Abstract
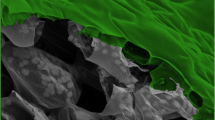
Phytophthora species, classified as oomycetes, are among the most destructive plant pathogens worldwide and pose a substantial threat to food security. Plant pathogens have developed various methods to breach the cuticle and walls of plant cells. For example, plant-pathogenic fungi use a ‘brute-force’ approach by producing a specialized and fortified invasion organ to generate invasive pressures. Unlike in fungi, the biomechanics of host invasion in oomycetes remains poorly understood. Here, using a combination of surface-deformation imaging, molecular-fracture sensors and modelling, we find that Phytophthora infestans, Phytophthora palmivora and Phytophthora capsici slice through the plant surface to gain entry into host tissues. To distinguish this mode of entry from the brute-force approach of fungi that use appressoria, we name this oomycete entry without appressorium formation ‘naifu’ invasion. Naifu invasion relies on polarized, non-concentric, force generation onto the surface at an oblique angle, which concentrates stresses at the site of invasion to enable surface breaching. Measurements of surface deformations during invasion of artificial substrates reveal a polarized mechanical geometry that we describe using a mathematical model. We confirm that the same mode of entry is used on real hosts. Naifu invasion uses actin-mediated polarity, surface adherence and turgor generation to enable Phytophthora to invade hosts without requiring specialized organs or vast turgor generation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw data associated with the figures in this manuscript have been archived at https://doi.org/10.4121/14115461 and are publicly available at https://data.4tu.nl/articles/dataset/Data_underlying_the_publication_Phytophthora_pathogens_exploit_slicing_action_for_host_invasion/14115461. Source data are provided with this paper.
Code availability
All data analysis algorithms used to perform the work presented in this manuscript were written by the authors using MatLab (v.2018b) and have been made publicly available at https://github.com/jorissprakel/Phytopthora_invasion.
References
Fisher, M. C. et al. Emerging fungal threats to animal, plant and ecosystem health. Nature 484, 186–194 (2012).
Savary, S. et al. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 3, 430–439 (2019).
Haverkort, A. J., Struik, P. C., Visser, R. G. F. & Jacobsen, E. Applied biotechnology to combat late blight in potato caused by Phytophthora infestans. Potato Res. 52, 249–264 (2009).
Haas, B. J. et al. Genome sequence and analysis of the Irish potato famine pathogen Phytophthora infestans. Nature 461, 393–398 (2009).
Kroon, L. P. N. M., Brouwer, H., de Cock, A. W. A. M. & Govers, F. The genus Phytophthora anno 2012. Phytopathology 102, 348–364 (2012).
Kamoun, S. et al. The Top 10 oomycete pathogens in molecular plant pathology. Mol. Plant Pathol. 16, 413–434 (2015).
Bastmeyer, M., Deising, H. B. & Bechinger, C. Force exertion in fungal infection. Annu Rev. Bioph Biom. 31, 321–341 (2002).
Latijnhouwers, M., de Wit, P. J. G. M. & Govers, F. Oomycetes and fungi: similar weaponry to attack plants. Trends Microbiol. 11, 462–469 (2003).
Talbot, N. J. Appressoria. Curr. Biol. 29, R144–R146 (2019).
Nezhad, A. S. & Geitmann, A. The cellular mechanics of an invasive lifestyle. J. Exp. Bot. 64, 4709–4728 (2013).
Foster, A. J. & Talbot, N. J. Getting a grip on blast. Nat. Microbiol 5, 1457–1458 (2020).
Howard, R. J. & Valent, B. Breaking and entering: host penetration by the fungal rice blast pathogen Magnaporthe grisea. Annu Rev. Microbiol. 50, 491–512 (1996).
Ryder, L. S. et al. A sensor kinase controls turgor-driven plant infection by the rice blast fungus. Nature 574, 423–427 (2019).
