Abstract
Ribosomes are multicomponent molecular machines that synthesize all of the proteins of living cells. Most of the genes that encode the protein components of ribosomes are therefore essential. A reduction in gene dosage is often viable albeit deleterious and is associated with human syndromes, which are collectively known as ribosomopathies1,2,3. The cell biological basis of these pathologies has remained unclear. Here, we model human ribosomopathies in Drosophila and find widespread apoptosis and cellular stress in the resulting animals. This is not caused by insufficient protein synthesis, as reasonably expected. Instead, ribosomal protein deficiency elicits proteotoxic stress, which we suggest is caused by the accumulation of misfolded proteins that overwhelm the protein degradation machinery. We find that dampening the integrated stress response4 or autophagy increases the harm inflicted by ribosomal protein deficiency, suggesting that these activities could be cytoprotective. Inhibition of TOR activity—which decreases ribosomal protein production, slows down protein synthesis and stimulates autophagy5—reduces proteotoxic stress in our ribosomopathy model. Interventions that stimulate autophagy, combined with means of boosting protein quality control, could form the basis of a therapeutic strategy for this class of diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary Information. The mass spectrometry dataset is available at ProteomeXchange under the identifier PXD023021. Accession numbers and names for the proteins identified by mass spectrometry are available at UniProt (https://www.uniprot.org/) and FlyBase (http://flybase.org/). Source data are provided with this paper.
References
Farley-Barnes, K. I., Ogawa, L. M. & Baserga, S. J. Ribosomopathies: old concepts, new controversies. Trends Genet. 35, 754–767 (2019).
Mills, E. W. & Green, R. Ribosomopathies: there’s strength in numbers. Science 358, eaan2755 (2017).
Aspesi, A. & Ellis, S. R. Rare ribosomopathies: insights into mechanisms of cancer. Nat. Rev. Cancer 19, 228–238 (2019).
Harding, H. P. et al. An integrated stress response regulates amino acid metabolism and resistance to oxidative stress. Mol. Cell 11, 619–633 (2003).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 168, 960–976 (2017).
Klauck, S. M. et al. Mutations in the ribosomal protein gene RPL10 suggest a novel modulating disease mechanism for autism. Mol. Psychiatry 11, 1073–1084 (2006).
Brooks, S. S. et al. A novel ribosomopathy caused by dysfunction of RPL10 disrupts neurodevelopment and causes X-Linked microcephaly in humans. Genetics 198, 723–733 (2014).
Hetman, M. & Slomnicki, L. P. Ribosomal biogenesis as an emerging target of neurodevelopmental pathologies. J. Neurochem. 148, 325–347 (2019).
Paolini, N. A. et al. A ribosomopathy reveals decoding defective ribosomes driving human dysmorphism. Am. J. Hum. Genet. 100, 506–522 (2017).
Alsop, R. J. et al. Structural abnormalities in the hair of a patient with a novel ribosomopathy. PLoS ONE 11, e0149619 (2016).
Marygold, S. J. et al. The ribosomal protein genes and minute loci of Drosophila melanogaster. Genome Biol. 8, R216 (2007).
Lambertsson, A. The Minute genes in Drosophila and their molecular functions. Adv. Genet. 38, 69–134 (1998).
Morata, G. & Ripoll, P. Minutes: mutants of Drosophila autonomously affecting cell division rate. Dev. Biol. 42, 211–221 (1975).
Moreno, E., Basler, K. & Morata, G. Cells compete for decapentaplegic survival factor to prevent apoptosis in Drosophila wing development. Nature 416, 755–759 (2002).
Johnston, L. A. Competitive interactions between cells: death, growth, and geography. Science 324, 1679–1682 (2009).
Milán, M., Campuzano, S. & García-Bellido, A. Developmental parameters of cell death in the wing disc of Drosophila. Proc. Natl Acad. Sci. USA 94, 5691–5696 (1997).
