Abstract
Type 1 interferon suppresses viral replication by upregulating the expression of interferon-stimulated genes with diverse antiviral properties1. The replication of human immunodeficiency virus type 1 (HIV-1) is naturally inhibited by interferon, with the steps between viral entry and chromosomal integration of viral DNA being notably susceptible2,3,4,5. The interferon-stimulated gene myxovirus resistance 2 has been defined as an effective postentry inhibitor of HIV-1, but is only partially responsible for interferon’s suppressive effect6,7,8. Using small interfering RNA-based library screening in interferon-α-treated cells, we sought to characterize further interferon-stimulated genes that target the pre-integration phases of HIV-1 infection, and identified human tripartite-containing motif 5α (TRIM5α) as a potent anti-HIV-1 restriction factor. Human TRIM5α, in contrast with many nonhuman orthologues, has not generally been ascribed substantial HIV-1 inhibitory function, a finding attributed to ineffective recognition of cytoplasmic viral capsids by TRIM5α2,9,10. Here, we demonstrate that interferon-α-mediated stimulation of the immunoproteasome, a proteasome isoform mainly present in immune cells and distinguished from the constitutive proteasome by virtue of its different catalytic β-subunits, as well as the proteasome activator 28 regulatory complex11,12,13, and the associated accelerated turnover of TRIM5α underpin the reprogramming of human TRIM5α for effective capsid-dependent inhibition of HIV-1 DNA synthesis and infection. These observations identify a mechanism for regulating human TRIM5α antiviral function in human cells and rationalize how TRIM5α participates in the immune control of HIV-1 infection.
Similar content being viewed by others
Main
Interferon-α (IFN-α) mobilizes the expression of hundreds of IFN-stimulated genes (ISGs), with the functions and viral substrates of many awaiting definition1. To identify ISGs that suppress HIV-1 replication, we designed a small interfering RNA (siRNA) library targeting 598 ISGs (plus two negative controls; Supplementary Table 1). Focusing on the early stages of infection (up to and including viral transcription), two cultures of IFN-α-responsive U87-MG CD4+ CXCR4+ cells were transfected with each siRNA, with one being maintained with 500 U ml−1 IFN-α for 24 h and one without. All cultures were then challenged with HIV-1/Nef-internal ribosome entry signal (IRES)-Renilla, a modified replication-competent reporter virus, and infection quantified by measuring Renilla luciferase activity at 48 h (Fig. 1a, unadjusted levels of infection indicated in left-hand panel and folds of IFN-α-mediated suppression indicated in right-hand panel; Supplementary Fig. 1). Three genes of well-established relevance to HIV-1 infection and whose suppression corresponded with markedly increased levels of infection in the presence of IFN-α were IFN regulatory factor 9 (IRF9), myxovirus resistance 2 (MX2) and tripartite-containing motif 5α (TRIM5α; Supplementary Fig. 2 displays the 14 genes with the strongest effects). IRF9, a transcription factor required for ISG induction4, and MX2, an established HIV-1 inhibitory ISG6,7, were anticipated finds, but TRIM5α was completely unexpected. Indeed, human TRIM5α has hithertofore been regarded as being virtually inactive against HIV-1; in contrast, nonhuman TRIM5α proteins, for example, from rhesus macaque, are potent HIV-1 restriction factors that recognize postentry viral capsids to induce their premature fragmentation and the inhibition of reverse transcription2,9.
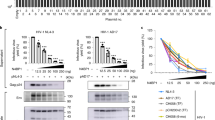
a, Dot plots of NL4-3/Nef-IRES-Renilla infectivity and IFN-α-induced fold inhibition at 48 h postinfection in U87-MG CD4+ CXCR4+ cells doubly transfected with siRNAs targeting 598 ISGs and two negative controls with or without the addition of 500 U ml−1 IFN-α for 24 h. Three notably influential ISGs are indicated in red. b, Percentage of GFP+ cells and IFN-α-induced fold inhibition in U87-MG CD4+ CXCR4+ cells infected with NL4-3/Nef-IRES-GFP after TRIM5α silencing using SMARTpool (n = 5) or individual siRNAs (n = 4) with or without added 500 U ml−1 IFN-α. c, Immunoblot analysis of TRIM5α expression in U87-MG CD4+ CXCR4+ cells after siRNA transfection, with α-tubulin serving as a loading control. One representative immunoblot from two independent experiments is shown. d, NL4-3/Nef-IRES-GFP infection and IFN-α-induced inhibition in U87-MG CD4+ CXCR4+ bulk (TRIM5α: n = 14) and clonal (TRIM5α_1: n = 8 and TRIM5α_2: n = 7) cell lines transduced to express TRIM5α-specific guide RNAs, with or without added 500 U ml−1 IFN-α. CRISPR–Cas9 control cells expressed an unrelated guide RNA (n = 14). e, Ablation of TRIM5α expression in CRISPR–Cas9-engineered U87-MG CD4+ CXCR4+ cells was verified by immunoblotting. α-Tubulin served as a loading control. One representative immunoblot from three independent experiments is shown. f, NL4-3/Nef-IRES-GFP infectivity and IFN-α-induced inhibition at 48 h postinfection in TRIM5α-deficient U87-MG CD4+ CXCR4+ cells (CRISPR TRIM5α, TRIM5α_1 and TRIM5α_2) or cells expressing an unrelated guide RNA (CRISPR control) transduced with EasiLV lentivirus vectors expressing luciferase (Luc) or an rTRIM5α with or without added 500 U ml−1 IFN-α (n = 3). g, NL4-3/Nef-IRES-Renilla infectivity and IFN-α-induced inhibition at 48 h postinfection in primary human CD4+ T cells transduced with shRNAs targeting TRIM5α or a control shRNA, and treated with or without 2,000 U ml−1 IFN-α for 24 h before infection (n = 5). h, U87-MG CD4+ CXCR4+ cells were transfected with control or TRIM5α-specific siRNAs and treated or not with 500 U ml−1 IFN-α for 24 h before 2-h infections with NL4-3/Nef-IRES-GFP (corresponding to 20 ng p24Gag). DNA was harvested at 48 h postinfection, and early reverse transcription products (strong stop) and IFN-α-induced inhibition were determined by qPCR (n = 5). Data are represented as the mean ± s.d. P values (95% confidence interval) were determined using two-sided unpaired (b, d, f, h) or paired (g) t-tests. AU, arbitrary units.