Howard, R. J., Ferrari, M. A., Roach, D. H. & Money, N. P. Penetration of hard substrates by a fungus employing enormous turgor pressures. Proc. Natl Acad. Sci. USA 88, 11281–11284 (1991).
Wilson, R. A. & Talbot, N. J. Under pressure: investigating the biology of plant infection by Magnaporthe oryzae. Nat. Rev. Microbiol. 7, 185–195 (2009).
Rocha, R. O., Elowsky, C., Pham, N. T. T. & Wilson, R. A. Spermine-mediated tight sealing of the Magnaporthe oryzae appressorial pore–rice leaf surface interface. Nat. Microbiol. 5, 1472–1480 (2020).
Chumley, F. G. Genetic analysis of melanin-deficient, nonpathogenic mutants of Magnaporthe grisea. Mol. Plant Microbe 3, 135 (1990).
Wang, T. et al. CgSCD1 is essential for melanin biosynthesis and pathogenicity of Colletotrichum gloeosporioides. Pathog. Basel Switz. 9, 141 (2020).
Wheeler, M. H. & Bell, A. A. Melanins and their importance in pathogenic fungi. Curr. Top. Med. Mycol. 2, 338–387 (1988).
Money, N. P., Davis, C. M. & Ravishankar, J. P. Biomechanical evidence for convergent evolution of the invasive growth process among fungi and oomycete water molds. Fungal Genet. Biol. 41, 872–876 (2004).
Meng, S., Torto-Alalibo, T., Chibucos, M. C., Tyler, B. M. & Dean, R. A. Common processes in pathogenesis by fungal and oomycete plant pathogens, described with Gene Ontology terms. BMC Microbiol. 9, S7 (2009).
Onoda, Y., Schieving, F. & Anten, N. P. R. A novel method of measuring leaf epidermis and mesophyll stiffness shows the ubiquitous nature of the sandwich structure of leaf laminas in broad-leaved angiosperm species. J. Exp. Bot. 66, 2487–2499 (2015).
Gibson, L. J. The hierarchical structure and mechanics of plant materials. J. R. Soc. Interface 9, 2749–2766 (2012).
Whisson, S. C., Boevink, P. C., Wang, S. & Birch, P. R. The cell biology of late blight disease. Curr. Opin. Microbiol. 34, 127–135 (2016).
Overdijk, E. J. R. et al. Interaction between the moss Physcomitrella patens and Phytophthora: a novel pathosystem for live‐cell imaging of subcellular defence. J. Microsc. 263, 171–180 (2016).
Pieterse, C. M. J., Risseeuw, E. P. & Davidse, L. C. An in planta induced gene of Phytophthora infestans codes for ubiquitin. Plant Mol. Biol. 17, 799–811 (1991).
Morris, B. M., Reid, B. & Gow, N. A. R. Electrotaxis of zoospores of Phytophthora palmivora at physiologically relevant field strengths. Plant Cell Environ. 15, 645–653 (1992).
Miao, J. et al. Resistance assessment for oxathiapiprolin in Phytophthora capsici and the detection of a point mutation (G769W) in PcORP1 that confers resistance. Front. Microbiol. 7, 615 (2016).
Reyssat, E., Tallinen, T., Merrer, M. L. & Mahadevan, L. Slicing softly with shear. Phys. Rev. Lett. 109, 244301 (2012).
Davis, D. A. et al. Force-induced activation of covalent bonds in mechanoresponsive polymeric materials. Nature 459, 68–72 (2009).
Celestine, A.-D. N. et al. Fracture-induced activation in mechanophore-linked, rubber toughened PMMA. Polymer 55, 4164–4171 (2014).
Bechinger, C. et al. Optical measurements of invasive forces exerted by appressoria of a plant pathogenic fungus. Science 285, 1896–1899 (1999).
Sun, Y., Tayagui, A., Shearer, H., Garrill, A. & Nock, V. A microfluidic platform with integrated sensing pillars for protrusive force measurements on Neurospora crassa. In 2018 IEEE Micro Electro Mechanical Systems 1116–1119 (IEEE, 2018).
