Abstract
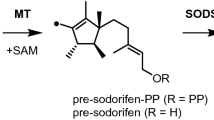
Terpenes constitute the largest class of natural products. Their skeletons are formed by terpene cyclases (TCs) from acyclic oligoprenyl diphosphates through sophisticated enzymatic conversions. These enzyme reactions start with substrate ionization through diphosphate abstraction, followed by a cascade reaction via cationic intermediates. Based on isotopic-labelling experiments in combination with a computational study, the cyclization mechanism for sodorifen, a highly methylated sesquiterpene from the soil bacterium Serratia plymuthica, was resolved. A peculiar problem in its biosynthesis lies in the formation of several methyl groups from chain methylene carbons. The underlying mechanism involves a methyltransferase-mediated cyclization and unprecedented ring contraction with carbon extrusion from the chain to form a methyl group. A terpene cyclase subsequently catalyses a fragmentation into two reactive intermediates, followed by hydrogen transfers between them and recombination of the fragments by [4 + 3] cycloaddition. This study solves the intricate mechanistic problem of extra methyl group formation in sodorifen biosynthesis.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The authors declare that the main data supporting the findings of this study are available within the Article, Supplementary Videos, Supplementary Data, and Supplementary Information. Crystallographic data for the structure reported in this Article have been deposited at the Cambridge Crystallographic Data Centre, under deposition number CCDC 2213524. Copies of the data can be obtained free of charge via https://www.ccdc.cam.ac.uk/structures/. Data and plasmids described in this study can be obtained from the corresponding author on reasonable request. Source data are provided with this paper.
References
Ryu, C.-M. et al. Bacterial volatiles promote growth in arabidopsis. Proc. Natl Acad. Sci. USA 100, 4927–4932 (2003).
Lee, H. H., Molla, M. N., Cantor, C. R. & Collins, J. J. Bacterial charity work leads to population-wide resistance. Nature 467, 82–86 (2010).
Chernin, L. et al. Quorum-sensing quenching by rhizobacterial volatiles. Environ. Microbiol. Rep. 3, 698–704 (2011).
Vespermann, A., Kai, M. & Piechulla, B. Rhizobacterial volatiles affect the growth of fungi and Arabidopsis thaliana. Appl. Environ. Microbiol. 73, 5639–5641 (2007).
von Reuss, S. H., Kai, M., Piechulla, B. & Francke, W. Octamethylbicyclo[3.2.1]octadienes from the rhizobacterium Serratia odorifera. Angew. Chem. Int. Ed. 49, 2009–2010 (2010).
Domik, D., Magnus, N. & Piechulla, B. Analysis of a new cluster of genes involved in the synthesis of the unique volatile organic compound sodorifen of Serratia plymuthica 4Rx13. FEMS Microbiol. Lett. 363, fnw139 (2016).
Domik, D. et al. A terpene synthase is involved in the synthesis of the volatile organic compound sodorifen of Serratia plymuthica 4Rx13. Front. Microbiol. 7, 737 (2016).
von Reuss, S. et al. Sodorifen biosynthesis in the rhizobacterium Serratia plymuthica involves methylation and cyclization of MEP-derived farnesyl pyrophosphate by a SAM-dependent C-methyltransferase. J. Am. Chem. Soc. 140, 11855–11862 (2018).
Duell, E. R. et al. Direct pathway cloning of the sodorifen biosynthetic gene cluster and recombinant generation of its product in E. coli. Microb. Cell. Fact. 18, 32 (2019).
Rabe, P. et al. Conformational analysis, thermal rearrangement and EI-MS-fragmentation mechanism of (1(10)E,4E,6S,7R)-germacradien-6-ol by 13C-labeling experiments. Angew. Chem. Int. Ed. 54, 13448–13451 (2015).
Eustaquio, A. S., Pojer, F., Noel, J. P. & Moore, B. S. Discovery and characterization of a marine bacterial SAM-dependent chlorinase. Nat. Chem. Biol. 4, 69–74 (2008).
Rabe, P. et al. Terpene cyclases from social amoebae. Angew. Chem. Int. Ed. 55, 15420–15423 (2016).