The restorative impact of each component TRIM5α siRNA of the SMARTpool against IFN-α-suppressed HIV-1 infection was confirmed using an HIV-1/Nef-IRES-green fluorescent protein (GFP) reporter virus, with IFN-α-mediated inhibition consistently decreasing by twofold to fourfold relative to the nonsilencing siControl (Fig. 1b) and TRIM5α expression being reliably silenced (Fig. 1c). Similar levels of rescue of infection were also observed in a bulk cell population and two independent knockout cell lines in which the endogenous TRIM5α alleles had been inactivated using clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein 9 (Cas9) genome editing (Fig. 1d,e and Supplementary Fig. 3) and across a range of IFN-α doses (Supplementary Fig. 4). Importantly, re-introduction of CRISPR-resistant human TRIM5α (rTRIM5α) cDNA into these cells completely restored IFN-α-mediated suppression, whereas no changes in HIV-1 susceptibility were detected with an irrelevant control gene (luciferase) (Fig. 1f). Short hairpin RNA (shRNA)-mediated silencing was then used to suppress TRIM5α expression in primary human CD4+ T cells. Relative to donor-matched control samples, reducing TRIM5α levels substantially impeded the IFN-α-mediated inhibition of HIV-1 infection, therefore confirming the importance of this IFN-α-regulated antiviral pathway in the principal in vivo cell target of HIV-1 infection (Fig. 1g and Supplementary Fig. 5). As noted earlier, it has been established that IFN-α is a potent and early inhibitor of HIV-1 reverse transcription following entry into susceptible cells3: TRIM5α’s substantial contribution to this effect was demonstrated by using quantitative polymerase chain reaction (qPCR) to measure the restoration of viral cDNA accumulation following TRIM5α silencing in IFN-α-treated cells (approximately fivefold rescue; Fig. 1h). The broad sensitivity of diverse strains to IFN-α-activated human TRIM5α was confirmed using siRNA-mediated silencing and ten HIV-1 isolates engineered to confer GFP expression following productive infection (extents of rescue ranging from twofold to fivefold; Supplementary Fig. 6).
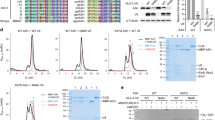
TRIM5α-mediated inhibition is initiated through the recognition of viral capsids in the cytoplasm, leading to their fragmentation and the suppression of reverse transcription (viral DNA synthesis). Binding is both sequence and structure specific, and is dependent on the capsid lattice (which is composed of multiple hexagonal and pentagonal assemblies of the viral Capsid (CA) protein)14 and the carboxy-terminal SPRY (a sequence repeat in dual-specificity kinase SPlA and RYanodine receptors) domain of TRIM5α2,9. The specificity of human TRIM5α for HIV-1 CA was demonstrated by the reduced inhibitory effect of IFN-α after TRIM5α silencing that was seen for virus-bearing wild-type HIV-1 CA (WTCA), but not for a matched HIV-1-based chimeric virus-carrying (nonrecognized) CA derived from a simian immunodeficiency virus (SIVCA) (Fig. 2a). The suppression of HIV-1 by human TRIM5α is not, therefore, the result of a generalized induction of TRIM5α antilentiviral activity by IFN-α.
a, Percentage of GFP+ cells and IFN-α-induced inhibition at 48 h postinfection in U87-MG CD4+ CXCR4+ cells challenged with GFP-encoding HIV-1-based vectors containing WTCA or CA from SIVMAC (SIVCA). TRIM5α was silenced using SMARTpool siRNAs and cells treated with or without 500 U ml−1 IFN-α before infection (n = 5). P values (95% confidence interval) were determined using two-sided unpaired t-tests. b, NL4-3/Nef-IRES-GFP infectivity and IFN-α-induced inhibition at 48 h postinfection in TRIM5α/MX2 doubly deficient U87-MG CD4+ CXCR4+ cells (Supplementary Fig. 7) transduced with EasiLV lentivirus vectors expressing luciferase, CD8, CRISPR-resistant MX2, TRIMCYP or CRISPR-resistant TRIM5α, TRIM5αΔSPRY, TRIM5α R332G/R335G or TRIM5α H43Y with or without added 500 U ml−1 IFN-α (n = 6). Numbers represent fold inhibition of HIV-1 infection after IFN-α treatment. Data are represented as the mean ± s.d.
The central importance of the SPRY domain for TRIM5α function in the presence of IFN-α was then confirmed in gene transduction-based experiments using U87-MG CD4+ CXCR4+ cells where endogenous TRIM5α and MX2 expression were ablated using CRISPR–Cas9 genome editing (Fig. 2b and Supplementary Figs. 7 and 8): luciferase and CD8 served as negative controls for not affecting infection (approximately fivefold suppression of infection in the presence of IFN-α), whereas a previously defined TRIM–cyclophilin (TRIMCYP) fusion protein and MX2 were positive controls6,7,10 whose inhibitory phenotypes do not depend on IFN-α (Fig. 2b). Consistent with our findings with the endogenous gene, ectopically expressed human TRIM5α inhibited HIV-1 infection very effectively when IFN-α was added (total suppression of ~14-fold). Removing the SPRY domain abolished this effect (approximately fivefold suppression), whereas including mutations (R332G and R335G) that have previously been shown to confer anti-HIV-1 activity15,16 still did so to a minor extent and without compromising activation by IFN-α (~14-fold total suppression).
