Abstract
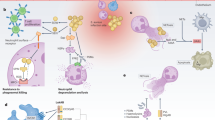
Most clonal lineages of Staphylococcus epidermidis are commensals present on human skin and in the nose. However, some globally spreading healthcare-associated and methicillin-resistant S. epidermidis (HA-MRSE) clones are major causes of difficult-to-treat implant or bloodstream infections. The molecular determinants that alter the lifestyle of S. epidermidis have remained elusive, and their identification might provide therapeutic targets. We reasoned that changes in surface-exposed wall teichoic acid (WTA) polymers of S. epidermidis, which potentially shape host interactions, may be linked to differences between colonization and infection abilities of different clones. We used a combined epidemiological and functional approach to show that while commensal clones express poly-glycerolphosphate WTA, S. epidermidis multilocus sequence type 23, which emerged in the past 15 years and is one of the main infection-causing HA-MRSE clones, contains an accessory genetic element, tarIJLM, that leads to the production of a second, Staphylococcus aureus-type WTA (poly-ribitolphosphate (RboP)). Production of RboP-WTA by S. epidermidis impaired in vivo colonization but augmented endothelial attachment and host mortality in a mouse sepsis model. tarIJLM was absent from commensal human sequence types but was found in several other HA-MRSE clones. Moreover, RboP-WTA enabled S. epidermidis to exchange DNA with S. aureus via siphovirus bacteriophages, thereby creating a possible route for the inter-species exchange of methicillin resistance, virulence and colonization factors. We conclude that tarIJLM alters the lifestyle of S. epidermidis from commensal to pathogenic and propose that RboP-WTA might be a robust target for preventive and therapeutic interventions against MRSE infections.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The whole-genome sequence data from this study were deposited in the NCBI Sequence Read Archive; the accession numbers can be found in Supplementary Table 1. Source data are provided with this paper. Additional information can be obtained from the corresponding author upon reasonable request.
References
Becker, K., Heilmann, C. & Peters, G. Coagulase-negative staphylococci. Clin. Microbiol. Rev. 27, 870–926 (2014).
Byrd, A. L., Belkaid, Y. & Segre, J. A. The human skin microbiome. Nat. Rev. Microbiol. 16, 143–155 (2018).
Lee, J. Y. H. et al. Global spread of three multidrug-resistant lineages of Staphylococcus epidermidis. Nat. Microbiol. 3, 1175–1185 (2018).
Li, M., Wang, X., Gao, Q. & Lu, Y. Molecular characterization of Staphylococcus epidermidis strains isolated from a teaching hospital in Shanghai, China. J. Med. Microbiol. 58, 456–461 (2009).
Meric, G. et al. Disease-associated genotypes of the commensal skin bacterium Staphylococcus epidermidis. Nat. Commun. 9, 5034 (2018).
Heilmann, C., Ziebuhr, W. & Becker, K. Are coagulase-negative staphylococci virulent?. Clin. Microbiol. Infect. 25, 1071–1080 (2019).
Weidenmaier, C. & Peschel, A. Teichoic acids and related cell-wall glycopolymers in Gram-positive physiology and host interactions. Nat. Rev. Microbiol. 6, 276–287 (2008).
Moller, A. G., Lindsay, J. A. & Read, T. D. Determinants of phage host range in Staphylococcus species. Appl. Environ. Microbiol. 85, e00209-19 (2019).
Chen, J. et al. Genome hypermobility by lateral transduction. Science 362, 207–212 (2018).
Haaber, J., Penades, J. R. & Ingmer, H. Transfer of antibiotic resistance in Staphylococcus aureus. Trends Microbiol. 25, 893–905 (2017).
Lee, A. S. et al. Methicillin-resistant Staphylococcus aureus. Nat. Rev. Dis. Primers 4, 18033 (2018).
Penades, J. R. & Christie, G. E. The phage-inducible chromosomal islands: a family of highly evolved molecular parasites. Annu. Rev. Virol. 2, 181–201 (2015).
