Abstract
The coral reef microbiome is central to reef health and resilience. Competitive interactions between opportunistic coral pathogens and other commensal microbes affect the health of coral. Despite great advances over the years in sequencing-based microbial profiling of healthy and diseased coral, the molecular mechanism underlying colonization competition has been much less explored. In this study, by examining the culturable bacteria inhabiting the gastric cavity of healthy Galaxea fascicularis, a scleractinian coral, we found that temperate phages played a major role in mediating colonization competition in the coral microbiota. Specifically, the non-toxigenic Vibrio sp. inhabiting the healthy coral had a much higher colonization capacity than the coral pathogen Vibrio coralliilyticus, yet this advantage was diminished by the latter killing the former. Pathogen-encoded LodAB, which produces hydrogen peroxide, triggers the lytic cycle of prophage in the non-toxicogenic Vibrio sp. Importantly, V. coralliilyticus could outcompete other coral symbiotic bacteria (for example, Endozoicomonas sp.) through LodAB-dependent prophage induction. Overall, we reveal that LodAB can be used by pathogens as an important weapon to gain a competitive advantage over lysogenic competitors when colonizing corals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this manuscript and its Supplementary Information files. The complete genome sequences of the strains used in this study and 16S rRNA gene amplicon sequencing reads have been deposited in GenBank under BioProject PRJNA416452 and PRJNA668462. The accession numbers of bacterial genomes are listed in the Supplementary Table 2. The accession numbers of sequencing reads of 16S rRNA gene amplicon sequencing are listed in Supplementary Table 7. Source data are provided with this paper.
Code availability
The software and codes used in this study are publicly available and customized parameters are indicated in Methods.
References
Carpenter, K. E. et al. One-third of reef-building corals face elevated extinction risk from climate change and local impacts. Science 321, 560–563 (2008).
Pollock, F. J. et al. Coral-associated bacteria demonstrate phylosymbiosis and cophylogeny. Nat. Commun. 9, 4921–4932 (2018).
Vega Thurber, R. et al. Metagenomic analysis of stressed coral holobionts. Environ. Microbiol. 11, 2148–2163 (2009).
Rosenberg, E. & Zilber-Rosenberg, I. Microbes drive evolution of animals and plants: the hologenome concept. mBio 7, e01395 (2016).
van Oppen, M. J. H. & Blackall, L. L. Coral microbiome dynamics, functions and design in a changing world. Nat. Rev. Microbiol. 17, 557–567 (2019).
Reshef, L., Koren, O., Loya, Y., Zilber-Rosenberg, I. & Rosenberg, E. The coral probiotic hypothesis. Environ. Microbiol. 8, 2068–2073 (2006).
Ainsworth, T. D., Thurber, R. V. & Gates, R. D. The future of coral reefs: a microbial perspective. Trends Ecol. Evol. 25, 233–240 (2010).
Lee, S. M. et al. Bacterial colonization factors control specificity and stability of the gut microbiota. Nature 501, 426–429 (2013).
Freter, R. The fatal enteric cholera infection in the guinea pig, achieved by inhibition of normal enteric flora. J. Infect. Dis. 97, 57–65 (1955).
Corr, S. C. et al. Bacteriocin production as a mechanism for the antiinfective activity of Lactobacillus salivarius UCC118. Proc. Natl. Acad. Sci. USA 104, 7617–7621 (2007).
Kamada, N. et al. Regulated virulence controls the ability of a pathogen to compete with the gut microbiota. Science 336, 1325–1329 (2012).
Khosravi, A. et al. Gut microbiota promote hematopoiesis to control bacterial infection. Cell Host Microbe 15, 374–381 (2014).
Li, J., Kuang, W. Q., Long, L. J. & Zhang, S. Production of quorum-sensing signals by bacteria in the coral mucus layer. Coral Reefs 36, 1235–1241 (2017).
Alagely, A., Krediet, C. J., Ritchie, K. B. & Teplitski, M. Signaling-mediated cross-talk modulates swarming and biofilm formation in a coral pathogen Serratia marcescens. ISME J. 5, 1609–1620 (2011).