TRIM5α is an E3 ubiquitin (Ub) ligase that is polyubiquitinated by cellular E2 Ub-conjugating enzymes as a prerequisite for suppressing reverse transcription and infection17,18,19. Consistent with previous results20, the integrity of TRIM5α’s ligase domain was confirmed as being necessary for viral inhibition through its inactivation by the H43Y mutation (Fig. 2b). Next, to determine the effects of IFN-α on TRIM5α ubiquitination and ensuing proteasome-mediated degradation, cells were cotransfected with vectors expressing FLAG-tagged TRIM5α and haemagglutinin (HA)-tagged Ub, and whole-cell lysates or immunoprecipitated FLAG-tagged proteins were analysed by immunoblotting (Fig. 3a). IFN-α did not substantially affect global levels of protein ubiquitination (lanes 1–8), but did induce a marked decrease in the level of ubiquitinated TRIM5α (lanes 10 and 14). This decrease was reversed by addition of the broadly acting proteasome inhibitor MG132 or the immunoproteasome-specific inhibitor ONX-0914 (lanes 15 and 16, respectively).
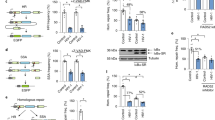
a, U87-MG CD4+ CXCR4+ cells were cotransfected with a vector expressing HA-Ub37 and pFLAG-TRIM5α or a control and treated with or without 500 U ml−1 IFN-α for 24 h in the presence of DMSO (negative control), the proteasome inhibitor MG132 (0.2 µM) or the immunoproteasome inhibitor ONX-0914 (1 µM). Whole-cell lysates and immunoprecipitated TRIM5α were analysed by immunoblotting using an anti-HA (Ub) antibody. One representative immunoblot from three independent experiments is shown. b, U87-MG CD4+ CXCR4+ cells were cotransfected with pFLAG‐TRIM5α and pFLAG-GFP. At 24 h after transfection, cells were treated with or without 500 U ml−1 IFN-α for 24 h in the presence of DMSO (n = 6), MG132 (n = 6) or ONX-0914 (n = 5), lysed and analysed by immunoblotting using antibodies specific for TRIM5α or FLAG. α-Tubulin served as a loading control. The bar graphs represent the relative band intensities for each protein normalized to α-tubulin. c, U87-MG CD4+ CXCR4+ cells were cultured with or without 500 U ml−1 IFN-α, then with DMSO (n = 8), MG132 (n = 4 without IFN-α and n = 8 with IFN-α) or ONX-0914 (n = 4 without IFN-α and n = 7 with IFN-α) for 24 h, and endogenous TRIM5α levels were determined by immunoblotting. α-Tubulin served as a loading control. The bar graph represents the relative band intensities for TRIM5α normalized to α-tubulin. d, NL4-3/Nef-IRES-GFP infectivity and IFN-α induced inhibition at 48 h postinfection in U87-MG CD4+ CXCR4+ cells with or without added 500 U ml−1 IFN-α for 24 h before infection in the presence of DMSO, MG132 or ONX-0914 (n = 4). e, NL4-3/Nef-IRES-GFP infectivity and IFN-α-induced inhibition at 48 h postinfection in U87-MG CD4+ CXCR4+ cells expressing an unrelated guide RNA (CRISPR control) or specific guide RNAs targeting TRIM5α (CRISPR TRIM5α; bulk Fig. 1d) treated with or without 500 U ml−1 IFN-α in the presence of DMSO, MG132 or ONX-0914 before infection (n = 5). Data are represented as the mean ± s.d. P values (95% confidence interval) were determined using two-sided unpaired t-tests.
Confirmation that IFN-α specifically promotes the proteolytic turnover of TRIM5α was obtained in three ways. First, analysis of cells co-expressing FLAG-tagged TRIM5α and FLAG-tagged GFP as a control showed that IFN-α induced the marked loss of TRIM5α, and that this was blocked by MG132 or ONX-0914 (Fig. 3b). Second, levels of endogenous TRIM5α in IFN-α-treated cells were increased by adding MG132 or ONX-0914 (Fig. 3c; of note, because TRIM5α is an ISG itself, its levels were modestly increased by IFN-α addition compared with untreated cells21, despite its accelerated turnover). Third, metabolic-labelling and pulse-chase analysis of FLAG-TRIM5α in transfected cells demonstrated a shortening in initial half-life from ~100 to ~70 min (Supplementary Fig. 9). As expected for proteasome-mediated degradation22, blocking the formation of lysine-48-linked polyubiquitin chains by co-expression of the K48R Ub mutant inhibited IFN-α-induced TRIM5α proteolysis (Supplementary Fig. 10).
The importance of proteasomal activity for IFN-α-mediated suppression was confirmed in viral challenge experiments, where both inhibitors substantially reversed the inhibitory influence of IFN-α (approximately threefold rescue; Fig. 3d and Supplementary Fig. 11). This effect was shown to be dependent on TRIM5α function because, in the presence of IFN-α, HIV-1 infection of cells lacking endogenous TRIM5α was not improved by MG132 or ONX-0914 (Fig. 3e and Supplementary Fig. 12).