Meric, G. et al. Ecological overlap and horizontal gene transfer in Staphylococcus aureus and Staphylococcus epidermidis. Genome Biol. Evol. 7, 1313–1328 (2015).
Li, M. et al. MRSA epidemic linked to a quickly spreading colonization and virulence determinant. Nat. Med. 18, 816–819 (2012).
Winstel, V. et al. Wall teichoic acid structure governs horizontal gene transfer between major bacterial pathogens. Nat. Commun. 4, 2345 (2013).
Fitzgerald, J. R. et al. Characterization of a putative pathogenicity island from bovine Staphylococcus aureus encoding multiple superantigens. J. Bacteriol. 183, 63–70 (2001).
Maiques, E. et al. Role of staphylococcal phage and SaPI integrase in intra- and interspecies SaPI transfer. J. Bacteriol. 189, 5608–5616 (2007).
Brown, S., Santa Maria, J. P. Jr. & Walker, S. Wall teichoic acids of Gram-positive bacteria. Annu. Rev. Microbiol. 67, 313–336 (2013).
Winstel, V., Xia, G. & Peschel, A. Pathways and roles of wall teichoic acid glycosylation in Staphylococcus aureus. Int. J. Med. Microbiol. 304, 215–221 (2014).
Krismer, B. et al. Nutrient limitation governs Staphylococcus aureus metabolism and niche adaptation in the human nose. PLoS Pathog. 10, e1003862 (2014).
Xia, G. et al. Glycosylation of wall teichoic acid in Staphylococcus aureus by TarM. J. Biol. Chem. 285, 13405–13415 (2010).
Dean, B. A., Williams, R. E., Hall, F. & Corse, J. Phage typing of coagulase-negative staphylococci and micrococci. J. Hyg. (Lond.) 71, 261–270 (1973).
Miragaia, M. et al. Comparison of molecular typing methods for characterization of Staphylococcus epidermidis: proposal for clone definition. J. Clin. Microbiol. 46, 118–129 (2008).
Sivadon, V. et al. Partial atlE sequencing of Staphylococcus epidermidis strains from prosthetic joint infections. J. Clin. Microbiol. 47, 2321–2324 (2009).
Mertens, A. & Ghebremedhin, B. Genetic determinants and biofilm formation of clinical Staphylococcus epidermidis isolates from blood cultures and indwelling devises. Eur. J. Microbiol. Immunol. 3, 111–119 (2013).
Sharma, P. et al. Multilocus sequence typing for interpreting blood isolates of Staphylococcus epidermidis. Interdiscip. Perspect. Infect. Dis. 2014, 787458 (2014).
Post, V. et al. Comparative genomics study of Staphylococcus epidermidis isolates from orthopedic-device-related infections correlated with patient outcome. J. Clin. Microbiol. 55, 3089–3103 (2017).
Both, A. et al. Distinct clonal lineages and within-host diversification shape invasive Staphylococcus epidermidis populations. PLoS Pathog. 17, e1009304 (2021).
Weiser, J. et al. Sub-inhibitory tigecycline concentrations induce extracellular matrix binding protein Embp dependent Staphylococcus epidermidis biofilm formation and immune evasion. Int. J. Med. Microbiol. 306, 471–478 (2016).
Wang, L. & Archer, G. L. Roles of CcrA and CcrB in excision and integration of staphylococcal cassette chromosome mec, a Staphylococcus aureus genomic island. J. Bacteriol. 192, 3204–3212 (2010).
Jiang, Y., Oliver, P., Davies, K. E. & Platt, N. Identification and characterization of murine SCARA5, a novel class A scavenger receptor that is expressed by populations of epithelial cells. J. Biol. Chem. 281, 11834–11845 (2006).
Baur, S. et al. A nasal epithelial receptor for Staphylococcus aureus WTA governs adhesion to epithelial cells and modulates nasal colonization. PLoS Pathog. 10, e1004089 (2014).
Widerstrom, M., McCullough, C. A., Coombs, G. W., Monsen, T. & Christiansen, K. J. A multidrug-resistant Staphylococcus epidermidis clone (ST2) is an ongoing cause of hospital-acquired infection in a Western Australian hospital. J. Clin. Microbiol. 50, 2147–2151 (2012).