Money, N. P. in Biology of the Fungal Cell (eds Howard, R. J. & Gow, N. A. R.) 237–249 (Springer, 2007).
Tayagui, A., Sun, Y., Collings, D. A., Garrill, A. & Nock, V. An elastomeric micropillar platform for the study of protrusive forces in hyphal invasion. Lab Chip 17, 3643–3653 (2017).
Heath, I. B. & Steinberg, G. Mechanisms of hyphal tip growth: tube dwelling amebae revisited. Fungal Genet. Biol. 28, 79–93 (1999).
Harold, F. M. Force and compliance: rethinking morphogenesis in walled cells. Fungal Genet. Biol. 37, 271–282 (2002).
Gossot, O. & Geitmann, A. Pollen tube growth: coping with mechanical obstacles involves the cytoskeleton. Planta 226, 405–416 (2007).
Ketelaar, T., Meijer, H. J. G., Spiekerman, M., Weide, R. & Govers, F. Effects of latrunculin B on the actin cytoskeleton and hyphal growth in Phytophthora infestans. Fungal Genet. Biol. 49, 1014–1022 (2012).
Robold, A. V. & Hardham, A. R. During attachment Phytophthora spores secrete proteins containing thrombospondin type 1 repeats. Curr. Genet. 47, 307–315 (2005).
Epstein, L. & Nicholson, R. L. in Biological Adhesives (eds Smith, A. M. & Callow, J. A.) 41–62 (Springer, 2006). https://doi.org/10.1007/978-3-540-31049-5_3
Hardham, A. R. & Shan, W. in Plant Relationships (ed. Deising, H. B.) 3–27 (2009).
Lee, S. & Vörös, J. An aqueous-based surface modification of poly(dimethylsiloxane) with poly(ethylene glycol) to prevent biofouling. Langmuir 21, 11957–11962 (2005).
Heuberger, M., Drobek, T. & Spencer, N. D. Interaction forces and morphology of a protein-resistant poly(ethylene glycol) layer. Biophys. J. 88, 495–504 (2005).
Marie, R., Beech, J. P., Vörös, J., Tegenfeldt, J. O. & Höök, F. Use of PLL-g-PEG in micro-fluidic devices for localizing selective and specific protein binding. Langmuir 22, 10103–10108 (2006).
MacDonald, E., Millward, L., Ravishankar, J. P. & Money, N. P. Biomechanical interaction between hyphae of two Pythium species (Oomycota) and host tissues. Fungal Genet. Biol. 37, 245–249 (2002).
Mendgen, K. & Deising, H. Infection structures of fungal plant pathogens—a cytological and physiological evaluation. New Phytol. 124, 193–213 (1993).
Huang, S., Vleeshouwers, V. G. A. A., Visser, R. G. F. & Jacobsen, E. An accurate in vitro assay for high-throughput disease testing of Phytophthora infestans in potato. Plant Dis. 89, 1263–1267 (2005).
Huang, W. R. H., Schol, C., Villanueva, S. L., Heidstra, R. & Joosten, M. H. A. J. Knocking out SOBIR1 in Nicotiana benthamiana abolishes functionality of transgenic receptor-like protein Cf-4. Plant Physiol. 185, kiaa047 (2020).
Boots, J. N. M., Fokkink, R., van der Gucht, J. & Kodger, T. E. Development of a multi-position indentation setup: mapping soft and patternable heterogeneously crosslinked polymer networks. Rev. Sci. Instrum. 90, 015108 (2019).
Johnson, K. L. Contact Mechanics (Cambridge Univ. Press, 1985).
Kots, K., Meijer, H. J. G., Bouwmeester, K., Govers, F. & Ketelaar, T. Filamentous actin accumulates during plant cell penetration and cell wall plug formation in Phytophthora infestans. Cell Mol. Life Sci. 74, 909–920 (2016).
Money, N. P. Osmotic pressure of aqueous polyethylene glycols: relationship between molecular weight and vapor pressure deficit. Plant Physiol. 91, 766–769 (1989).