The immunoproteasome is predominantly expressed in cells of haematopoietic origin in response to inflammatory cytokines such as IFN, and differs from the constitutive proteasome with respect to the identity of its three proteolytic β-subunits and dependence on the PA28 regulatory complex11,12,23 (Fig. 4a). Its functional divergence from the constitutive proteasome is incompletely understood, although roles in major histocompatibility complex class I antigen processing, T cell differentiation and cytokine modulation have been described24,25. In light of the ability of ONX-0914, a selective inhibitor of β5i/PSMB8, to block IFN-α-induced TRIM5α function (Fig. 3e), the expression patterns of the immunoproteasome β-subunits and the PA28A/PA28B subunits of the PA28 regulatory complex were analysed by immunoblotting (Fig. 4b). With some variation, all five proteins accumulated to higher levels in the presence of IFN-α.
a, On IFN-α stimulation, the catalytic subunits (β1, β2 and β3) of the constitutive proteasome are replaced by inducible catalytic subunits (β1i, β2i and β5i), and expression of the proteasome activator PA28 subunits A and B is induced. b, Protein levels of immunoproteasome components and PA28 subunits in U87-MG CD4+ CXCR4+ cells after siRNA transfection with or without 500 U ml−1 IFN-α were determined by immunoblotting. α-Tubulin served as a loading control. Immunoblot analysis of the panel was performed once in entirety. c, NL4-3/Nef-IRES-GFP infectivity and IFN-α-induced inhibition at 48 h postinfection in U87-MG CD4+ CXCR4+ cells after siRNA-mediated silencing of the indicated genes with or without addition of 500 U ml−1 IFN-α 24 h before infection (n = 3). d, NL4-3/Nef-IRES-GFP infectivity and IFN-α-induced inhibition at 48 h postinfection after siRNA-mediated silencing of PA28A or TRIM5α in U87-MG CD4+ CXCR4+ cells expressing an unrelated guide RNA (CRISPR control) or specific guide targeting TRIM5α (CRISPR TRIM5α). A total of 500 U ml−1 IFN-α was added for 24 h before infection (n = 9). Numbers represent changes in the folds of inhibition. Data are represented as the mean ± s.d. P values (95% confidence interval) were determined using two-sided unpaired t-tests.
These five immunoproteasome components, as well as TRIM5α, were then individually silenced using siRNA (Fig. 4b), and the effects on IFN-α-induced viral suppression determined (Fig. 4c and Supplementary Fig. 13). All knockdowns conferred a degree of relief from inhibition by IFN-α, with PA28A silencing being the most effective to the extent that the magnitude of rescue matched that observed with silencing of TRIM5α itself. Finally, the functional interdependence of the IFN-α-activated anti-HIV-1 phenotypes of human TRIM5α and the immunoproteasome was established by the substantial diminution in the level of rescue from IFN-α inhibition that was observed after PA28A silencing in cells that lacked TRIM5α (Fig. 4d and Supplementary Fig. 14).
TRIM5α restriction has been widely perceived to be important for preventing zoonotic retroviral infections, whereas being ineffective in controlling viruses in their natural hosts2,9. Although there have been sporadic reports of human TRIM5α affecting HIV-1 infection either by suppression of certain HLA-associated CTL escape mutant viruses26 or by inducing autophagy in Langerhans cells27, our findings demonstrate broad, nonstrain-specific inhibition of HIV-1 infection by human TRIM5α. Importantly, we have shown that TRIM5α function is operative in CD4+ T cells and is dependent on IFN-α and activation of the immunoproteasome. Given that IFN levels are elevated during the acute and chronic phases of natural HIV-1 infection4, we surmise that TRIM5α contributes to the immune control of HIV-1 in infected humans, a conclusion consistent with noted associations between favourable clinical outcomes and elevated TRIM5α expression28 or specific TRIM5α alleles29,30. Finally, these results further indicate that the functionality of a specific protein can be fine-tuned by the proteasomal landscape of a cell.
Methods
HIV-1 molecular clones and retroviral vectors
The NL4-3/Nef-IRES-Renilla and NL4-3/Nef-IRES-GFP reporter viruses, the HIV-1-based lentiviral vectors containing WTCA or CA from SIVMAC (SIVCA), and the infectious provirus molecular clones HIV-1 NL4.3, THRO, WITO, CH106.c, REJO.c, SUMA, pAD17, WARO, R66201 and IDU034.2 have been described previously.31,32,33,34,35
Cells
U87-MG CD4+ CXCR4+ cells, which stably express the CD4 and CXCR4 receptors, have been described6, and 293T cells were obtained from the American Type Culture Collection. All cell lines were maintained in complete DMEM-GlutaMAX (Gibco) supplemented with 10% foetal bovine serum (FBS; Gibco), 100 U ml−1 penicillin and 100 g ml−1 streptomycin. Human primary CD4+ T cells were isolated from peripheral blood mononuclear cells of healthy donors (approved by the Guy’s Research Ethics Committee, Ref 03/02/06) obtained by density gradient centrifugation through LymphoPrep (Axis-Shield) and isolated by negative selection using the CD4+ T Cell Isolation Kit (Miltenyi Biotec) according to the manufacturer’s instructions. Cells were activated using Dynabeads Human T-Activator CD3/CD28 (Thermo Fisher) and 50 U ml−1 recombinant interleukin-2 (rIL-2) (Roche) for 48 h in RPMI 1640-GlutaMAX medium containing 10% heat-inactivated autologous serum, 100 U ml−1 penicillin and 100 U ml−1 streptomycin. Cells were maintained after activation in medium containing 30 U ml−1 IL-2. Cells were treated with IFN-α-2b (INTRON A; Merck, Sharpe & Dohme Corp.) for 24 h before infection. U87-MG CD4+ CXCR4+ cells were treated with dimethylsulfoxide (DMSO; Sigma), the proteasome inhibitor MG132 (0.2 µM; ab141003; Abcam) or the immunoproteasome inhibitor ONX-0914 (1 µM; A4011; ApexBio).
siRNA library screen
To identify ISGs that modulate HIV-1 infection, we performed an ISGs siRNA screen using a custom siRNA library (Dharmacon) containing 600 siRNA targets (Supplementary Table 1). U87-MG CD4+ CXCR4+ cells were plated at 1.2 × 104 per well in a 96-well plate and were reverse transfected with siRNA (10 nM). Twenty-four hours later, cells were treated with or without IFN-α (500 U ml−1), and after 12-h incubation, a second siRNA transfection was performed. At 24 h after IFN-α treatment, cells were challenged with NL4-3/Nef-IRES-Renilla (15 ng p24Gag), and infectivity was determined 48 h later by measuring Renilla activity.