Coelho, C. M. A. Growth and cell survival are unevenly impaired in pixie mutant wing discs. Development 132, 5411–5424 (2005).
Ulirsch, J. C. et al. The genetic landscape of Diamond-Blackfan anemia. Am. J. Hum. Genet. 103, 930–947 (2018).
Kucinski, I., Dinan, M., Kolahgar, G. & Piddini, E. Chronic activation of JNK JAK/STAT and oxidative stress signalling causes the loser cell status. Nat. Commun. 8, 136 (2017).
Lee, C.-H. et al. A regulatory response to ribosomal protein mutations controls translation, growth, and cell competition. Dev. Cell 46, 456–469 (2018).
Akdemir, F., Christich, A., Sogame, N., Chapo, J. & Abrams, J. M. p53 directs focused genomic responses in Drosophila. Oncogene 26, 5184–5193 (2007).
Baillon, L., Germani, F., Rockel, C., Hilchenbach, J. & Basler, K. Xrp1 is a transcription factor required for cell competition-driven elimination of loser cells. Sci. Rep. 8, 17712 (2018).
Ji, Z. et al. Drosophila RpS12 controls translation, growth, and cell competition through Xrp1. PLOS Genet. 15, e1008513 (2019).
Danilova, N. & Gazda, H. T. Ribosomopathies: how a common root can cause a tree of pathologies. Dis. Model. Mech. 8, 1013–1026 (2015).
Wartlick, O. et al. Dynamics of dpp signaling and proliferation control. Science 331, 1154–1159 (2011).
Nienhaus, U., Aegerter-Wilmsen, T. & Aegerter, C. M. In-vivo imaging of the Drosophila wing imaginal disc over time: novel insights on growth and boundary formation. PLoS ONE 7, e47594 (2012).
Liu, J., Xu, Y., Stoleru, D. & Salic, A. Imaging protein synthesis in cells and tissues with an alkyne analog of puromycin. Proc. Natl Acad. Sci. USA 109, 413–418 (2012).
Lacsina, J. R. et al. Premature translational termination products are rapidly degraded substrates for MHC Class I presentation. PLoS ONE 7, e51968 (2012).
Mediani, L. et al. Defective ribosomal products challenge nuclear function by impairing nuclear condensate dynamics and immobilizing ubiquitin. EMBO J. 38, e101341 (2019).
Wenger, T. et al. Autophagy inhibition promotes defective neosynthesized proteins storage in ALIS, and induces redirection toward proteasome processing and MHCI-restricted presentation. Autophagy 8, 350–363 (2012).
Seguin, S. J. et al. Inhibition of autophagy, lysosome and VCP function impairs stress granule assembly. Cell Death Differ. 21, 1838–1851 (2014).
Martin, D. D. O., Ladha, S., Ehrnhoefer, D. E. & Hayden, M. R. Autophagy in Huntington disease and huntingtin in autophagy. Trends Neurosci. 38, 26–35 (2015).
Serpionov, G. V., Alexandrov, A. I., Antonenko, Y. N. & Ter-Avanesyan, M. D. A protein polymerization cascade mediates toxicity of non-pathological human huntingtin in yeast. Sci. Rep. 5, 18407 (2015).
Busch, A. et al. Mutant huntingtin promotes the fibrillogenesis of wild-type huntingtin: a potential mechanism for loss of huntingtin function in Huntington’s disease. J. Biol. Chem. 278, 41452–41461 (2003).
Bjørkøy, G. et al. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J. Cell Biol. 171, 603–614 (2005).
Nezis, I. P. et al. Autophagic degradation of dBruce controls DNA fragmentation in nurse cells during late Drosophila melanogaster oogenesis. J. Cell Biol. 190, 523–531 (2010).
Pederson, T. Ribosomal protein mutations in Diamond‐Blackfan anemia: might they operate upstream from protein synthesis? FASEB J. 21, 3442–3445 (2007).