Vleeshouwers, V. G. A. A. et al. A laboratory assay for Phytophthora infestans resistance in various solanum species reflects the field situation. Eur. J. Plant Pathol. 105, 241–250 (1999).
Vallieres, C. et al. Discovery of (meth)acrylate polymers that resist colonization by fungi associated with pathogenesis and biodeterioration. Sci. Adv. 6, eaba6574 (2020).
Acknowledgements
This research is supported by research programme ECHO with project number 712.016.001, financed by the Dutch Research Council (NWO) (to J. Bronkhorst and J.S.); NWO Science domain (NWO-ENW) project GSGT.GSGT.2018.024 (to M.K.); and the European Research Council (ERC) project CoG-SOFTBREAK (to J.v.d.G). We thank V. Vleeshouwers (WUR Plant Breeding) for supplying seedlings.
Author information
Authors and Affiliations
Contributions
J. Bronkhorst, T.K., F.G. and J.S. conceived and designed the project. J. Bronkhorst, M.K., S.v.V., J.M.C. and K.K. performed experimental work. J. Bronkhorst, S.v.V., J. Buijs and J.S. conceived image analysis routines and performed the analysis. J. Bronkhorst, J.v.d.G. and J.S. performed mathematical modelling. J. Bronkhorst, T.K., F.G. and J.S. wrote the paper with assistance and input of all coauthors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Nicholas Money, Richard Wilson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Spiropyran fracture sensor reveals locus of surface invasion by the pathogen.
Additional examples of time series that reveal the time and locus of crack nucleation by invasion of P. infestans - 14:3 eGFP into a force-sensor elastomer substrate. We define t = 0 min as the moment of crack nucleation. All scale bars represent 5 μm.
Extended Data Fig. 2 Additional time series of surface deformations during invasion of elastomer substrates by P. infestans – 14:3 eGFP.
Colors indicate surface deformations in micrometers. Red: positive deformations due to adhesion; blue: negative deformations due to indentation. Arrows in b point to the location of the cyst, showing weak indentation but no adhesion. Scale bars: 5 μm.
Extended Data Fig. 3 Additional time series of surface deformations during invasion by P. palmivora – wt.
Colors indicate surface deformations in micrometers. Red: positive deformations due to adhesion, blue: negative deformations due to indentation. Scale bars: 5 μm.
Extended Data Fig. 4 Time series of surface deformations during invasion on elastomer substrates by P. infestans – wt.
Colors indicate surface deformations in micrometers. Red: positive deformations due to adhesion, Blue: negative deformations due to indentation. Scale bars: 5 μm.
Extended Data Fig. 5 Time series of surface deformations pathogenic invasion on elastomer substrates by P. capsici – wt.
Colors indicate surface deformations in micrometers. Red: positive deformations due to adhesion, blue: negative deformations due to indentation. Scale bars: 5 μm.
Extended Data Fig. 6 Mechanical invasion exhibits three stages.
Two additional examples of total indentation force Fi as a function of time (t) for the mechanical invasion of P. infestans - 14:3 eGFP into 0.58 MPa elastomer substrates, exhibiting three temporal stages (see Fig. 1a): I) germ tube growth is observed but no detectable surface deformations occur, II) pressure application increases until a fracture nucleates, III) pathogen invades into the crack opening, during which the crack propagates at relatively constant force.
Extended Data Fig. 7 Leaf infection assays.
a, Detached leaf assay to test the effect of cytoskeletal disruption with LatB on P.inf-wt infectivity on Nb-sobir1 leaves. b, Detached leaf assay to test the effect of a non-adhesion coating on P.inf-wt infectivity on Nb-sobir1 leaves. Leaf images are deep-red images that reveal lesions as bright orange-white zones (a-b). Schematic illustrations created with BioRender.com. c, Infectivity, determined as the percentage of inoculated spots that show a clear lesion 7dpi, for various controls and treatments, showing clear reduction in pathogenicity for high LatB concentrations and the non-adhesion coating. Numbers N indicate the total number of leaves tested for each sample.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10, Methods, Discussion, Tables 1 and 2 and table of experimental details.
