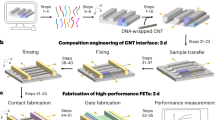
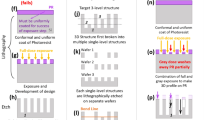
Abstract
Lithographic scaling of periodic three-dimensional patterns is critical for advancing scalable nanomanufacturing. Current state-of-the-art quadruple patterning or extreme-ultraviolet lithography produce a line pitch down to around 30 nm, which might be further scaled to sub-20 nm through complex post-fabrication processes. Herein, we report the use of three-dimensional (3D) DNA nanostructures to scale the line pitch down to 16.2 nm, around 50% smaller than state-of-the-art results. We use a DNA modular epitaxy approach to fabricate 3D DNA masks with prescribed structural parameters (geometry, pitch and critical dimensions) along a designer assembly pathway. Single-run reactive ion etching then transfers the DNA patterns to a Si substrate at a lateral critical dimension of 7 nm and a vertical critical dimension of 2 nm. The nanolithography guided by DNA modular epitaxy achieves a smaller pitch than the projected values for advanced technology nodes in field-effect transistors, and provides a potential complement to the existing lithographic tools for advanced 3D nanomanufacturing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available within the article and its Supplementary Information files and from the corresponding authors upon reasonable request.
References
Lu, W. & Lieber, C. M. Nanoelectronics from the bottom up. Nat. Mater. 6, 841–850 (2007).
Soukoulis, C. M. & Wegener, M. Past achievements and future challenges in the development of three-dimensional photonic metamaterials. Nat. Photon. 5, 523–530 (2011).
Kildishev, A. V., Boltasseva, A. & Shalaev, V. M. Planar photonics with metasurfaces. Science 339, 1232009 (2013).
Fu, J., Schoch, R. B., Stevens, A. L., Tannenbaum, S. R. & Han, J. A patterned anisotropic nanofluidic sieving structure for continuous-flow separation of DNA and proteins. Nat. Nanotechnol. 2, 121–128 (2007).
Loh, O. Y. & Espinosa, H. D. Nanoelectromechanical contact switches. Nat. Nanotechnol. 7, 283–295 (2012).
Chau, R. et al. Benchmarking nanotechnology for high-performance and low-power logic transistor applications. IEEE Trans. Nanotechnol. 4, 153–158 (2005).
James, D. IEDM 2017: Intel’s 10 nm Platform Process (2017); https://sst.semiconductor-digest.com/chipworks_real_chips_blog/2017/12/
Vincent, B., Lariviere, S., Wilson, C., Kim, R. H. & Ervin, J. Virtual fabrication and advanced process control improve yield for SAQP process assessment with 16 nm half-pitch. In Proc. SPIE, Adv. Etch. Technol. Nanopatterning VIII 109630Q (SPIE, 2019).
The International Technology Roadmap for Semiconductors 2.0: 2015 Edition (Semiconductor Industry Association, 2015).
Wagner, C. & Harned, N. EUV lithography: lithography gets extreme. Nat. Photon. 4, 24–26 (2010).
Chen, Y. Nanofabrication by electron beam lithography and its applications: a review. Microelectron. Eng. 135, 57–72 (2015).
Park, M., Harrison, C., Chaikin, P. M., Register, R. A. & Adamson, D. H. Block copolymer lithography: periodic arrays of 1011 holes in 1 square centimeter. Science 276, 1401–1404 (1997).
Chou, S. Y., Krauss, P. R. & Renstrom, P. J. Imprint lithography with 25-nanometer resolution. Science 272, 85–87 (1996).
Wu, B. & Kumar, A. Extreme ultraviolet lithography and three dimensional integrated circuit—a review. Appl. Phys. Rev. 1, 011104 (2014).
Seeman, N. C. DNA in a material world. Nature 421, 427–431 (2003).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Dietz, H., Douglas, S. M. & Shih, W. M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725–730 (2009).
Wei, B., Dai, M. J. & Yin, P. Complex shapes self-assembled from single stranded DNA tiles. Nature 485, 623–626 (2012).
Ke, Y., Ong, L. L., Shih, W. M. & Yin, P. Three-dimensional structures self-assembled from DNA bricks. Science 338, 1177–1183 (2012).
Ke, Y. et al. DNA brick crystals with prescribed depths. Nat. Chem. 6, 994–1002 (2014).
Ong, L. L. et al. Programmable self-assembly of three-dimensional nanostructures from 10,000 unique components. Nature 552, 72–77 (2017).
Maune, H. T. et al. Self-assembly of carbon nanotubes into two-dimensional geometries using DNA origami templates. Nat. Nanotechnol. 5, 61–66 (2009).
Liu, N. & Liedl, T. DNA-assembled advanced plasmonic architectures. Chem. Rev. 118, 3032–3053 (2018).
Sun, W. et al. Precise pitch-scaling of carbon nanotube arrays within three-dimensional DNA nanotrenches. Science 368, 874–877 (2020).