Shi, Z. et al. Heterogeneous ribosomes preferentially translate distinct subpools of mRNAs genome-wide. Mol. Cell 67, 71–83 (2017).
Grentzmann, G., Ingram, J. A., Kelly, P. J., Gesteland, R. F. & Atkins, J. F. A dual-luciferase reporter system for studying recoding signals. RNA 4, 479–486 (1998).
Manuvakhova, M., Keeling, K. & Bedwell, D. M. Aminoglycoside antibiotics mediate context-dependent suppression of termination codons in a mammalian translation system. RNA 6, 1044–1055 (2000).
Albert, B. et al. A ribosome assembly stress response regulates transcription to maintain proteome homeostasis. eLife 8, e45002 (2019).
Tye, B. W. et al. Proteotoxicity from aberrant ribosome biogenesis compromises cell fitness. eLife 8, e43002 (2019).
Nagaraj, N. et al. Deep proteome and transcriptome mapping of a human cancer cell line. Mol. Syst. Biol. 7, 548 (2011).
Pelletier, J., Thomas, G. & Volarević, S. Ribosome biogenesis in cancer: new players and therapeutic avenues. Nat. Rev. Cancer 18, 51–63 (2017).
Wiśniewski, J. R., Hein, M. Y., Cox, J. & Mann, M. A ‘proteomic ruler’ for protein copy number and concentration estimation without spike-in standards. Mol. Cell. Proteom. 13, 3497–3506 (2014).
An, H. & Harper, J. W. Ribosome abundance control via the ubiquitin–proteasome system and autophagy. J. Mol. Biol. https://doi.org/10.1016/j.jmb.2019.06.001 (2019).
Pillet, B., Mitterer, V., Kressler, D. & Pertschy, B. Hold on to your friends: dedicated chaperones of ribosomal proteins. BioEssays 39, e201600153 (2017).
Sung, M. K. et al. A conserved quality-control pathway that mediates degradation of unassembled ribosomal proteins. eLife 5, e19105 (2016).
Blanco, J., Cooper, J. C. & Baker, N. E. Roles of C/EBP class bZip proteins in the growth and cell competition of Rp (‘Minute’) mutants in Drosophila. eLife 9, e50535 (2020).
Malzer, E. et al. Coordinate regulation of eIF2α phosphorylation by PPP1R15 and GCN2 is required during Drosophila development. J. Cell Sci. 126, 1406–1415 (2013).
King, M. A. et al. Rapamycin inhibits polyglutamine aggregation independently of autophagy by reducing protein synthesis. Mol. Pharmacol. 73, 1052–1063 (2008).
Conn, C. S. & Qian, S.-B. Nutrient signaling in protein homeostasis: an increase in quantity at the expense of quality. Sci. Signal. 6, ra24 (2013).
Xie, J. et al. Regulation of the elongation phase of protein synthesis enhances translation accuracy and modulates lifespan. Curr. Biol. 29, 737–749 (2019).
Baumgartner, M., Dinan, M. P., Langton, P. F., Kucinski, I. & Piddini, E. Proteotoxic stress is a driver of the loser status and of cell competition. Nat. Cell Biol. https://doi.org/10.1038/s41556-020-00626-1 (2020).
Bové, J., Martínez-Vicente, M. & Vila, M. Fighting neurodegeneration with rapamycin: mechanistic insights. Nat. Rev. Neurosci. 12, 437–452 (2011).
Doulatov, S. et al. Drug discovery for Diamond-Blackfan anemia using reprogrammed hematopoietic progenitors. Sci. Transl. Med. 9, eaah5645 (2017).
Cortez, L. & Sim, V. The therapeutic potential of chemical chaperones in protein folding diseases. Prion 8, 197–202 (2014).
Mayor-Ruiz, C. et al. Rational discovery of molecular glue degraders via scalable chemical profiling. Nat. Chem. Biol. 16, 1199–1207 (2020).