Ziebuhr, W. et al. Nosocomial infections by Staphylococcus epidermidis: how a commensal bacterium turns into a pathogen. Int. J. Antimicrob. Agents 28, S14–S20 (2006).
Du, X. et al. Molecular analysis of Staphylococcus epidermidis strains isolated from community and hospital environments in China. PLoS ONE 8, e62742 (2013).
Zhang, Y. Q. et al. Genome-based analysis of virulence genes in a non-biofilm-forming Staphylococcus epidermidis strain (ATCC 12228). Mol. Microbiol. 49, 1577–1593 (2003).
Mack, D., Siemssen, N. & Laufs, R. Parallel induction by glucose of adherence and a polysaccharide antigen specific for plastic-adherent Staphylococcus epidermidis: evidence for functional relation to intercellular adhesion. Infect. Immun. 60, 2048–2057 (1992).
Otto, M. Staphylococcal biofilms. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.GPP3-0023-2018 (2018).
Weidenmaier, C. et al. Lack of wall teichoic acids in Staphylococcus aureus leads to reduced interactions with endothelial cells and to attenuated virulence in a rabbit model of endocarditis. J. Infect. Dis. 191, 1771–1777 (2005).
Weidenmaier, C. et al. Role of teichoic acids in Staphylococcus aureus nasal colonization, a major risk factor in nosocomial infections. Nat. Med. 10, 243–245 (2004).
Schade, J. & Weidenmaier, C. Cell wall glycopolymers of Firmicutes and their role as nonprotein adhesins. FEBS Lett. 590, 3758–3771 (2016).
Weidenmaier, C. et al. Differential roles of sortase-anchored surface proteins and wall teichoic acid in Staphylococcus aureus nasal colonization. Int. J. Med. Microbiol. 298, 505–513 (2008).
Foster, T. J. Surface proteins of Staphylococcus epidermidis. Front. Microbiol. 11, 1829 (2020).
Zipperer, A. et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature 535, 511–516 (2016).
Iwase, T. et al. Staphylococcus epidermidis Esp inhibits Staphylococcus aureus biofilm formation and nasal colonization. Nature 465, 346–349 (2010).
Ladner, J. T., Grubaugh, N. D., Pybus, O. G. & Andersen, K. G. Precision epidemiology for infectious disease control. Nat. Med. 25, 206–211 (2019).
Wirth, T. et al. Niche specialization and spread of Staphylococcus capitis involved in neonatal sepsis. Nat. Microbiol. 5, 735–745 (2020).
van Dalen, R. et al. Do not discard Staphylococcus aureus WTA as a vaccine antigen. Nature 572, E1–E2 (2019).
Tormo, M. A. et al. Staphylococcus aureus pathogenicity island DNA is packaged in particles composed of phage proteins. J. Bacteriol. 190, 2434–2440 (2008).
Winstel, V., Kuhner, P., Rohde, H. & Peschel, A. Genetic engineering of untransformable coagulase-negative staphylococcal pathogens. Nat. Protoc. 11, 949–959 (2016).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Thomas, J. C. et al. Improved multilocus sequence typing scheme for Staphylococcus epidermidis. J. Clin. Microbiol. 45, 616–619 (2007).
Kaya, H. et al. SCC mecFinder, a web-based tool for typing of staphylococcal cassette chromosome mec in Staphylococcus aureus using whole-genome sequence data. mSphere https://doi.org/10.1128/mSphere.00612-17 (2018).
Siguier, P., Perochon, J., Lestrade, L., Mahillon, J. & Chandler, M. ISfinder: the reference centre for bacterial insertion sequences. Nucleic Acids Res. 34, D32–D36 (2006).