Krediet, C. J., Ritchie, K. B., Alagely, A. & Teplitski, M. Members of native coral microbiota inhibit glycosidases and thwart colonization of coral mucus by an opportunistic pathogen. ISME J. 7, 980–990 (2013).
Thompson, F. L., Hoste, B., Thompson, C. C., Huys, G. & Swings, G. The coral bleaching Vibrio shiloi Kushmaro et al. 2001 is a later synonym of Vibrio mediterranei Pujalte and Garay 1986. Syst. Appl. Microbiol. 24, 516–519 (2001).
Santoro, E. P. et al. Coral microbiome manipulation elicits metabolic and genetic restructuring to mitigate heat stress and evade mortality. Sci. Adv. 7, eabg3088 (2021).
Tang, K. H. et al. Antagonism between coral pathogen Vibrio coralliilyticus and other bacteria in the gastric cavity of scleractinian coral Galaxea fascicularis. Sci. China-Earth Sci. 63, 157–166 (2020).
Zhou, Y. Q. et al. Identification of bacteria-derived urease in the coral gastric cavity. Sci. China-Earth Sci. 63, 1553–1563 (2020).
Chen, B. et al. Microbiome community and complexity indicate environmental gradient acclimatisation and potential microbial interaction of endemic coral holobionts in the South China Sea. Sci. Total Environ. 765, 142690 (2021).
Tout, J. et al. Increased seawater temperature increases the abundance and alters the structure of natural Vibrio populations associated with the coral Pocillopora damicornis. Front. Microbiol. 6, 432 (2015).
Savary, R. et al. Fast and pervasive transcriptomic resilience and acclimation of extremely heat-tolerant coral holobionts from the northern Red Sea. Proc. Natl. Acad. Sci. USA 118, e2023298118 (2021).
Vezzulli, L. et al. Vibrio infections triggering mass mortality events in a warming Mediterranean Sea. Environ. Microbiol. 12, 2007–2019 (2010).
Rosenberg, E. & Falkovitz, L. The Vibrio shiloi/Oculina patagonica model system of coral bleaching. Annu. Rev. Microbiol. 58, 143–159 (2004).
Gibbin, E. et al. Vibrio coralliilyticus infection triggers a behavioural response and perturbs nutritional exchange and tissue integrity in a symbiotic coral. ISME J. 13, 989–1003 (2019).
Kimes, N. E. et al. Temperature regulation of virulence factors in the pathogen Vibrio coralliilyticus. ISME J. 6, 835–846 (2012).
Banin, E., Vassilakos, D., Orr, E., Martinez, R. J. & Rosenberg, E. Superoxide dismutase is a virulence factor produced by the coral bleaching pathogen Vibrio shiloi. Curr. Microbiol. 46, 418–422 (2003).
Meron, D. et al. Role of flagella in virulence of the coral pathogen Vibrio coralliilyticus. Appl. Environ. Microbiol. 75, 5704–5707 (2009).
Rubio-Portillo, E. et al. Virulence as a side effect of interspecies interaction in Vibrio coral pathogens. mBio 11, e00201-20 (2020).
Rubio-Portillo, E., Yarza, P., Penalver, C., Ramos-Espla, A. A. & Anton, J. New insights into Oculina patagonica coral diseases and their associated Vibrio spp. communities. ISME J. 8, 1794–1807 (2014).
Bourne, D. G. et al. Microbial disease and the coral holobiont. Trends Microbiol. 17, 554–562 (2009).
Ben-Haim, Y., Zicherman-Keren, M. & Rosenberg, E. Temperature-regulated bleaching and lysis of the coral Pocillopora damicornis by the novel pathogen Vibrio coralliilyticus. Appl. Environ. Microbiol. 69, 4236–4242 (2003).
Gavish, A. R., Shapiro, O. H., Kramarsky-Winter, E. & Vardi, A. Microscale tracking of coral–vibrio interactions. ISME Commun. 1, 18 (2021).