Rabe, P. et al. Mechanistic investigantions on two bacterial diterpene cyclases: spiroviolene synthase and tsukubadiene synthase. Angew. Chem. Int. Ed. 56, 2776–2779 (2017).
Rinkel, J. & Dickschat, J. S. Addressing the chemistry of germacrene A by isotope labeling experiments. Org. Lett. 21, 2426–2429 (2019).
Rinkel, J., Lauterbach, L., Rabe, P. & Dickschat, J. S. Two diterpene synthases for spiroalbatene and cembrene A from Allokutzneria albata. Angew. Chem. Int. Ed. 57, 3238–3241 (2018).
Cornforth, J. W., Cornforth, R. H., Donninger, C. & Popjak, G. Studies on the biosynthesis of cholesterol XIX. Steric course of hydrogen eliminations and of C–C bond formations in squalene biosynthesis. Proc. R. Soc. London, Ser. B 163, 492–514 (1966).
Rinkel, J. et al. Mechanisms of the diterpene cyclases β-pinacene synthase from Dictyostelium discoideum and hydropyrene synthase from Streptomyces clavuligerus. Chem. Eur. J. 23, 10501–10505 (2017).
Reetz, M. T. Dytropic rearrangements, a new class of orbital-symmetry controlled reactions. Type I. Angew. Chem. Int. Ed. 11, 129–130 (1972).
Hugelshofer, C. L. & Magauer, T. Dyotropic rearrangements in natural product total synthesis and biosynthesis. Nat. Prod. Rep. 34, 228–234 (2017).
Gutierrez, O. & Tantillo, D. J. Analogies between synthetic and biosynthetic reactions in which [1,2]-alkyl shifts are combined with other events: dyotropic, Schmidt, and carbocation rearrangements. J. Org. Chem. 77, 8845–8850 (2012).
Li, H. & Dickschat, J. S. Isotopic labelling experiments and enzymatic preparation of iso-casbenes with casbene synthase from Ricinus communis. Org. Chem. Front. 9, 795–801 (2022).
Mitsuhashi, T., Rinkel, J., Okada, M., Abe, I. & Dickschat, J. S. Mechanistic characterization of two chimeric sesterterpene synthases from Penicillium. Chem. Eur. J. 23, 10053–10057 (2017).
Cornforth, J. W., Cornforth, R. H., Popjak, G. & Yengoyan, L. Studies on the biosynthesis of cholesterol XX. Steric course of decarboxylation of 5-pyrophosphomevalonate and of the carbon to carbon bond formation in the biosynthesis of farnesyl pyrophosphate. J. Biol. Chem. 241, 3970–3987 (1966).
Adamo, C. & Barone, V. Exchange functionals with improved long-range behavior and adiabatic connection methods without adjustable parameters: the mPW and mPW1PW models. J. Chem. Phys. 108, 664–675 (1998).
Matsuda, S. P. T., Wilson, W. K. & Xiong, Q. Mechanistic insights into triterpene synthesis from quantum mechanical calculations. Detection of systematic errors in B3LYP cyclization energies. Org. Biomol. Chem. 4, 530–543 (2006).
Hong, Y. J. & Tantillo, D. J. A maze of dyotropic rearrangements and triple shifts: carbocation rearrangements connecting stemarene, stemodene, betaerdene, aphidicolene, and scopadulanol. J. Org. Chem. 83, 3780–3793 (2018).
Quan, Z., Hou, A., Goldfuss, B. & Dickschat, J. S. Mechanism of the bifunctional multiple product sesterterpene synthase AcAS from Aspergillus calidoustus. Angew. Chem. Int. Ed. 61, e202117273 (2022).
Lemfack, M. C. et al. Reaction mechanism of the farnesyl pyrophosphate C-methyltransferase towards the biosynthesis of pre-sodorifen pyrophosphate by Serratia plymuthica 4Rx13. Sci. Rep. 11, 3182 (2021).
Burger, U., Delay, A. & Mazenod, F. 1,2,3,4,5-Pentamethyl-5-acetyl-cyclopentadien-1,3, ein ungewöhnliches Keton. Helv. Chim. Acta 57, 2106–2111 (1974).
Pace, V. et al. Efficient access to all-carbon quaternary and tertiary α-functionalized homoallyl-type aldehydes from ketones. Angew. Chem. Int. Ed. 56, 12677–12682 (2017).
