1
MEF_input—Proteome deregulation following Mof KO. Two-tailed t-test applied.
Supplementary Table 2
ESC_input—Proteome deregulation following Mof KO. Two-tailed t-test applied.
Supplementary Table 3
MEF_Acetylome deregulation following Mof KO. Two-tailed t-test applied.
Supplementary Table 4
ESC_Acetylome deregulation following Mof KO. Two-tailed t-test applied.
Supplementary Table 5
Gene Ontology analysis.
Supplementary Table 6
Histone acetyl (lysine) sites multiplicity comparison for Mof KO.
Supplementary Table 7
Statistics source data.
Supplementary Video 1
Live-cell imaging of a control MEF cell transduced with GFP–NLS (green). Time intervals of 15 min are shown.
Supplementary Video 2
Live-cell imaging of a Mof KO MEF transduced with GFP–NLS (green). Time intervals of 24 min are shown.
Supplementary Video 3
Live-cell imaging of a Kansl2 KO MEF transduced with GFP–NLS (green). Time intervals of 15 min are shown.
Supplementary Video 4
Live-cell imaging of a Mof KO MEF transduced with GFP–H2B (green). Time intervals of 15 min are shown.
Supplementary Video 5
Live-cell imaging of a Mof KO MEF transduced with GFP–NLS (green). Time intervals of 2 min are shown. Rupture of the main nucleus is observed.
Supplementary Video 6
Live-cell imaging of a Mof KO MEF transduced with GFP–NLS (green). Time intervals of 2 min are shown. Rupture of the main nucleus is observed.
Supplementary Video 7
Live-cell imaging of a Mof KO MEF transduced with GFP–NLS (green). Time intervals of 2 min are shown. Rupture of the nuclear abnormality followed by main nuclei rupture is observed.
Supplementary Video 8
Live-cell imaging of a Mof KO MEF transduced with GFP–NLS (green). Time intervals of 2 min are shown. Continuous rupture of the nuclear abnormality is observed.
Supplementary Video 9
Live-cell imaging of a WT lamin A MEF transduced with GFP–NLS (green). Time intervals of 15 min are shown.
Supplementary Video 10
Live-cell imaging of a K311R lamin A K311R MEF transduced with GFP–NLS (green). Time intervals of 15 min are shown.
Supplementary Video 11
Live-cell imaging of a Mof KO MEF transduced with GFP-H3K27me3-mintbody (green). Time intervals of 15 min are shown.
Supplementary Video 12
3D reconstruction of a Mof KO MEF (4 d) where the main nuclei (dark blue) with NERDI as well as the MN (light blue) are visible.
Supplementary Note
Mof KO SILAC proteome and acetylome data analysis.
Rights and permissions
About this article
Cite this article
Karoutas, A., Szymanski, W., Rausch, T. et al. The NSL complex maintains nuclear architecture stability via lamin A/C acetylation. Nat Cell Biol 21, 1248–1260 (2019). https://doi.org/10.1038/s41556-019-0397-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0397-z
This article is cited by
-
Lamin B1 curtails early human papillomavirus infection by safeguarding nuclear compartmentalization and autophagic capacity
Cellular and Molecular Life Sciences (2024)
-
Heritable transcriptional defects from aberrations of nuclear architecture
Nature (2023)
-
Disrupting the phase separation of KAT8–IRF1 diminishes PD-L1 expression and promotes antitumor immunity
Nature Cancer (2023)
-
COX17 acetylation via MOF–KANSL complex promotes mitochondrial integrity and function
Nature Metabolism (2023)
-
Deformation of the nucleus by TGFβ1 via the remodeling of nuclear envelope and histone isoforms
Epigenetics & Chromatin (2022)