Plasmids
pEasiLV plasmids expressing luciferase, CD8, MX2 or TRIMCYP have been described previously6. cDNAs encoding TRIM5α and TRIM5α isoform X7 lacking the SPRY domain were amplified using the SuperScript III One-Step RT–PCR System with Platinum Taq polymerase (Invitrogen) from 20 ng RNA obtained from IFN-α-treated (500 U ml−1) U87-MG CD4+ CXCR4+ cells and inserted into pEasiLV-MCS6 between the AgeI and XhoI restriction sites. The TRIM5α R332G/R335G and H43Y mutants were obtained by site-directed mutagenesis (SDM). CRISPR-resistant TRIM5α constructs (rTRIM5α) were produced by SDM using primers designed to silently mutate the guide RNA target sequence. pFLAG-TRIM5α was generated by subcloning a cDNA encoding TRIM5α with an amino-terminal FLAG tag into pCAGGS (Addgene) using the EcoRI and XhoI sites. pCAGGS-expressing FLAG-tagged GFP, pFLAG-GFP, was derived from a plasmid encoding HA-tagged GFP36. The vectors encoding HA-tagged Ub and the K48R and K63R mutant derivatives have been described previously.37,38
Knockout cells
To generate CRISPR–Cas9 cells, we cloned sequences encoding specific guide RNAs targeting TRIM5α, MX2 or red fluorescent protein (RFP), as a control, into BsmBI-linearized lentiviral vector pLentiCRISPRv239,40 using the oligonucleotides (forward and reverse) 5′-caccgCCTCCTCCTTTACATTAACC-3′ and 5′-aaacGGTTAATGTAAAGGAGGAGGc-3′ for TRIM5α, 5′-caccgGACAACCAGCCCCGAGACAT-3′ and 5′-aaacATGTCTCGGGGCTGGTTGTCc-3′ for MX2 or 5′-caccgCTCAGTTCCAGTACGGCTCCA-3′ and 5′-aaacTGGAGCCGTACTGGAACTGAGc-3′ for RFP. Vectors expressing the indicated guide RNAs were produced by cotransfection of 293T cells with pLentiCRISPRv2, p8.91 and vesicular stomatitis virus G protein (VSV-G) vectors at a ratio of 1.5:1.5:0.375, respectively. U87-MG CD4+ CXCR4+ cells were transduced with the TRIM5α targeting or RFP control lentiviral vectors and selected using 1 μg ml−1 puromycin for 3 days. Single-cell clones (TRIM5α_1 and TRIM5α_2) were derived from the bulk (TRIM5α) population by single-cell sorting in 96-well plates and assayed for the loss of TRIM5α by immunoblotting. TRIM5α gene disruption was validated by PCR amplification and sequencing of the targeted genomic region. To generate TRIM5α/MX2 double-knockout cells, we further transduced the TRIM5α_1 line (Fig. 1d) with a high concentration of the MX2 targeting vector, and the loss of MX2 expression in the bulk population was confirmed by immunoblotting.
Viral production
All viruses were generated by transfection of 10-cm 293T cultures using TransIT-2020 transfection reagent (Mirus); the medium was changed after 6 h, and virus-containing medium was filtered and collected 48 h after transfection. NL4-3/Nef-IRES-GFP and NL4-3/Nef-IRES-Renilla viral particles were produced by transfection of 10 µg of proviral plasmid. For analysis of HIV-1 reverse transcription products and shRNA-mediated TRIM5α silencing, virus-containing supernatants were DNase (RQ1 RNase free DNase; Promega) treated for 1 h at 37 °C, and viruses were purified through a 20% (weight/volume) sucrose cushion at 145,370g for 75 min at 4 °C, resuspended in PBS and stored in aliquots at −80 °C. Lentiviral vectors stocks were produced by cotransfection of 293T cells with plasmids encoding Gag-Pol (p8.91 or p8.91-SIVCA), a mini viral genome bearing a cytomegalovirus (CMV)-immediate early GFP expression cassette (pRRL.sin.cPPT.CMV/eGFP), and the VSV-G envelope (pMD.G), at a ratio of 4:4:2, respectively3. To generate GFP-encoding derivatives of full-length molecular clones NL4.3, THRO, WITO, CH106.c, REJO.c, SUMA, pAD17, WARO, R66201 and IDU034.2, we cotransfected provirus plasmids with the GFP reporter vector pCSGW and VSV-G at a ratio of 4.5:3:3, respectively41. Virus titres were determined by challenging U87-MG CD4+ CXCR4+ cells with different viral dilutions and by quantifying the number of GFP+ cells after 48 h by flow cytometry (FACSCalibur; BD Biosciences; a typical example is displayed in Supplementary Fig. 15), and p24Gag content was quantified using an HIV-1 p24 ELISA detection kit (Perkin-Elmer). To produce EasiLV particles, we cotransfected 293T cells with pEasiLV, p8.91, pptTRKrab and VSV-G at a ratio of 1:1:0.5:0.25, respectively6. At 48 h after transfection, viral particles were harvested, filtered and used directly to transduce U87-MG CD4+ CXCR4+ cells. Six hours after transduction, the medium was replaced with fresh medium containing 0.5 µg ml−1 doxycycline (Sigma) to induce transgene expression. At 72 h after transduction, the percentage of E2-Crimson-positive cells was scored (typically > 80%) by flow cytometry.