Dess, D. B. & Martin, J. C. Readily accessible 12-I-51 oxidant for the conversion of primary and secondary alcohols to aldehydes and ketones. J. Org. Chem. 48, 4155–4156 (1983).
Horner, L., Hoffmann, H., Wippel, H. G. & Klahre, G. Phosphinoxyde als olefiniemgsreagenzien. Chem. Ber. 92, 2499–2505 (1959).
Wadsworth, W. S. & Emmons, W. D. The utility of phosphonate carbanions in olefin synthesis. J. Am. Chem. Soc. 83, 1733–1738 (1961).
Wise, M. L., Savage, T. J., Katahira, E. & Croteau, R. Monoterpene synthases from common sage (Salvia officinalis): cDNA Isolation, characterization, and functional expression of (+)-sabinene synthase, 1,8-cineol synthase, and (+)-bornyl diphosphate synthase. J. Biol. Chem. 273, 14891–14899 (1998).
Nakano, C., Kim, H.-K. & Ohnishi, Y. Identification of the first bacterial monoterpene cyclase, a 1,8-cineole synthase, that catalyzes the direct conversion of geranyl diphosphate. ChemBioChem 12, 1988–1991 (2011).
Jamieson, C. S., Ohashi, M., Liu, F., Tang, Y. & Houk, K. N. The expanding world of biosynthetic pericyclases: cooperation of experiment and theory for discovery. Nat. Prod. Rep. 36, 698–713 (2019).
Zhang, B. et al. Enzyme-catalysed [6 + 4] cycloadditions in the biosynthesis of natural products. Nature 568, 122–126 (2019).
Harmata, M. The (4 + 3)-cycloaddition reaction: simple allylic cations as dienophiles. Chem. Commun. 46, 8886–8903 (2010).
Giets, R. D. & Schiestl, R. H. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 31–34 (2007).
Dickschat, J. S., Pahirulzaman, K. A. K., Rabe, P. & Klapschinski, T. A. An improved technique for the rapid chemical characterisation of bacterial terpene cyclases. ChemBioChem 15, 810–814 (2014).
Grimme, S., Ehrlich, S. & Goerigk, L. Effect of the damping function in dispersion corrected density functional theory. J. Comp. Chem. 32, 1456–1465 (2011).
Frisch, M. J. et al. Gaussian 16, revision B.01 (Gaussian, 2016).
Lauterbach, L., Goldfuss, B. & Dickschat, J. S. Two diterpene synthases from Chryseobacterium: chryseodiene synthase and wanjudiene synthase. Angew. Chem. Int. Ed. 59, 11943–11947 (2020).
Grimme, S. Supramolecular binding thermodynamics by dispersion-corrected density functional theory. Chem. Eur. J. 18, 9955–9964 (2012).
Luchini, G., Alegre-Requena, J. V., Guan, Y., Funes-Ardoiz, I. & Paton, R. S. GoodVibes v.3.0.1 (2019); https://patonlab.com/robert-paton/
Acknowledgements
This work was funded by the Deutsche Forschungsgemeinschaft (DFG) (DI1536/11-1, project number 469042295) and supported by the computing centre of the University of Cologne (RRZK), providing CPU time on the DFG-funded supercomputer CHEOPS. We thank P. Garbeva (Wageningen) for strain S. plymuthica PRI-2C, B. Piechulla (Rostock) and S. von Reuss (Neuchatel) for discussions, and Andreas Schneider (Bonn) for HPLC separations.
Author information
Authors and Affiliations
Contributions
H.X. and L.L. performed syntheses and labelling experiments. B.G. performed density functional theory computations. G.S. solved the X-ray structure of (4S,5S)-12. J.S.D. designed and supervised research and wrote the manuscript with additions by all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–73, Tables 1–6, Schemes 1–8, and experimental and synthetic procedures.
Supplementary Data 1
Computed structure of P1* in Supplementary Fig. 42.
Supplementary Data 2
Computed structure of P1*-P3*-TS in Supplementary Fig. 42.
Supplementary Data 3
Computed structure of P3* in Supplementary Fig. 42.
Supplementary Data 4
Computed structure of P3*-TS in Supplementary Fig. 42.