Shapiro, O. H. et al. Vortical ciliary flows actively enhance mass transport in reef corals. Proc. Natl. Acad. Sci. USA 111, 13391–13396 (2014).
Shapiro, O. H., Kramarsky-Winter, E., Gavish, A. R., Stocker, R. & Vardi, A. A coral-on-a-chip microfluidic platform enabling live-imaging microscopy of reef-building corals. Nat. Commun. 7, 10860 (2016).
Chen, D. D. et al. Identification and characterization of microsatellite markers for scleractinian coral Galaxea fascicularis and its symbiotic zooxanthellae. Conservation. Genet. Resour. 5, 741–743 (2013).
Parks, D. H. et al. A complete domain-to-species taxonomy for bacteria and archaea. Nat. Biotechnol. 38, 1079–1086 (2020).
Liu, X. et al. Symbiosis of a P2-family phage and deep-sea Shewanella putrefaciens. Environ. Microbiol. 21, 4212–4232 (2019).
Wang, P. et al. Eliminating mcr-1-harbouring plasmids in clinical isolates using the CRISPR/Cas9 system. J. Antimicrob. Chemother. 74, 2559–2565 (2019).
Zeng, Z. et al. Cold adaptation regulated by cryptic prophage excision in Shewanella oneidensis. ISME J. 10, 2787–2800 (2016).
Wang, X. et al. Cryptic prophages help bacteria cope with adverse environments. Nat. Commun. 1, 147 (2010).
Bardwell, J. C., McGovern, K. & Beckwith, J. Identification of a protein required for disulfide bond formation in vivo. Cell 67, 581–589 (1991).
Wang, X., Kim, Y. & Wood, T. K. Control and benefits of CP4-57 prophage excision in Escherichia coli biofilms. ISME J. 3, 1164–1179 (2009).
Wood, T. K., Gonzalez Barrios, A. F., Herzberg, M. & Lee, J. Motility influences biofilm architecture in Escherichia coli. Appl. Microbiol. Biotechnol. 72, 361–367 (2006).
Song, S., Guo, Y., Kim, J. S., Wang, X. & Wood, T. K. Phages mediate bacterial self-recognition. Cell Rep. 27, 737–749 (2019).
Krediet, C. J., Carpinone, E. M., Ritchie, K. B. & Teplitski, M. Characterization of the gacA-dependent surface and coral mucus colonization by an opportunistic coral pathogen Serratia marcescens PDL100. FEMS Microbiol. Ecol. 84, 290–301 (2013).
Guo, Y., Lin, J. & Wang, X. Rapid detection of temperate bacteriophage using a simple motility assay. Environ. Microbiol. Rep. 13, 728–734 (2021).
Tang, K. et al. Prophage Tracer: precisely tracing prophages in prokaryotic genomes using overlapping split-read alignment. Nucleic Acids Res. 49, e128 (2021).
Ding, J. Y., Shiu, J. H., Chen, W. M., Chiang, Y. R. & Tang, S. L. Genomic insight into the host–endosymbiont relationship of Endozoicomonas montiporae CL-33(T) with its coral host. Front. Microbiol. 7, 251 (2016).
Yang, C. S. et al. Endozoicomonas montiporae sp. nov., isolated from the encrusting pore coral Montipora aequituberculata. Int. J. Syst. Evol. Microbiol. 60, 1158–1162 (2010).
Schreiber, L., Kjeldsen, K. U., Obst, M., Funch, P. & Schramm, A. Description of Endozoicomonas ascidiicola sp nov., isolated from Scandinavian ascidians. Syst. Appl. Microbiol. 39, 313–318 (2016).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Lu, S. N. et al. CDD/SPARCLE: the conserved domain database in 2020. Nucleic Acids Res. 48, D265–D268 (2020).
Mai-Prochnow, A. et al. Hydrogen peroxide linked to lysine oxidase activity facilitates biofilm differentiation and dispersal in several Gram-negative bacteria. J. Bacteriol. 190, 5493–5501 (2008).