Sahl, J. W. et al. NASP: an accurate, rapid method for the identification of SNPs in WGS datasets that supports flexible input and output formats. Micro. Genom. 2, e000074 (2016).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Delcher, A. L., Phillippy, A., Carlton, J. & Salzberg, S. L. Fast algorithms for large-scale genome alignment and comparison. Nucleic Acids Res. 30, 2478–2483 (2002).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Arndt, D. et al. PHASTER: a better, faster version of the PHAST phage search tool. Nucleic Acids Res. 44, W16–W21 (2016).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15 (2015).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Guindon, S. & Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Anisimova, M., Gil, M., Dufayard, J. F., Dessimoz, C. & Gascuel, O. Survey of branch support methods demonstrates accuracy, power, and robustness of fast likelihood-based approximation schemes. Syst. Biol. 60, 685–699 (2011).
Gerwig, G. J., Kamerling, J. P. & Vliegenthart, J. F. Determination of the absolute configuration of mono-saccharides in complex carbohydrates by capillary G.L.C. Carbohydr. Res. 77, 10–17 (1979).
Geiger, T. et al. The stringent response of Staphylococcus aureus and its impact on survival after phagocytosis through the induction of intracellular PSMs expression. PLoS Pathog. 8, e1003016 (2012).
Bruckner, R. A series of shuttle vectors for Bacillus subtilis and Escherichia coli. Gene 122, 187–192 (1992).
Wang, L. et al. SarZ is a key regulator of biofilm formation and virulence in Staphylococcus epidermidis. J. Infect. Dis. 197, 1254–1262 (2008).
Kretschmer, D., Rautenberg, M., Linke, D. & Peschel, A. Peptide length and folding state govern the capacity of staphylococcal beta-type phenol-soluble modulins to activate human formyl-peptide receptors 1 or 2. J. Leukoc. Biol. 97, 689–697 (2015).
Acknowledgements
We thank M. Marschal for clinical S. epidermidis strains and strain information; F. Oesterhelt for the help with microscopy; R. Stemmler, P. Kühner and L. Li for phage propagation and WTA isolation; K. Jakob and H. Käßner for technical support with NMR spectroscopy and chemical composition analysis; R. Rosenstein and D. Gerlach for helpful discussions; A. Tooming-Klunderud for PacBio sequencing of strain E73; A. Schneider for help with LC–MS analysis of strain E73 WTA; J. Y. H. Lee for sending us the strain US06; D. A. Robinson for sending us the strain DAR1907; and J. Doškař for sending us the phage ΦPH15. This work was financed by grants from the Deutsche Forschungsgemeinschaft (DFG) SFB766 to C.W., C.M. and A.P.; TRR34 to C.W. and A.P.; TRR156 (project ID 246807620) to A.P.; PE 805/7-1 to A.P.; the German Center of Infection Research to B.K., S.H., C.W. and A.P.; the National Natural Science Foundation of China 81861138043 to M.L.; and the Damp Foundation (HAPDICS project 2013-19) to H.R. B.K., S.H., C.M. and A.P. were supported by infrastructural funding from the Cluster of Excellence EXC 2124 ‘Controlling Microbes to Fight Infections’ (project ID 390838134) from the DFG.
Author information
Authors and Affiliations
Contributions
X.D. conceived the study and performed and analysed most of the bacteriological, molecular and cell culture experiments. J. Larsen performed most of the bioinformatics experiments (with support from M.S.). M.L., A.B. and H.R. collected and typed the S. epidermidis strains. M.L. provided important intellectual support and performed the animal experiments with the help of Y.L. and J. Liu. The WTA structure was analysed via MS by A.W. and C.M. and via NMR microscopy by P.M.S.C. and K.A.D. C.S., E.L., J.S. and C.W. contributed to the cell culture experiments. B.K. and S.H. helped with the design of the cloning experiments. V.W. and A.P. conceived and supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Victor Torres, Wilma Ziebuhr and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phage adsorption assay confirms presence of the RboP-WTA phage receptor on tarIJLM2-positive S. epidermidis isolates.