Poernbacher, I. et al. Lessons in genome engineering: opportunities, tools and pitfalls. Preprint at bioRxiv https://doi.org/10.1101/710871 (2019).
Kondo, S. & Ueda, R. Highly improved gene targeting by germline-specific Cas9 expression in Drosophila. Genetics 195, 715–721 (2013).
Baena-Lopez, L. A., Alexandre, C., Mitchell, A., Pasakarnis, L. & Vincent, J. P. Accelerated homologous recombination and subsequent genome modification in Drosophila. Development 140, 4818–4825 (2013).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Germani, F., Bergantinos, C. & Johnston, L. A. Mosaic analysis in Drosophila. Genetics 208, 473–490 (2018).
Nezis, I. P. et al. Ref(2)P, the Drosophila melanogaster homologue of mammalian p62, is required for the formation of protein aggregates in adult brain. J. Cell Biol. 180, 1065–1071 (2008).
Acknowledgements
We thank T. E. Rusten (University of Oslo) for the gift of anti-p62 antibodies and F. Zhang (Broad Institute of MIT and Harvard) for the PX459 vector. We also acknowledge the staff at the Developmental Studies Hybridoma Bank, created by the NICHD of the NIH and maintained at the University of Iowa for the provision of antibodies. Drosophila stocks obtained from the Bloomington Drosophila Stock Center (NIH P40OD018537) were used in this study. We also thank M. Cockman, I. McGough and P. Ratcliffe (all at the Crick Institute), as well as M. Pieterse (Mill, The Netherlands) for discussions. This work was supported by a Wellcome Trust Investigator award (no. 206341/Z/17/Z to J.P.V.) and the Francis Crick Institute, which receives its core funding from Cancer Research UK (no. FC001204), the UK Medical Research Council (no. FC001204) and the Wellcome Trust (no. FC001204).
Author information
Authors and Affiliations
Contributions
This project was conceived by C.R.-A., H.N. and J.-P.V.; H.N. created the RPS23R67K strain, as well as the RPS26attP-KO strain, which was used as the starting point for generating RPS26cKO by C.A. and C.R.-A.; H.N. also performed the developmental timing measurements. D.J.H. generated the RPS23R67K HEK293 cells. C.A. and C.R.-A. generated the translation fidelity reporter. J.K. and A.P.S. generated and analysed the mass spectrometry data. The manuscript was written by C.R.-A. and J.-P.V.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phenotypes of Minute heterozygotes.
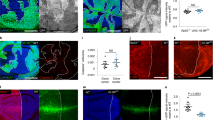
(a, b) Scutellar region of control and RPS23R67K/+ flies showing the short-bristle phenotype that characterises Minute heterozygotes. c, Cumulative distribution of pupariation time for control (n = 143 larvae) and RPS23R67K/+ (n = 134 larvae). Error bars represent standard deviation (d) Control mosaic imaginal disc harbouring wildtype clones (2X GFP) and their wild type twin clones (absence of GFP), induced by heat shock-mediated expression of FLP (hs-FLP). Note the low number of Dcp1-positive cells (red and grey). e, Mosaic imaginal discs harbouring wild type clones (2X GFP) in a RPS23R67K/+ background (1X GFP), also induced with hs-FLP. Here the twin clones (RPS23R67K/R67K) are rapidly eliminated and the wild type cells outcompete the RPS23R67K/+ cells, which undergo a high rate of apoptosis (Dcp1, red and grey). (f, g) JNK signalling (indicated by expression of the TRE-GFP reporter and apoptosis (Dcp1) in RPS23R67K/+ are fully supressed by a wild type copy of RPS23 from a genomic duplication (g). (h–j) JNK signalling and apoptosis in a panel of heterozygous Minute mutants (RPL5, RPL14, and RPS13). Scale bars represent 50 µm. Genotypes for each figure panel are available in Supplementary Table 1. Source data is available for this figure.