Campillo-Brocal, J. C., Lucas-Elio, P. & Sanchez-Amat, A. Identification in Marinomonas mediterranea of a novel quinoprotein with glycine oxidase activity. MicrobiologyOpen 2, 684–694 (2013).
Chacon-Verdu, M. D., Gomez, D., Solano, F., Lucas-Elio, P. & Sanchez-Amat, A. LodB is required for the recombinant synthesis of the quinoprotein l-lysine-epsilon-oxidase from Marinomonas mediterranea. Appl. Microbiol. Biotechnol. 98, 2981–2989 (2014).
Gomez, D., Lucas-Elio, P., Solano, F. & Sanchez-Amat, A. Both genes in the Marinomonas mediterranea lodAB operon are required for the expression of the antimicrobial protein lysine oxidase. Mol. Microbiol. 75, 462–473 (2010).
Piewngam, P. et al. Pathogen elimination by probiotic Bacillus via signalling interference. Nature 562, 532–537 (2018).
Zipperer, A. et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature 535, 511–516 (2016).
Selva, L. et al. Killing niche competitors by remote-control bacteriophage induction. Proc. Natl. Acad. Sci. USA 106, 1234–1238 (2009).
Regev-Yochay, G., Trzcinski, K., Thompson, C. M., Malley, R. & Lipsitch, M. Interference between Streptococcus pneumoniae and Staphylococcus aureus: in vitro hydrogen peroxide-mediated killing by Streptococcus pneumoniae. J. Bacteriol. 188, 4996–5001 (2006).
Paul, J. H. Prophages in marine bacteria: dangerous molecular time bombs or the key to survival in the seas? ISME J. 2, 579–589 (2008).
Frazao, N., Sousa, A., Lassig, M. & Gordo, I. Horizontal gene transfer overrides mutation in Escherichia coli colonizing the mammalian gut. Proc. Natl. Acad. Sci. USA 116, 17906–17915 (2019).
Yu, M. et al. Purification and characterization of antibacterial compounds of Pseudoalteromonas flavipulchra JG1. Microbiology-SGM 158, 835–842 (2012).
James, S. G., Holmstrom, C. & Kjelleberg, S. Purification and characterization of a novel antibacterial protein from the marine bacterium D2. Appl. Environ. Microbiol. 62, 2783–2788 (1996).
Lucas-Elio, P., Gomez, D., Solano, F. & Sanchez-Amat, A. The antimicrobial activity of marinocine, synthesized by Marinomonas mediterranea, is due to hydrogen peroxide generated by its lysine oxidase activity. J. Bacteriol. 188, 2493–2501 (2006).
Imlay, J. A. & Linn, S. Mutagenesis and stress responses induced in Escherichia coli by hydrogen peroxide. J. Bacteriol. 169, 2967–2976 (1987).
Los, J. M., Los, M., Wegrzyn, G. & Wegrzyn, A. Differential efficiency of induction of various lambdoid prophages responsible for production of Shiga toxins in response to different induction agents. Microb. Pathog. 47, 289–298 (2009).
Knowles, B. et al. Lytic to temperate switching of viral communities. Nature 531, 466–470 (2016).
Howard-Varona, C., Hargreaves, K. R., Abedon, S. T. & Sullivan, M. B. Lysogeny in nature: mechanisms, impact and ecology of temperate phages. ISME J. 11, 1511–1520 (2017).
Arkin, A. P. et al. KBase: The United States Department of Energy Systems Biology Knowledgebase. Nat. Biotechnol. 36, 566–569 (2018).
Luo, P., He, X. Y., Liu, Q. T. & Hu, C. Q. Developing universal genetic tools for rapid and efficient deletion mutation in Vibrio species based on suicide T-vectors carrying a novel counterselectable marker, vmi480. PLoS ONE 10, e0144465 (2015).
Wang, P. et al. Development of an efficient conjugation-based genetic manipulation system for Pseudoalteromonas. Microb. Cell Fact. 14, 11 (2015).