The binding rate of RboP-WTA specific phage Φ11 (a, c, d, e) and of GroP-WTA specific Φ187 (b) to the indicated strains is shown. RP62A, ATCC12228, and 1457 are S. epidermidis laboratory strains (light gray), only expressing GroP-WTA. S. aureus strains RN4220 or PS187 (dark gray) express RboP-WTA or GroP-WTA, respectively. Means ± s.d. of three independent experiments are shown. One-way ANOVA with Dunnett’s post-test (two-sided) was used to analyze significant differences vs. mutant in (d) and (e). Unpaired t-test was used to analyze significant differences between wt and the hybrid strain in (c).
Extended Data Fig. 2 While tarL2 is efficiently transcribed in the tarIJLM2-positive isolates E6, E45, and E73, tarL1 is not or only very weakly expressed in all tested S. epidermidis strains during growth in human serum (a) or synthetic nasal medium 3 (SNM3) (b).
The control strains RP62A, 1457, and 12228 lack tarIJLM2 and were negative in the PCR reaction. Values represent means of two independent experiments. They were normalized for strongly and constitutively expressed housekeeping genes gyrB, rho, and tpiA. BD, below detection limit.
Extended Data Fig. 3 NMR and MS spectroscopy confirm that WTA from tarIJLM2-positive S. epidermidis strains contains both, glycerol (Gro) and ribitol (Rbo), while WTA of tarIJLM2-lacking strains contains only Gro.
a, Part of the HSQC-DEPT spectra of the methylene region of Rbo/Gro in the poly-RboP/GroP of the indicated WTA samples are shown. Signals corresponding to different substitutions of the polymers are shifted depending on the substitution. The presence of poly-RboP is clearly seen in the case of the WTA samples E73 wild type (wt), E73ΔIJLM2 (pRB-IJLM2), and 1457 (pRB-IJLM2) with signals at d 4.13;4.09/65.7, 4.07;3.98/67.8 and 4.20;4.16/64.7 (H/C), which are absent in the case of the other WTA samples. b, S. epidermidis E73 produces both, GroP (blue lines, [M + H]+ = 173.022 m/z) and RboP (red line, [M + H]+ = 233.042 m/z) in different growth phase as shown by LC-MS. c, another tarIJLM2-positive S. epidermidis E1 also produces both, GroP (blue lines, [M + H]+ = 173.022 m/z) and RboP (red line, [M + H]+ = 233.042 m/z) as shown by LC-MS.
Extended Data Fig. 4 RboP-WTA production does not increase the overall WTA amounts and does not affect S. epidermidis growth behavior or microscopic appearance.
a, Total WTA phosphate amount per cell wall dry weight of the indicated S. epidermidis strains. Means ± s.d. of three independent experiments are shown. None of the minor differences is significant. b, Growth curves of S. epidermidis E73 with or without tarIJLM2 in TSB. Means of three independent experiments are shown. c, Microscopic images of E73 with or without tarIJLM2. Scale bars: 5 µm.
Extended Data Fig. 5 Detailed phylogenetic distribution of tarIJLM2-5 among selected S. epidermidis clones and evolutionary relation of tarL from different gene clusters.