Supplementary Data 5
Computed structure of P4* in Supplementary Fig. 42.
Supplementary Data 6
Computed structure of P4*-TS in Supplementary Fig. 42.
Supplementary Data 7
Computed structure of P5* in Supplementary Fig. 42.
Supplementary Data 8
Computed structure of P1*-NH3 in Supplementary Fig. 42.
Supplementary Data 9
Computed structure of P1*-NH3_TS in Supplementary Fig. 42.
Supplementary Data 10
Computed structure of P2*-NH3 in Supplementary Fig. 42.
Supplementary Data 11
Computed structure of P2*-NH3-TS in Supplementary Fig. 42.
Supplementary Data 12
Computed structure of P3*-NH3 in Supplementary Fig. 42.
Supplementary Data 13
Computed structure of S1 in Supplementary Fig. 43.
Supplementary Data 14
Computed structure of S1-TS in Supplementary Fig. 43.
Supplementary Data 15
Computed structure of S2 in Supplementary Fig. 43.
Supplementary Data 16
Computed structure of S2-TS in Supplementary Fig. 43.
Supplementary Data 17
Computed structure of S3 in Supplementary Fig. 43.
Supplementary Data 18
Computed structure of S3-TS in Supplementary Fig. 43.
Supplementary Data 19
Computed structure of S4 in Supplementary Fig. 43.
Supplementary Data 20
Computed structure of S4-TS in Supplementary Fig. 43.
Supplementary Data 21
Computed structure of S5a in Supplementary Fig. 43.
Supplementary Data 22
Computed structure of S5b in Supplementary Fig. 43.
Supplementary Data 23
Computed structure of S5-TS in Supplementary Fig. 43.
Supplementary Data 24
Computed structure of S6 in Supplementary Fig. 43.
Supplementary Data 25
Computed structure of S7a in Supplementary Fig. 43.
Supplementary Data 26
Computed structure of S7b in Supplementary Fig. 43.
Supplementary Data 27
Computed structure of S7-TS2 in Supplementary Fig. 43.
Supplementary Data 28
Computed structure of S7-TS1 in Supplementary Fig. 43.
Supplementary Data 29
Computed structure of S8 in Supplementary Fig. 43.
Supplementary Data 30
Computed structure of S8-TS in Supplementary Fig. 43.
Supplementary Data 31
Computed structure of S9 in Supplementary Fig. 43.
Supplementary Data 32
CIF file for the X-ray structure of (4S,5S)-12.
Supplementary Data 33
checkCIF reports for the X-ray structure of (4S,5S)-12.
Supplementary Video 1
Video of the transition state P1*-P3*-TS in Supplementary Fig. 42.
Supplementary Video 2
Video of the transition state P3*-TS in Supplementary Fig. 42.
Supplementary Video 3
Video of the transition state P4*-TS in Supplementary Fig. 42.
Supplementary Video 4
Video of the transition state P1*-NH3-TS in Supplementary Fig. 42.
Supplementary Video 5
Video of the transition state P2*-NH3-TS in Supplementary Fig. 42.
Supplementary Video 6
Video of the transition state S1-TS in Supplementary Fig. 43.
Supplementary Video 7
Video of the transition state S2-TS in Supplementary Fig. 43.
Supplementary Video 8
Video of the transition state S3-TS in Supplementary Fig. 43.
Supplementary Video 9
Video of the transition state S4-TS in Supplementary Fig. 43.
Supplementary Video 10
Video of the transition state S5-TS in Supplementary Fig. 43.
Supplementary Video 11
Video of the transition state S7-TS2 in Supplementary Fig. 43.
Supplementary Video 12
Video of the transition state St-TS1 in Supplementary Fig. 43.
Supplementary Video 13
Video of the transition state S8-TS in Supplementary Fig. 43.
Source data
Source Data Fig. 1
Unprocessed SDS-PAGE gel of MT and TC
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, H., Lauterbach, L., Goldfuss, B. et al. Fragmentation and [4 + 3] cycloaddition in sodorifen biosynthesis. Nat. Chem. 15, 1164–1171 (2023). https://doi.org/10.1038/s41557-023-01223-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-023-01223-z
This article is cited by
-
Mining the noncanonical terpenome
Nature Chemical Biology (2023)