Bertani, L. E. & Bertani, G. Preparation and characterization of temperate, non-inducible bacteriophage P2 (host: Escherichia coli). J. Gen. Virol. 6, 201–212 (1970).
Garneau, J. R., Depardieu, F., Fortier, L. C., Bikard, D. & Monot, M. PhageTerm: a tool for fast and accurate determination of phage termini and packaging mechanism using next-generation sequencing data. Sci Rep. 7, 8292 (2017).
Pratt, L. A. & Kolter, R. Genetic analysis of Escherichia coli biofilm formation: roles of flagella, motility, chemotaxis and type I pili. Mol. Microbiol. 30, 285–293 (1998).
Page, A. J. et al. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31, 3691–3693 (2015).
Brynildsrud, O., Bohlin, J., Scheffer, L. & Eldholm, V. Rapid scoring of genes in microbial pan-genome-wide association studies with Scoary. Genome Biol. 17, 238 (2016).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
Yilmaz, P. et al. The SILVA and ‘All-species Living Tree Project (LTP)’ taxonomic frameworks. Nucleic Acids Res. 42, D643–D648 (2014).
Chong, J., Liu, P., Zhou, G. & Xia, J. Using MicrobiomeAnalyst for comprehensive statistical, functional, and meta-analysis of microbiome data. Nat. Protoc. 15, 799–821 (2020).
Nagpal, S., Singh, R., Yadav, D. & Mande, S. S. MetagenoNets: comprehensive inference and meta-insights for microbial correlation networks. Nucleic Acids Res. 48, W572–W579 (2020).
Acknowledgements
X.W. was supported by the National Natural Science Foundation of China (grant nos. 42188102, 91951203 and 31625001), the National Key R&D Programme of China (grant no. 2018YFC1406500), Local Innovative and Research Teams Project of Guangdong Pearl River Talents Programme (grant no. 2019BT02Y262), Guangdong Major Project of Basic and Applied Basic Research (grant no. 2019B030302004) and the Key Special Project for Introduced Talents Team of Southern Marine Science and Engineering Guangdong Laboratory (Guangzhou) (grant no. GML2019ZD0407). T.L. was supported by K.C. Wong Education Foundation (grant no. GJTD-2020-12). P.W. was supported by the National Natural Science Foundation of China (grant no. 32070175) and Youth Innovation Promotion Association CAS (grant no. 2021345).
Author information
Authors and Affiliations
Contributions
X.W. and K.T. designed the study. W.W., K.T., P.W., Z.Z., T.X., W.Z., T.L., Y.W. and X.W. performed the experiments. W.W., K.T. and P.W. analysed the data. X.W., K.T. and Y.W. contributed materials/analysis tools. X.W., P.W., K.T. and W.W. wrote the manuscript. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
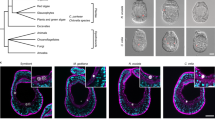
Extended Data Fig. 1 The culture-dependent and culture-independent analysis of G. fascicularis microbiota.
(a) Bacterial isolates were identified by 16S rRNA gene sequencing. Data from our cultivable 405 isolates were reanalyed. The number after each genus indicates the number of isolates obtained. Vibrio and Pseudoalteromonas species were assigned according to the top hit using NCBI 16S_ribosomal_RNA database. During isolation from different coral colonies, three gammaproteobacterial genera, Vibrio, Pseudoalteromonas and Halomonas, were consistently isolated. Vibrio and Pseudoalteromonas accounted for ~50% of the total 405 isolates from either healthy or heat-stressed corals sampled from different locations around Hainan Island. Among 121 Vibrio isolates, 81 isolates were classified as V. alginolyticus and two isolates were classified as V. coralliilyticus. Among 73 Pseudoalteromonas isolates, 29 isolates were classified as P. flavipulchra. (b) Bacteria with different colony morphologies isolated from gastric fluid samples on 2216E plates (the geographical locations of these corals are shown below each plate). The red arrow indicates transparent spots formed on Vibrio colonies. (c) Distribution and relative abundance of major microbial family in G. fascicularis polyps (n = 2) at 26 °C and 32 °C based on 16S rRNA gene V4 amplicon sequencing. The relative abundance of Vibrio and Pseudoalteromonas accounted for ~5% of total reads of the 16S rRNA gene amplicon in G. fascicularis polyps and up to ~20% after heating for 7 days with the occurrence of tissue necrosis.