Extended Data Fig. 2 Xrp1 is required for activation of apoptosis in RPS23R67K/+.
a, Imaginal disc of a RPS23R67K/+ larva expressing an RNAi transgene against Xrp1 in the anterior compartment (marked with anti-Ci). The number of Dcp1-positive cells is lower in the anterior than in the posterior compartment where Xrp1 activity is unaffected. Scale bars represent 50 µm. Genotypes for each figure panel are available in Supplementary Table 1. Experiments were repeated independently three times with similar results.
Extended Data Fig. 3 Manipulation of Rheb activity affects OPP incorporation in wild type and RPS23R67K/ + imaginal discs.
(a–d) Wing imaginal discs (wild type and RPS23R67K/+, as indicated) overexpressing Rheb or RhebRNAi in the anterior compartment (left hand side of the disc). The discs were explanted and incubated for a 15 min in 1 µM OPP before staining for puromycilated peptides (grey scale). Rheb overexpression stimulated OPP incorporation in both genotypes, while RhebRNAi had the opposite effect. Scale bars represent 50 µm. Genotypes for each figure panel are available in Supplementary Table 1. Experiments were repeated independently three times with similar results.
Extended Data Fig. 4 RPS23R67K/ + imaginal discs develop tumours upon inhibition of apoptosis.
(a,b) Wing disc from a RPS23R67K/+ larva expressing P35, a baculovirus-derived inhibitor of effector caspases in the pouch (under the control of rotund-GAL4). Note the overgrowth characterised by epithelial folds and ectopic Wingless expression (green), shown in grey scale at higher magnification in b. Formation of these tumours shows that RP-deficient cells are not inherently incapable of growth. Tumour formation may be relevant to the increase cancer risk associated with human ribosomopathies as well as to the observation that ribosomal protein genes are frequently deleted in human cancers, often in concert with the loss of TP533. Scale bars represent 50 µm. Genotypes for each figure panel are available in Supplementary Table 1. Experiments were repeated independently three times with similar results.
Extended Data Fig. 5 A toxic form of Huntingtin (HTT96Q) triggers apoptosis.
(a,b) Expression of HTT96Q throughout the pouch (with rotund-gal4) triggers an increased rate of apoptosis relative to that seen with GFP expression, which is expected to be innocuous. c, Quantification of Dcp1 coverage in the two genotypes shown in panels a and b (n = 5 discs per genotype). Error bars denote standard deviation. For statistical analysis, a two-tailed unpaired t-test was carried out. **P = 4.11E-03. Scale bars represent 50 µm. Genotypes for each figure panel are available in Supplementary Table 1. Source data is available for this figure.
Extended Data Fig. 6 Accumulation of p62 and P-eIF2α in RPS26KO/+.
(a, b) RPS26cKO was inactivated (and tubulin-mCherry deleted) by crossing to hedgehog-GAL4, UAS-FLP. In the resulting RPS26KO/+ posterior compartment, immunoreactivity against p62 and P-eIF2α was higher than in the control anterior compartment. Scale bars represent 50 µm. Genotypes for each figure panel are available in Supplementary Table 1. Experiments were repeated independently three times with similar results.
Extended Data Fig. 7 RP deficiency alters the activity of an autophagy reporte.
a, Cartoon showing progression through autophagy as monitored by the GFP-mCherry-Atg8a reporter. Yellow indicates the simultaneous presence of GFP and mCherry in the phagophore and autophagosome. Autolysosomes only retain the red colour because of the drop in pH, which quenches GFP fluorescence. (b, c) Fluorescence from GFP-mCherry-Atg8a, expressed with tubulin-GAL4 in wild type or RPS23R67K/+. Single fluorescence channels are also shown in grey. Scale bars represent 50 µm. Genotypes for each figure panel are available in Supplementary Table 1. Experiments were repeated independently three times with similar results.