Zhao, M. et al. DNA-directed nanofabrication of high-performance carbon nanotube field-effect transistors. Science 368, 878–881 (2020).
Sun, W. et al. Casting inorganic structures with DNA molds. Science 346, 1258361 (2014).
Liu, X. et al. Complex silica composite nanomaterials templated with DNA origami. Nature 559, 593–598 (2018).
Jin, Z. et al. Metallized DNA nanolithography for encoding and transferring spatial information for graphene patterning. Nat. Commun. 4, 1663 (2013).
Shen, B. et al. Plasmonic nanostructures through DNA-assisted lithography. Sci. Adv. 4, eaap8978 (2018).
Hong, F., Zhang, F., Liu, Y. & Yan, H. DNA origami: scaffolds for creating higher order structures. Chem. Rev. 117, 12584–12640 (2017).
Tikhomirov, G., Petersen, P. & Qian, L. Fractal assembly of micrometre-scale DNA origami arrays with arbitrary patterns. Nature 552, 67–71 (2017).
Diagne, C. T., Brun, C., Gasparutto, D., Baillin, X. & Tiron, R. DNA origami mask for sub-ten-nanometer lithography. ACS Nano 10, 6458–6463 (2016).
Surwade, S. P., Zhao, S. & Liu, H. Molecular lithography through DNA-mediated etching and masking of SiO2. J. Am. Chem. Soc. 133, 11868–11871 (2011).
Surwade, S. P. et al. Nanoscale growth and patterning of inorganic oxides using DNA nanostructure templates. J. Am. Chem. Soc. 135, 6778–6781 (2013).
Schulman, R. & Winfree, E. Synthesis of crystals with a programmable kinetic barrier to nucleation. Proc. Natl Acad. Sci. USA 104, 15236–15241 (2007).
Barish, R. D., Schulman, R., Rothemund, P. W. K. & Winfree, E. An information-bearing seed for nucleating algorithmic self-assembly. Proc. Natl Acad. Sci. USA 106, 6054–6059 (2009).
Woods, D. et al. Diverse and robust molecular algorithms using reprogrammable DNA self-assembly. Nature 567, 366–372 (2019).
Zhang, Y., Reinhardt, A., Wang, P., Song, J. & Ke, Y. Programming the nucleation of DNA brick self-assembly with a seeding strand. Angew. Chem. Int. Ed. Engl. 132, 8594–8600 (2020).
Hooks, D. E., Fritz, T. & Ward, M. D. Epitaxy and molecular organization on solid substrates. Adv. Mater. 13, 227–241 (2001).
Peng, X., Schlamp, M. C., Kadavanich, A. V. & Alivisatos, A. P. Epitaxial growth of highly luminescent CdSe/CdS core/shell nanocrystals with photostability and electronic accessibility. J. Am. Chem. Soc. 119, 7019–7029 (1997).
Choi, J. K., Sargsyan, G., Shabbir-Hussain, M., Holmes, A. E. & Balaz, M. Chiroptical detection of condensed nickel(ii)-Z-DNA in the presence of the B-DNA via porphyrin exciton coupled circular dichroism. J. Phys. Chem. B 115, 10182–10188 (2011).
Raley, A., Mack, C., Thibaut, S., Liu, E. & Ko, A. Benchmarking of EUV lithography line/space patterning versus immersion lithography multipatterning schemes at equivalent pitch. In Proc. SPIE, International Conference on Extreme Ultraviolet Lithography 2018 1080915 (SPIE, 2018).
Wei, B., Ong, L. L., Chen, J., Jaffe, A. S. & Yin, P. Complex reconfiguration of DNA nanostructures. Angew. Chem. Int. Ed. Engl. 53, 7475–7479 (2014).
Rueger, N. R. et al. Role of steady state fluorocarbon films in the etching of silicon dioxide using CHF3 in an inductively coupled plasma reactor. J. Vac. Sci. Technol. A 15, 1881–1889 (1997).
Khazi, I., Muthiah, U. & Mescheder, U. 3D free forms in c-Si via grayscale lithography and RIE. Microelectron. Eng. 193, 34–40 (2018).
Yamazaki, K., Kurihara, K., Namatsu, H., Yamazaki, K. & Kurihara, K. Supercritical resist dryer. J. Vac. Sci. Technol. B 18, 780–784 (2000).
Gopinath, A., Miyazono, E., Faraon, A. & Rothemund, P. W. K. Engineering and mapping nanocavity emission via precision placement of DNA origami. Nature 535, 401–405 (2016).
Brun, C. et al. Metallic conductive nanowires elaborated by PVD metal deposition on suspended DNA bundles. Small 13, 1700956 (2017).
Zhou, F. et al. Programmably shaped carbon nanostructure from shape-conserving carbonization of DNA. ACS Nano 10, 3069–3077 (2016).
Shen, J., Sun, W. & Yin, P. Scalable nucleic acid-based nanofabrication. US patent 2017/0190573 A1 (2017).