shRNA-mediated TRIM5α silencing
TRIM5α silencing in primary human CD4+ T cells was achieved using a modified version of the HIV-1-based lentiviral vector pHRSIREN-S-SBP-∆LNGFR-W42, where the selectable marker for antibody-free magnetic cell sorting sequence (SBP-∆LNGFR) was replaced by the E2-Crimson fluorescent reporter gene for flow cytometry analysis, generating the pHRSIREN-S-E2-W vector. The shRNA targeting sequence for TRIM5α43 (shTRIM5) was 5′-TGGCTTCTGGAATCCTGGTTAA-3′, and the scrambled shRNA sequence42 (shControl) used as negative control was 5′-GTTATAGGCTCGCAAAAGG-3′. Lentiviral vectors were produced by cotransfection of 293T cells with p8.91, VSV-G and the pHRSIREN-S-E2-W vectors at a ratio of 1:0.5:1.5, respectively. Lentiviral particles were concentrated by ultracentrifugation before use for transduction of primary human CD4+ T cells at 2,000g for 2 h. At 48 h after transduction, E2-crimson-positive cells were sorted by flow cytometry (typically > 90%) and cultured in fresh medium. When indicated, 2.5 × 104 cells per well in 96-well plates were treated with IFN-α (2,000 U ml−1) for 24 h before infection with NL4-3/Nef-IRES-Renilla (30 ng p24Gag) at 2,000g for 2 h, and infectivity was measured 48 h after infection by measuring Renilla activity.
Quantification of mRNA expression
A total of 106 activated primary human CD4+ T cells were collected with or without 24-h treatment with IFN-α, and total RNA was extracted using the RNeasy Mini Kit (Qiagen). cDNA was generated using 500 ng RNA and the High-Capacity cDNA Reverse Transcription Kit (Thermo Fisher), and qPCR of selected genes was performed using the following TaqMan gene expression assays (Thermo Fisher): TRIM5α (Hs01552559_m1), MX2 (Hs01550808_m1) and GAPDH (Hs99999905_m1). Expression levels of target genes were normalized to GAPDH.
Measurement of HIV-1 reverse transcription products
A total of 105 U87-MG CD4+ CXCR4+ were reverse transfected with 10 nM of a SMARTpool siRNA targeting human TRIM5α or a nontargeting siRNA (Dharmacon). Cells were treated with or without 500 U ml−1 IFN-α and transfected again 12 h after treatment. At 24 h after IFN-α treatment, cells were challenged with NL4-3/Nef-IRES-GFP (20 ng p24Gag) for 2 and 48 h after infection; cells were collected, and total DNA extraction was performed using the DNeasy kit (Qiagen). Strong-stop cDNA products were detected using primers that amplify the regions between nucleotides 500 and 635 of the provirus: oHC64 (5′-TAACTAGGGAACCCACTGC-3′) and oHC65 (5′-GCTAGAGATTTTCCACACTG-3′) with probe oHC66 (5′-FAM-ACACAACAGACGGGCACACACTA-TAMRA-3′), where FAM is 6-carboxyfluorescein and TAMRA is 6-carboxytetramethylrhodamine. qPCR reactions were performed in triplicate using TaqMan Universal PCR master mix, 900 nM of each primer and 250 nM of probe. After 10 min at 95 °C, reactions were cycled through 15 s at 95 °C, followed by 1 min at 60 °C for 40 repeats, carried out using the QuantStudio 5 Real-Time PCR System (Applied Biosystems). NL4-3/Nef-IRES-GFP plasmid was diluted in salmon sperm DNA solution (20 ng ml−1) to create dilution standards to calculate relative cDNA copy numbers and confirm the linearity of the assays.
siRNA-mediated knockdown
U87-MG CD4+ CXCR4+ cells were plated at 1.2 × 104 per well in a 96-well plate and were reverse transfected with 10 nM SMARTpool siRNAs (Dharmacon) targeting TRIM5α (M-007100-00-0005), PSMB8 (M-006022-01-0005), PSMB9 (M-006023-02-0005), PSMB10 (M-006019-01-0005), PA28A (M-012254-00-0005), PA28B (M-011370-00-0005) or individual siRNAs targeting TRIM5α (D-007100-01-0002, D-007100-02-0002, D-007100-03-0002, D-007100-04-0002). A nontargeting siRNA (D-001210-03-05) was used as a negative control. At 24 h after the first transfection, cells were treated with or without IFN-α (500 U ml−1), and a second siRNA transfection was performed 12 h after treatment. At 24 h after IFN-α treatment, cells were challenged with the indicated viruses, and infectivity was determined 48 h later. To confirm knockdown efficiency, cell lysates were harvested 24 h after IFN-α treatment and were subjected to SDS–PAGE and immunoblot assays.
Immunoblotting
Cells were washed with PBS, lysed in sample buffer, boiled for 3 min, resolved by SDS–PAGE and analysed by immunoblotting using primary antibodies specific for human MX2 (sc-47197 (N-17); Santa Cruz Biotechnology), human TRIM5α (ab4389; Abcam), human PSMB8 (ab3329; Abcam), human PSMB9 (ab3328; Abcam), human PSMB10 (ab77735; Abcam), human PA28A (ab155091; Abcam), human PA28B (ab183727; Abcam), FLAG (horseradish peroxidase (HRP)-conjugated M2; Sigma), HA (HRP-conjugated 3F10; Sigma) or human α-tubulin (DM1A; Sigma), and detected using either HRP-conjugated secondary antibodies and chemiluminescence (ECL+ western blotting substrate; Pierce) or IRDye-800CW-labelled secondary antibodies and the LI-COR infrared imaging technology (LI-COR UK Ltd.).