Extended Data Fig. 2 Va43009 was less pathogenic than Vc43001.
(a) The percentage of tissue loss was measured as the area of tissue lost relative the total area of the coral fragment of the immerse-based infection assay in Fig. 1. (b) G. fascicularis fragments before 32 °C treatment (upper panel) and under 32 °C treatment for 1 day (middle panel) and 2 days (lower panel). Three coral fragments were prepared in the same way as those used in the immersion infection assay. For each coral fragment, only the right side was injected. Red arrows indicate the side (two sides were separated by elastic bands on each fragment) that was injected with Va43009 and Vc43001 or seawater (control). A total of 0.2–1 ×108 cells were injected into the gastric cavity of one corallite, and 5-8 corallites in the center of the right region received an injection. The coral fragments were acclimatized separately at 26 °C for 2 days before temperate was increased to 32 °C. After 1 day, the right side of the coral fragment that had been injected with Vc43001 showed stronger signs of necrosis than the left side. After 2 days, the whole fragments lost tissue on the side injected with Vc43001, suggesting that Vc43001 could achieve a successful infection in the context of the native microbiota in the coral gastric cavity. There was no obvious difference between the right side injected with Va43009 and the left side after 1 day, and sporadic tissue loss of the coenenchyma appeared similar on both sides (~10%) after 2 days. (c) Quantitation of the bleaching rate of the corals in panel b. The percentage of tissue loss was calculated as the area of lost tissue divided by the area of the coral fragment.
Extended Data Fig. 3 Construction and verification of mutant strains.
(a) The GfP2 prophage deletion mutant ΔGfP2 in Va43009. (b) The gfp gene fused with the mcp gene of GfP2 in Va43009; integration of the gfp gene with the gentamicin promoter (PGm) in the chromosome of strain Va43009 (c) and strain ΔGfP2 (d). (e) In-frame deletion lodAB in strain Vc43001. (f) Integration of the gfp gene with the gentamicin promoter (PGm) in the chromosome of strain Vc43001.
Extended Data Fig. 4 GfP2 excision alters the promoter sequence but not dsbA expression.
(a) A schematic diagram illustrating the excision of the prophage GfP2 from the Va43009 chromosome and the formation of GfP2 replicative form (RF) molecules in the cytoplasm of Va43009. The PattR promoter flanking the attR site drives the expression of dsbA when the GfP2 prophage is integrated, and the PattB promoter flanking the attB site drives the expression of dsbA when GfP2 is excised. (b) Sequence analysis of the PattR and PattB promoters. The -35 and -10 boxes predicted from BPROM in Softberry and the Shine–Dalgarno (SD) sequences are marked. The same sequence in the two promoters is highlighted in red, the coding region of dsbA is italicized, and the start codon of dsbA is shown in bold letters. The core phage attachment sites (45 bp) are in red letters and underlined. (c) A schematic of lacZ reporter constructs containing the PattR and PattB promoters shown in (b), with a qualitative lacZ assay using X-gal plates shown. 1–4 are four independent colonies containing PattR in ER2738, and 5-8 are four independent colonies containing PattB in ER2738. (d) Quantitative results showing PattR and PattB promoter activity in both Va43009ΔGfP2 and E. coli ER2738 hosts according to β-gal activity. Data are shown as the mean ± SD. Each pair of groups was compared using an unpaired t test with 95% confidence intervals (n = 3; two-tailed p value).
Extended Data Fig. 5 The addition of NAC reduced the ability of Vc43001 to activate GfP2 in Va43009.