Acknowledgements
We thank J.D. Deng, L. Xie and S. Stoilova-McPhie at the Center for Nanoscale Systems of Harvard University for valuable discussion and help with cryo-EM work. This work is supported by NSF (CMMI-1333215, CMMI-1344915 and CBET-1729397), AFOSR (MURI FATE, FA9550–15–1–0514) and internal funding support from the Wyss Institute to P.Y. D.L. is supported by a Merck Fellowship of the Life Sciences Research Foundation.
Author information
Authors and Affiliations
Contributions
J.S. conceived, designed and conducted the lithography study and wrote the manuscript. W.S. conceived, designed and conducted the DNA modular epitaxy study and wrote the manuscript. D.L. performed cryo-EM analysis. D.L. and T.S. analysed the data and co-wrote the manuscript. P.Y. conceived and supervised the study and wrote the paper. All authors reviewed, edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
A patent based on this work51 was issued to J.S., W.S. and P.Y.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of DNA mask 12H-grid.
a, SEM image section of DNA mask 12H-grid. b, The corresponding SEM line-scan profiles along the x axis. c, The histogram of mask taller-line width. d, The histogram of taller-line pitch. e, The corresponding SEM line-scan profiles along the z axis. f, The histogram of mask lower-line width. g, The histogram of lower-line pitch. h, SEM images of randomly selected DNA mask 12H-grid.
Extended Data Fig. 2 Characterization of DNA mask 12H-b.
a, SEM image section of DNA mask 12H-b with labeled scan-lines. b, The corresponding SEM line-scan profiles along the x axis. c, The histogram of DNA line width. d, The histogram of DNA line pitch. e, SEM images of randomly selected DNA mask 12H-b.
Extended Data Fig. 3 Characterization of DNA mask 10H-b.
a, SEM image section of DNA mask 10H-b with labeled scan-lines. b, The corresponding SEM line-scan profiles along the x axis. c, The histogram of DNA line width. d, The histogram of DNA line pitch. e, SEM images of randomly selected DNA mask 10H-b.
Extended Data Fig. 4 Characterization of DNA mask 8H-b.
a, SEM image section of DNA mask 8H-b-0.5H2 with labeled scan-lines. b, The corresponding SEM line-scan profiles along the x axis. c, The histogram of DNA line width. d, The histogram of DNA line pitch. e, SEM images of randomly selected DNA mask 8H-b-0.5H2.
Extended Data Fig. 5 Characterization of DNA mask 6H-b.
a, SEM image section of DNA mask 6H-b with labeled scan-lines. b, The corresponding SEM line-scan profiles along the x axis. c, The histogram of DNA line width. d, The histogram of DNA line pitch. e, SEM images of randomly selected DNA mask 6H-b.
Extended Data Fig. 6 Characterization of silicon pattern Si-12H-b.
a, SEM image section of silicon pattern Si-12H-b with labeled scan-line profiles. b, The corresponding SEM line-scan profiles along the x axis. c, The histogram of silicon line width. d, The histogram of silicon line pitch. e, SEM images of randomly selected silicon pattern Si-12H-b.
Extended Data Fig. 7 Characterization of silicon pattern Si-10H-b.
a, SEM image section of silicon pattern Si-10H-b with labeled scan-line profiles. b, The corresponding SEM line-scan profiles along the x axis. c, The histogram of silicon line width. d, The histogram of silicon line pitch. e, SEM images of randomly selected silicon pattern Si-10H-b.
Extended Data Fig. 8 Characterization of silicon pattern Si-8H-b-0.5H2.
a, SEM image section of silicon pattern Si-8H-b-0.5H2 with labeled scan-line profiles. b, The corresponding SEM line-scan profiles along the x axis. c, The histogram of silicon line width. d, The histogram of silicon line pitch. e, SEM images of randomly selected silicon pattern Si-8H-b-0.5H2.
Extended Data Fig. 9 Characterization of silicon pattern Si-6H-b.
a, SEM image section of silicon pattern Si-6H-b with labeled scan-lines. b, The corresponding SEM line-scan profiles along the x axis. c, The histogram of silicon line width. d, The histogram of silicon line pitch. e, SEM images of randomly selected silicon pattern Si-6H-b.
Extended Data Fig. 10 Characterization of silicon pattern Si-12H-grid.
a, SEM image section of silicon pattern Si-12H-grid. b, The corresponding SEM line-scan profiles along the x axis. c, The histogram of taller-Si-line width. d, The histogram of taller-Si-line pitch. e, The corresponding SEM line-scan profiles along the z axis. f, The histogram of lower-Si-line width. g, The histogram of lower-Si-line pitch. h, SEM images of randomly selected silicon pattern Si-12H-grid.
Supplementary information
Supplementary Information
Supplementary Methods, Figs. 1–24 and Tables 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shen, J., Sun, W., Liu, D. et al. Three-dimensional nanolithography guided by DNA modular epitaxy. Nat. Mater. 20, 683–690 (2021). https://doi.org/10.1038/s41563-021-00930-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41563-021-00930-7