Extended Data Fig. 8 Translation fidelity is unaffected in RPS23R67K/+.
a, The stop codon readthrough reporter comprises 10X Upstream Activator Sequences (UAS), which confers GAL4 responsiveness, the 5’ UTR from Syn21, the coding region of Firefly luciferase (Fluc), a STOP codon (UGAC), a flexible linker, the coding region of Nanoluciferase (Nluc), and the 3’UTR from p10. b, Quantification of the Nluc/Fluc ratio, measured from whole larvae lysates and normalised to that in control larvae. Statistical analysis: 4 replicates for each condition. Error bars denote standard deviation. A two-tailed unpaired t-test was carried out. P>0.05, no significant increase was seen in RPS23R67K/+ larvae. Genotypes for each figure panel are available in Supplementary Table 1. Source data is available for this figure.
Extended Data Fig. 9 Validating the effect of GADD34 overexpression.
a, P-eIF2α immunoreactivity in a wild type imaginal disc. b, This is reduced by GADD34 overexpression (GADD34OE) driven by nubbin-GAL4. (c) P-eIF2α immunoreactivity is similarly decreased in RPS23R67K/+ larvae overexpressing GADD34. d, Schematic representation of the domain where GADD34 was overexpressed. (e,f) GADD34 overexpression causes a mild but significant increase in wing size in otherwise wild type flies but not in RPS23R67K heterozygotes. Note that the wing of RPS23R67K heterozygotes is smaller than that of wildtype. (g-i) GADD34 overexpression exacerbates the formation of HTT25Q punctae in RPS23R67K heterozygotes. Statistics: error bars denote standard deviation. In f, n = 12 adult wings for each genotype. In i, from left to right, n = 10 and 8 discs. A two-tailed unpaired t-test was carried out. P-values in f, from top to bottom: 2.48E-01, 1.31E-07 and 7.09E-06. P-value in i, 9.65E-04. Scale bars represent 50 µm. Genotypes for each figure panel are available in Supplementary Table 1. Source data is available for this.
Extended Data Fig. 10 Effect of proteasome inhibition on the rate of apoptosis in RP-deficient tissues.
a, Extent of apoptosis (coverage of Dcp1 immuno-reactivity) in the pouch of discs of genotypes indicated. Rpt6RNAi denotes rotund-gal4-driven expression of a Rpt6RNAi transgene.. This particular Rpt6RNAi transgene had only a minor effect on apoptosis in wildtype tissue. Expression of this RNAi transgene did not enhance the rate of apoptosis in RPS23R67K heterozygotes. b, The effect of stronger proteasome knockdown (with Rpn2RNAi) on the rate of apoptosis in RP-deficient tissue could not be assessed because it triggered extensive apoptosis in otherwise wild type imaginal discs. Statistics: error bars denote standard deviation. n = 9 discs for each genotype. A two-tailed unpaired t-test was carried out. P > 0.05, no significant difference was seen. Genotypes for each figure panel are available in Supplementary Table 1. Source data is available for this figure.
Supplementary information
Supplementary Tables 1 and 2
Supplementary Table 1: genotypes analysed. Supplementary Table 2: proteins highlighted in Fig. 3.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Recasens-Alvarez, C., Alexandre, C., Kirkpatrick, J. et al. Ribosomopathy-associated mutations cause proteotoxic stress that is alleviated by TOR inhibition. Nat Cell Biol 23, 127–135 (2021). https://doi.org/10.1038/s41556-020-00626-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-00626-1
This article is cited by
-
Cell competition in development, homeostasis and cancer
Nature Reviews Molecular Cell Biology (2023)
-
Loss of Paip1 causes translation reduction and induces apoptotic cell death through ISR activation and Xrp1
Cell Death Discovery (2023)
-
Nacα protects the larval fat body from cell death by maintaining cellular proteostasis in Drosophila
Nature Communications (2023)
-
To not love thy neighbor: mechanisms of cell competition in stem cells and beyond
Cell Death & Differentiation (2023)
-
The PECAn image and statistical analysis pipeline identifies Minute cell competition genes and features
Nature Communications (2023)