The excision rate of GfP2 and the number of GfP2 RFs when coculturing Va43009 and Vc43001 with or without N-acetyl L-cysteine (NAC) (an inhibitor of reactive oxygen species). Data are shown as the mean ± SD. Each pair of groups was compared using an unpaired t test with 95% confidence intervals (n = 3; two-tailed p value).
Extended Data Fig. 6 Construction of chromosomally tagged antibiotic resistance gene strains and growth measurements in monoculture or coculture.
PCR verification of the mutant strains Vc43001 Gmr, Vc43001ΔlodAB Gm (a) and Va43009 Cmr (b). (c-f) Growth of different strains. Chromosome tagging of antibiotic resistance genes did not affect the growth of these strains. (g) Comparison of monoculture growth of strains. Data are the same as in panel c-f. (h-j) Comparison of the growth of cocultured strains. Growths of strains in coculture are highlighted in red. Growth of strains in monoculture is shown black, the data for which are the same as those in panel g. Data are shown as the mean ± SD. n = 3 for c-j.
Extended Data Fig. 7 Analysis of colonized bacteria using agar plates with corresponding antibiotics.
(a) Boxplots of CFUs of Va43009 and Vc40001 from polyps in each group were determined in 2216E plates containing chloramphenicol and gentamicin, respectively. n = 5 (Va43009 in Group 3; Va43009 and Vc43001ΔlodAB in Group 4); n = 6 (Va43009 in Group 1; Vc43001 in Group 2; and Vc43001 in Group 3). Statistically significant differences were detected by Kruskal–Wallis test with Dunn’s correction. (b) Ratio of Va43009:Vc43001 in Group 3 and Va43009:Vc43001ΔlodAB in Group 4. Statistically significant differences were detected by for multiple comparisons and Mann–Whitney test with 95% confidence intervals. n = 6, two-tailed p value. In boxplots, centre lines indicate median values and the whiskers extend from the 25th or 75th percentiles to the minimum or maximum values.
Extended Data Fig. 8 Vc43001 outcompetes other coral symbiotic bacteria through LodAB-dependent prophage induction.
(a) Competition between Vc43001/Vc43001ΔlodAB and other coral symbiotic bacteria. Data are shown as the mean ± SD. Each pair of groups was compared using an unpaired t test with 95% confidence intervals (n = 5; two-tailed p value). (b) The induction of prophage by Vc43001 but not by Vc43001ΔlodAB was evaluated by qPCR to quantify the phage structural gene and gyrB gene. Data are shown as the mean ± SD. Each pair of groups was compared using an unpaired t test with 95% confidence intervals (n = 3; two-tailed p value).
Extended Data Fig. 9 Schematic diagram of interplay between coral pathogen and commensal bacteria.
LodAB-mediated prophage induction plays an important role in colonization competition of gastric bacteria, affecting coral health.
Supplementary information
Supplementary Information
Supplementary Text (Results, Materials and Methods), Figs. 1–11, Discussion and Tables 1–7.
Supplementary Data 1 and 2
Supplementary Data 1, List of P2 major capsid proteins retrieved from IMG/M database. Supplementary Data 2, List of LodA homologues and domain annotation.
Supplementary Data 3
Source data for Supplementary Information.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3b
Unprocessed gels.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed gels.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Rights and permissions
About this article
Cite this article
Wang, W., Tang, K., Wang, P. et al. The coral pathogen Vibrio coralliilyticus kills non-pathogenic holobiont competitors by triggering prophage induction. Nat Ecol Evol 6, 1132–1144 (2022). https://doi.org/10.1038/s41559-022-01795-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-022-01795-y
This article is cited by
-
The coral microbiome in sickness, in health and in a changing world
Nature Reviews Microbiology (2024)
-
Viral predation pressure on coral reefs
BMC Biology (2023)
-
Removal of competing bacteria in coral microbiome through “trojan virus”: A newly discovered mechanism of coral pathogenicity
Science China Earth Sciences (2023)