Immunoprecipitation
A total of 3 × 106 U87-MG CD4+ CXCR4+ cells were seeded in 10-cm tissue culture plates and transfected with the indicated plasmids expressing FLAG-TRIM5α and/or HA-Ub at a ratio of 1:0.5, using TransIT-2020 transfection reagent according to the manufacturer’s instructions. After 24 h, indicated cells were treated with fresh medium containing or not IFN-α (500 U ml−1) and DMSO, MG132 (0.2 µM) or ONX-0914 (1 µM). At 24 h after treatment, cells were washed twice with PBS, harvested with lysis buffer (1× PBS, 0.5% Triton X-100 and protease inhibitor cocktail; Roche) and disrupted by brief sonication. Lysates were cleared by centrifugation at 10,000g for 10 min at 4 °C, FLAG-TRIM5α was immunoprecipitated using anti-FLAG M2 magnetic beads (M8823-5ML; Sigma) for 2 h at 4 °C, and the beads were washed a further four times in lysis buffer before adding sample buffer (200 mM Tris-HCl, pH 6.8, 5.2% SDS, 20% glycerol, 0.1% bromophenol blue, 5% β-mercaptoethanol). HA-tagged proteins (ubiquitinated proteins) were resolved by SDS–PAGE and detected by immunoblotting using HRP-conjugated anti-HA antibodies.
S35 pulse-chase labelling and radio-immunoprecipitation
U87-MG CD4+ CXCR4+ cells were seeded at 2 × 105 per well in six-well plates and transfected with pFLAG-TRIM5α using TransIT-2020 transfection reagent according to the manufacturer’s instructions. After 24 h, indicated cells were treated with fresh medium containing or not IFN-α (500 U ml−1). Twenty-four hours later, cells were washed twice with PBS and incubated for 20 min in cysteine-methionine-depleted DMEM (Gibco) at 37 °C. The medium was replaced by depletion medium containing 0.25 mCi ml−1 S35-labelled cysteine-methionine, and the cells maintained for 10 min at 37 °C. Cells were then washed twice with PBS and incubated in DMEM. The cells were harvested at various times with lysis buffer (1× PBS, 0.5% Triton X-100 and protease inhibitor cocktail), incubated on ice for 10 min and clarified by centrifugation at 10,000g for 10 min at 4 °C. FLAG-TRIM5α was immunoprecipitated using anti-FLAG M2 magnetic beads, resolved by SDS–PAGE and exposed on a phosphor image screen over several days before development on a Typhoon Trio PhosphorImager (GE Healthcare).
Statistical analysis
Results in bar charts are expressed as means ± s.d. for experimental replicates in each case. Differences between the experimental groups were evaluated, where indicated, by paired or unpaired two-tailed t-tests.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available from the corresponding author upon request.
References
Schoggins, J. W. et al. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 472, 481–485 (2011).
Malim, M. H. & Bieniasz, P. D. HIV restriction factors and mechanisms of evasion. Cold Spring Harb. Perspect. Med. 2, a006940 (2012).
Goujon, C. & Malim, M. H. Characterization of the alpha interferon-induced postentry block to HIV-1 infection in primary human macrophages and T cells. J. Virol. 84, 9254–9266 (2010).
Doyle, T., Goujon, C. & Malim, M. H. HIV-1 and interferons: who’s interfering with whom? Nat. Rev. Microbiol. 13, 403–413 (2015).
Cheney, K. M. & McKnight, A. Interferon-alpha mediates restriction of human immunodeficiency virus type-1 replication in primary human macrophages at an early stage of replication. PLoS ONE 5, e13521 (2010).
Goujon, C. et al. Human MX2 is an interferon-induced post-entry inhibitor of HIV-1 infection. Nature 502, 559–562 (2013).
Kane, M. et al. MX2 is an interferon-induced inhibitor of HIV-1 infection. Nature 502, 563–566 (2013).
Liu, Z. et al. The interferon-inducible MxB protein inhibits HIV-1 infection. Cell Host Microbe 14, 398–410 (2013).
Stremlau, M. et al. The cytoplasmic body component TRIM5alpha restricts HIV-1 infection in Old World monkeys. Nature 427, 848–853 (2004).
Sayah, D. M., Sokolskaja, E., Berthoux, L. & Luban, J. Cyclophilin A retrotransposition into TRIM5 explains owl monkey resistance to HIV-1. Nature 430, 569–573 (2004).
Akiyama, K. et al. Replacement of proteasome subunits X and Y by LMP7 and LMP2 induced by interferon-gamma for acquirement of the functional diversity responsible for antigen processing. FEBS Lett. 343, 85–88 (1994).
Ma, C. P., Slaughter, C. A. & DeMartino, G. N. Identification, purification, and characterization of a protein activator (PA28) of the 20 S proteasome (macropain). J. Biol. Chem. 267, 10515–10523 (1992).
Groettrup, M., Kirk, C. J. & Basler, M. Proteasomes in immune cells: more than peptide producers? Nat. Rev. Immunol. 10, 73–78 (2010).
Wagner, J. M. et al. General model for retroviral capsid pattern recognition by trim5 proteins. J. Virol. 92, e01563-17 (2018).
Pham, Q. T., Bouchard, A., Grutter, M. G. & Berthoux, L. Generation of human TRIM5alpha mutants with high HIV-1 restriction activity. Gene Ther. 17, 859–871 (2010).
Yap, M. W., Nisole, S. & Stoye, J. P. A single amino acid change in the SPRY domain of human Trim5alpha leads to HIV-1 restriction. Curr. Biol. 15, 73–78 (2005).
Fletcher, A. J. et al. TRIM5alpha requires Ube2W to anchor Lys63-linked ubiquitin chains and restrict reverse transcription. EMBO J. 34, 2078–2095 (2015).
Campbell, E. M. et al. TRIM5alpha-mediated ubiquitin chain conjugation is required for inhibition of hiv-1 reverse transcription and capsid destabilization. J. Virol. 90, 1849–1857 (2016).
Diaz-Griffero, F. et al. Rapid turnover and polyubiquitylation of the retroviral restriction factor TRIM5. Virol. 349, 300–315 (2006).