a, Maximum-likelihood phylogeny of 33 tarL genes from representative S. epidermidis (Se) and S. aureus (Sa) isolates and other Staphylococcus spp. isolates that carried tarIJL with or without tarM, including S. warneri (Sw), S. hominis (Sh), S. capitis (Sc), and uncharacterised Staphylococcus species (Ssp) (Source Data). The tarL2 gene found in S. epidermidis is more closely related to the two tarL genes in S. aureus than to S. epidermidis tarL1. The S. epidermidis tarL3, tarL4, and tarL5 genes, but not tarL2, were also found in other Staphylococcus species. Phylogenetic reconstruction was carried out using the maximum-likelihood program PhyML with a GTR model of nucleotide substitution, and support for the nodes was assessed using aBayes (see Methods). The tree was midpoint rooted. The scale bar denotes substitutions per variable sites. b, Maximum-likelihood phylogeny of 261 S. epidermidis isolates, comprising 25 tarIJLM-positive S. epidermidis isolates collected in this study, nine tarIJLM-positive S. epidermidis genomes from in the NCBI Reference Sequence Database (accessed July 3 2018), and a global collection of 227 S. epidermidis isolates originating from 96 healthcare institutions across 24 countries3 was inferred from an alignment of 39,298 single-nucleotide polymorphisms (Supplementary Table 2). Four tarIJLM-negative isolates (BPH0677, BPH0704, BPH0737, and SEI) were manually removed from the phylogeny due to their extreme divergence, and clades containing isolates with the same or closely related STs were collapsed to reduce complexity. The tarIJLM2 genes in E73 (Fig. 1b) were used as queries in BLASTN searches. The multilocus sequence types (STs) are indicated. Phylogenetic reconstruction was carried out using the maximum-likelihood program PhyML with a GTR model of nucleotide substitution, and support for the nodes was assessed using aBayes (see Methods). The tree was rooted according to work of Lee et al.3 The scale bar denotes substitutions per variable sites. Expanded subtrees illustrate the phylogenetic relationships of isolates belonging to ST5/ST87, ST10, ST23/single-locus variant (SLV), and ST2/ST188. The presence of tarIJLM2-5 is indicated in colors. A single ST23 isolate, VCU117, carried both, tarIJLM2 and tarIJLM3.
Extended Data Fig. 6 The gene cluster tarIJLM3 and tarIJLM4 lead to RboP-WTA synthesis in HA-MRSE ST2 and ST5 strains.
a, The tested tarIJLM3-positive (DAR1907 and RJ8, both ST2) and tarIJLM4-positive S. epidermidis strains (US06, ST5) contain both, GroP (blue lines, [M + H]+ = 173.022) and RboP (red line, [M + H]+ = 233.042) as WTA constituents; counts per second (CPS) measured by LC-MS are shown. b, Transduction with SaPIbov1 via Φ11. c, The binding rate of RboP-WTA-specific phage Φ11. d, The binding rate of GroP-WTA-specific phage Φ187. Means ± s.d. of three independent experiments are shown in (b-d).
Extended Data Fig. 7 RboP-WTA impairs S. epidermidis 1457 binding to A549 but promotes its binding to HUVECs.
a, The scavenger receptor inhibitor polyinosinic acid (poly-I) inhibits the increased binding of S. epidermidis E73∆IJLM2 to A549. b, RboP-WTA production in the GroP-WTA S. epidermidis 1457 decreases its binding to the human airway epithelial cell line A549 but promotes its binding to HUVECs (c). Means ± s.d. of three independent experiments are shown in (a-c). One-way ANOVA with Dunnett’s post-test (two-sided) was used to analyze significant differences vs. PBS condition in (a). Significant differences (P ≤ 0.05) were calculated by unpaired t-test between wt and the hybrid strain in (b) and (c).
Extended Data Fig. 8 The presence of plasmid pT181 does not affect the growth behavior of S. epidermidis E73 wild type (wt) compared to E73Δ IJLM2.
The indicated bacterial strains were grown in TSB medium. Means of two independent experiments are shown.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Rights and permissions
About this article
Cite this article
Du, X., Larsen, J., Li, M. et al. Staphylococcus epidermidis clones express Staphylococcus aureus-type wall teichoic acid to shift from a commensal to pathogen lifestyle. Nat Microbiol 6, 757–768 (2021). https://doi.org/10.1038/s41564-021-00913-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-021-00913-z
This article is cited by
-
Modulation of MRSA virulence gene expression by the wall teichoic acid enzyme TarO
Nature Communications (2023)
-
Cryptic susceptibility to penicillin/β-lactamase inhibitor combinations in emerging multidrug-resistant, hospital-adapted Staphylococcus epidermidis lineages
Nature Communications (2023)
-
Staphylococcus epidermidis and its dual lifestyle in skin health and infection
Nature Reviews Microbiology (2023)
-
Commensal production of a broad-spectrum and short-lived antimicrobial peptide polyene eliminates nasal Staphylococcus aureus
Nature Microbiology (2023)
-
Multi-species host range of staphylococcal phages isolated from wastewater
Nature Communications (2021)