Supplementary Video 1
Three-dimensional reconstruction of P. infestans invasion into a potato stem: Several fluorescent pathogens (P. infestans 14-3-GFP, green) invade the stem of an etiolated potato cultivar (Bintje), with plant plastids shown in pink, at distinct oblique angles. Scale: the 3D volume has a size of 130 × 60 × 50 μm (l × w × h). Stem surface is located at approximately 35 μm from the base of the volume.
Supplementary Video 2
Side view of P. infestans invasion into an elastomer substrate: Time series of maximum-intensity projections in the xz-plane showing the invasion of P. infestans 14-3-GFP (green) into a fluorescent elastomer substrate (red).
Supplementary Video 3
Top view of P. infestans invasion into a fracture-reporting substrate: Time series of maximum-intensity projections of P. infestans 14-3-GFP (green) invasion in a fracture-reporting elastomer surface, which is covalently modified to carry the molecular mechanosensor spiropyran (magenta). This video reveals the initial stage of surface-fracture nucleation.
Supplementary Video 4
Surface-deformation maps for P. infestans 14-3-GFP invasion: Right: time series of reconstructred surface-deformation maps for P. infestans 14-3-GFP into a fluorescent elastomer substrate. Colour bar indicates the surface-deformation amplitude in μm. Left: corresponding GFP channel.
Supplementary Video 5
Surface-deformation maps for P. palmivora WT invasion: Right: time series of reconstructred surface-deformation maps for P. palmivora WT into a fluorescent elastomer substrate. Colour bar indicates the surface-deformation amplitude in μm.
Supplementary Video 6
Surface-deformation maps for P. infestans WT invasion: Right: time series of reconstructred surface-deformation maps for P. infestans WT into a fluorescent elastomer substrate. Colour bar indicates the surface-deformation amplitude in μm. Left: corresponding bright-field channel (black rectangle indicates the corresponding area for surface-deformation analysis).
Supplementary Video 7
Surface-deformation maps for P. capsici WT invasion: Right: time series of reconstructred surface-deformation maps for P. capsici WT into a fluorescent elastomer substrate. Colour bar indicates the surface-deformation amplitude in μm.
Supplementary Video 8
Fitting of surface-deformation profiles with model: Time series of surface-deformation profiles (symbols) and fits to the mathematical model (lines) for P. infestans 14-3-GFP invasion into a fluorescent elastomer substrate.
Source data
Source Data Fig. 1
Statistical source data for Fig. 1d–i.
Source Data Fig. 2
Statistical source data for Fig. 2d,e.
Source Data Fig. 3
Statistical source data for Fig. 3b,d,e.
Source Data Fig. 4
Statistical source data for Fig. 4k,l.
Source Data Extended Data Fig. 6
Statistical source data for Extended Data Fig. 6a,b.
Source Data Extended Data Fig. 7
Statistical source data for Extended Data Fig. 7c.
Rights and permissions
About this article
Cite this article
Bronkhorst, J., Kasteel, M., van Veen, S. et al. A slicing mechanism facilitates host entry by plant-pathogenic Phytophthora. Nat Microbiol 6, 1000–1006 (2021). https://doi.org/10.1038/s41564-021-00919-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-021-00919-7
This article is cited by
-
Genome-wide association analysis reveals a novel pathway mediated by a dual-TIR domain protein for pathogen resistance in cotton
Genome Biology (2023)
-
Cell geometry regulates tissue fracture
Nature Communications (2023)
-
A molecular mechanosensor for real-time visualization of appressorium membrane tension in Magnaporthe oryzae
Nature Microbiology (2023)
-
Phytophthora cuts into plant hosts
Nature Reviews Microbiology (2021)
-
2021 in review
Nature Microbiology (2021)