Sawyer, S. L., Wu, L. I., Akey, J. M., Emerman, M. & Malik, H. S. High-frequency persistence of an impaired allele of the retroviral defense gene TRIM5alpha in humans. Curr. Biol. 16, 95–100 (2006).
Carthagena, L. et al. Human TRIM gene expression in response to interferons. PLoS ONE 4, e4894 (2009).
Komander, D. & Rape, M. The ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012).
Preckel, T. et al. Impaired immunoproteasome assembly and immune responses in PA28-/- mice. Science 286, 2162–2165 (1999).
Kloetzel, P. M. Antigen processing by the proteasome. Nat. Rev. Mol. Cell Biol. 2, 179–187 (2001).
Muchamuel, T. et al. A selective inhibitor of the immunoproteasome subunit LMP7 blocks cytokine production and attenuates progression of experimental arthritis. Nat. Med. 15, 781–787 (2009).
Granier, C. et al. Pressure from TRIM5alpha contributes to control of HIV-1 replication by individuals expressing protective HLA-B alleles. J. Virol. 87, 10368–10380 (2013).
Ribeiro, C. M. et al. Receptor usage dictates HIV-1 restriction by human TRIM5alpha in dendritic cell subsets. Nature 540, 448–452 (2016).
Sewram, S. et al. Human TRIM5alpha expression levels and reduced susceptibility to HIV-1 infection. J. Infect. Dis. 199, 1657–1663 (2009).
Celerino da Silva, R. et al. TRIM5 gene polymorphisms in HIV-1-infected patients and healthy controls from Northeastern Brazil. Immunol. Res. 64, 1237–1242 (2016).
vanManen, D. et al. The effect of Trim5 polymorphisms on the clinical course of HIV-1 infection. PLoS Pathog. 4, e18 (2008).
Schindler, M. et al. Down-modulation of mature major histocompatibility complex class II and up-regulation of invariant chain cell surface expression are well-conserved functions of human and simian immunodeficiency virus nef alleles. J. Virol. 77, 10548–10556 (2003).
Naldini, L. et al. In vivo gene delivery and stable transduction of nondividing cells by a lentiviral vector. Science 272, 263–267 (1996).
Cordeil, S. et al. Evidence for a different susceptibility of primate lentiviruses to type I interferons. J. Virol. 87, 2587–2596 (2013).
Ochsenbauer, C. et al. Generation of transmitted/founder HIV-1 infectious molecular clones and characterization of their replication capacity in CD4 T lymphocytes and monocyte-derived macrophages. J. Virol. 86, 2715–2728 (2012).
Parrish, N. F. et al. Phenotypic properties of transmitted founder HIV-1. Proc. Natl Acad. Sci. USA 110, 6626–6633 (2013).
Dicks, M. D. et al. Oligomerization requirements for MX2-mediated suppression of HIV-1 infection. J. Virol. 90, 22–32 (2015).
Treier, M., Staszewski, L. M. & Bohmann, D. Ubiquitin-dependent c-Jun degradation in vivo is mediated by the delta domain. Cell 78, 787–798 (1994).
Dart, A. E. et al. PAK4 promotes kinase-independent stabilization of RhoU to modulate cell adhesion. J. Cell. Biol. 211, 863–879 (2015).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Schaller, T. et al. Effects of inner nuclear membrane proteins SUN1/UNC-84A and SUN2/UNC-84B on the EARLY steps of HIV-1 infection. J. Virol. 91, e00463-17 (2017).
Matheson, N. J., Peden, A. A. & Lehner, P. J. Antibody-free magnetic cell sorting of genetically modified primary human CD4+T cells by one-step streptavidin affinity purification. PLoS ONE 9, e111437 (2014).
Pertel, T. et al. TRIM5 is an innate immune sensor for the retrovirus capsid lattice. Nature 472, 361–365 (2011).
Acknowledgements
We thank D. Pollpeter, M. Dicks, S. Papaioannou, C. Wells and S. Wolinsky for the generous provision of reagents and helpful discussions. The work was supported by the UK Medical Research Council (grant no. G1000196), the Wellcome Trust (grant no. 106223/Z/14/Z), and the Department of Health via a National Institute for Health Research comprehensive Biomedical Research Centre award to Guy’s and St. Thomas’ NHS Foundation Trust in partnership with King’s College London and King’s College Hospital NHS Foundation Trust. J.M.J.-G. is a Long-Term Fellow of the European Molecular Biology Organization (EMBO) (ALTF 663-2016).
Author information
Authors and Affiliations
Contributions
J.M.J.-G., L.A., and M.H.M. conceived the siRNA screen. J.M.J.-G. and M.H.M. designed the study and wrote the manuscript with input from all co-authors. J.M.J.-G. carried out the experiments and analysed the data. L.A. and G.B. contributed to the execution of experiments and provided reagents. M.H.M. supervised all aspects of the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–24 and Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Jimenez-Guardeño, J.M., Apolonia, L., Betancor, G. et al. Immunoproteasome activation enables human TRIM5α restriction of HIV-1. Nat Microbiol 4, 933–940 (2019). https://doi.org/10.1038/s41564-019-0402-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0402-0
This article is cited by
-
Autophagy-enhancing ATG16L1 polymorphism is associated with improved clinical outcome and T-cell immunity in chronic HIV-1 infection
Nature Communications (2024)
-
TRIM5α restricts poxviruses and is antagonized by CypA and the viral protein C6
Nature (2023)
-
TRIM5α recruits HDAC1 to p50 and Sp1 and promotes H3K9 deacetylation at the HIV-1 LTR
Nature Communications (2023)
-
HIV-1 capsid variability: viral exploitation and evasion of capsid-binding molecules
Retrovirology (2021)
-
MX2-mediated innate immunity against HIV-1 is regulated by serine phosphorylation
Nature Microbiology (2021)